Mature Synchrotron Resources for Structural Biology (P30) Program
The Mature Synchrotron Resources (MSRs) for Structural Biology (new NOFO forecast PAR-26-029) connect biomedical researchers with state-of-the-art X-ray beamlines, hands-on user support, and expert training to generate and analyze structural and cellular biology data.
MSRs are accessible to all biomedical researchers whose projects are vetted through a peer review process. Explore the Available MSRs below to find available techniques and learn how to request access
What MSRs Provide

Support
MSRs provide assistance with data collection, processing, and analysis. This includes guidance on sample preparation and shipping, as well as support throughout the experimental process.

Access
User access is available on an equal-opportunity, nationwide basis, with supported techniques including fiber diffraction, macromolecular crystallography, small- and wide-angle X-ray scattering, soft X-ray tomography, X-ray spectroscopy and imaging, and X-ray footprinting with mass spectrometry.

Training
Training is available through on-site and remote sessions, workshops, and instructional videos.
Available MSRs
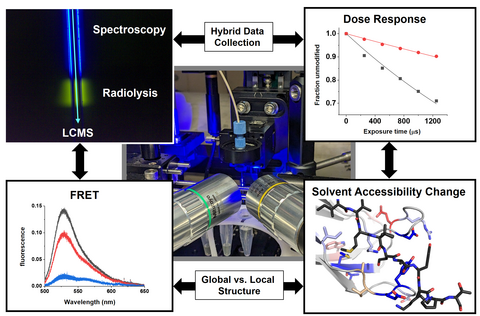
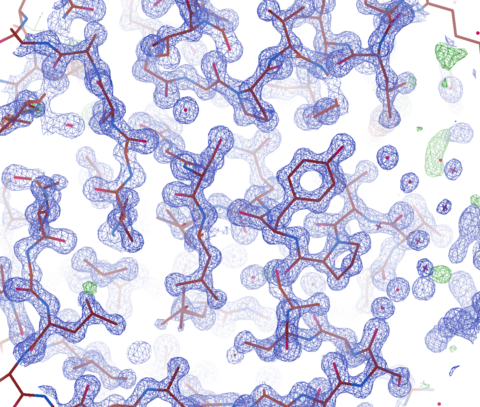
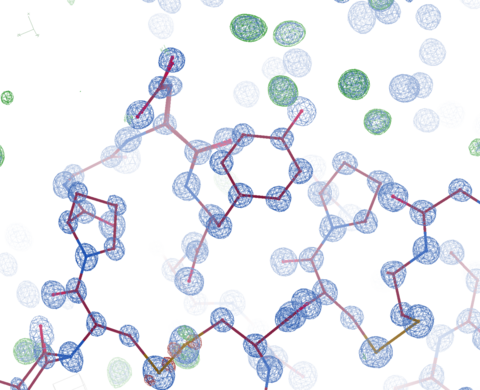
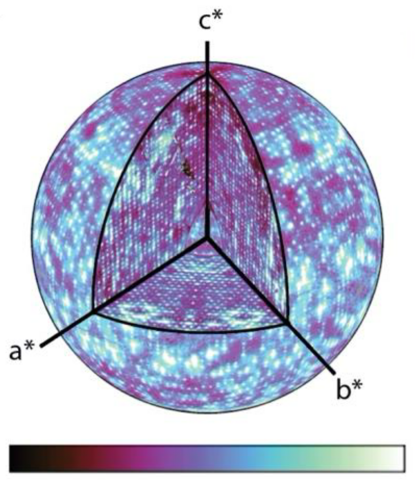
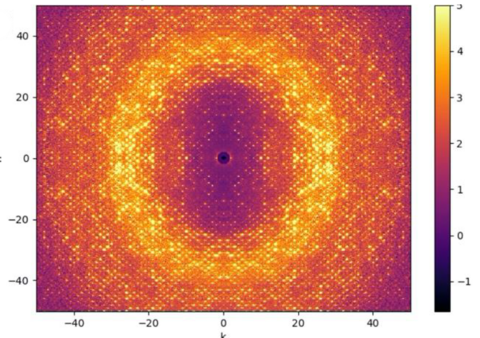
ALS-ENABLE @ ALS/Lawrence Berkeley National Laboratory
Macromolecular Crystallography, Small-Angle X-ray Scattering,
X-ray Footprinting with Mass Spectrometry.
Images courtesy of Dr. Paul Adams
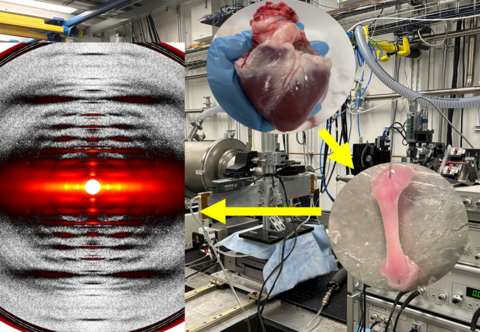
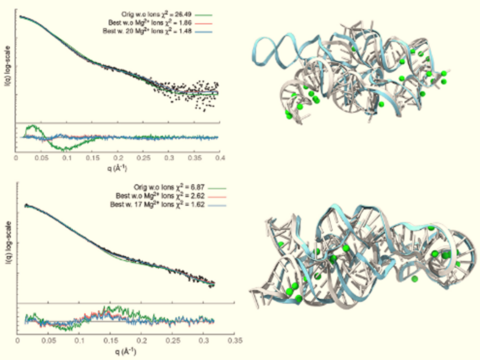
BioCAT @ APS/Argonne National Laboratory
Small-Angle X-ray Scattering, Fiber Diffraction
Images courtesy of Dr. Thomas Irving
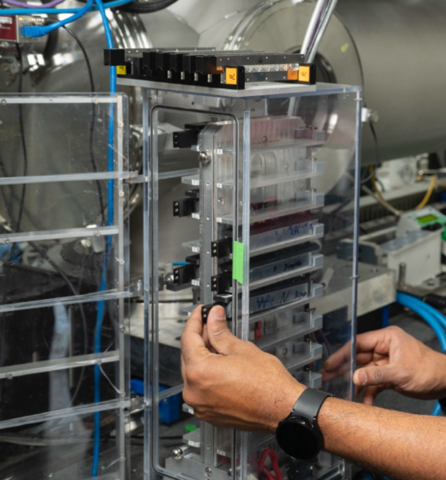
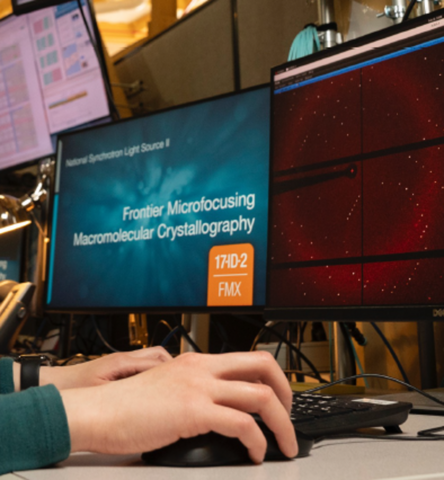
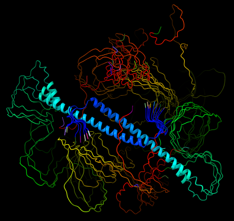
CBMS @ NSLS-II/Brookhaven National Laboratory
Macromolecular Crystallography, Small- and Wide-Angle X-ray
Scattering
Images courtesy of Dr. Sean McSweeney
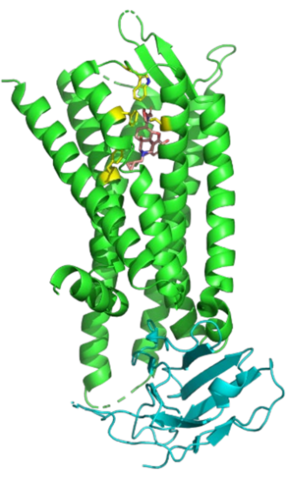
GM/CA @ APS/Argonne National Laboratory
Macromolecular Crystallography
Images courtesy of Dr. Robert Fischetti and Dr. Janet Smith
MacCHESS @ CHESS/Cornell University
Macromolecular Crystallography, Small-Angle X-ray Scattering
Images courtesy of Dr. Richard Cerione
NCXT @ ALS/Lawrence Berkeley National Laboratory
Soft X-ray Tomography
NE-CAT @ APS/Argonne National Laboratory
Macromolecular Crystallography
Images courtesy of Dr. Frank Murphy
SMB @ SSRL/SLAC National Accelerator Laboratory
Macromolecular Crystallography, Small-Angle X-ray Scattering,
X-ray Spectroscopy, and Imaging
Images courtesy of Dr. Keith Hodgson
For Current MSR Awardees
Guidance on preparing and submitting annual Research Performance Progress Reports (RPPRs) is available below.
RPPR Guidance for Funded Resources Word Template [DOCX]
Program Contacts
For more information or assistance contact the program staff: