Switch to List View
Image and Video Gallery
This is a searchable collection of scientific photos, illustrations, and videos. The images and videos in this gallery are licensed under Creative Commons Attribution Non-Commercial ShareAlike 3.0. This license lets you remix, tweak, and build upon this work non-commercially, as long as you credit and license your new creations under identical terms.
6612: Ciclo circadiano de un adolescente típico
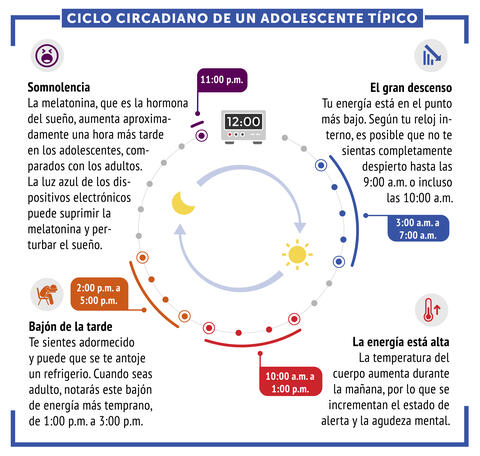
6612: Ciclo circadiano de un adolescente típico
Los ritmos circadianos son cambios físicos, mentales y conductuales que siguen un ciclo de 24 horas. Los ritmos circadianos típicos conducen a un nivel alto de energía durante la mitad del día (de 10 a.m. a 1 p.m.) y un bajón por la tarde. De noche, los ritmos circadianos hacen que la hormona melatonina aumente, lo que hace que la persona se sienta somnolienta.
Vea 6611 para la versión en inglés de esta infografía.
Vea 6611 para la versión en inglés de esta infografía.
NIGMS
View Media
2523: Plasma membrane
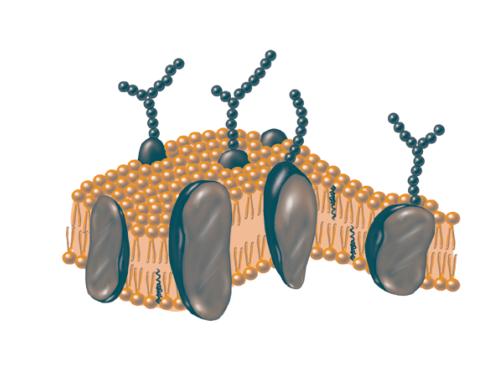
2523: Plasma membrane
The plasma membrane is a cell's protective barrier. See image 2524 for a labeled version of this illustration. Featured in The Chemistry of Health.
Crabtree + Company
View Media
5751: Genetically identical mycobacteria respond differently to antibiotic 1
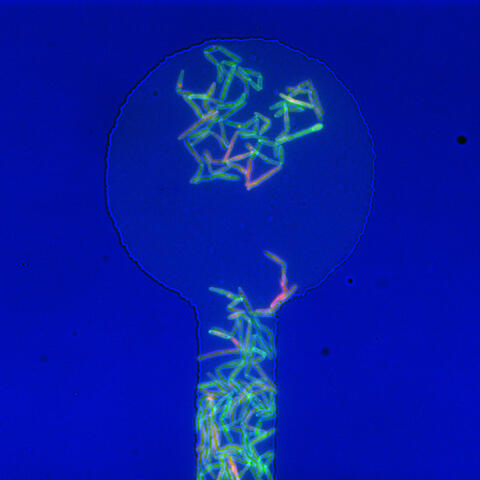
5751: Genetically identical mycobacteria respond differently to antibiotic 1
Antibiotic resistance in microbes is a serious health concern. So researchers have turned their attention to how bacteria undo the action of some antibiotics. Here, scientists set out to find the conditions that help individual bacterial cells survive in the presence of the antibiotic rifampicin. The research team used Mycobacterium smegmatis, a more harmless relative of Mycobacterium tuberculosis, which infects the lung and other organs and causes serious disease.
In this image, genetically identical mycobacteria are growing in a miniature growth chamber called a microfluidic chamber. Using live imaging, the researchers found that individual mycobacteria will respond differently to the antibiotic, depending on the growth stage and other timing factors. The researchers used genetic tagging with green fluorescent protein to distinguish cells that can resist rifampicin and those that cannot. With this gene tag, cells tolerant of the antibiotic light up in green and those that are susceptible in violet, enabling the team to monitor the cells' responses in real time.
To learn more about how the researchers studied antibiotic resistance in mycobacteria, see this news release from Tufts University. Related to video 5752.
In this image, genetically identical mycobacteria are growing in a miniature growth chamber called a microfluidic chamber. Using live imaging, the researchers found that individual mycobacteria will respond differently to the antibiotic, depending on the growth stage and other timing factors. The researchers used genetic tagging with green fluorescent protein to distinguish cells that can resist rifampicin and those that cannot. With this gene tag, cells tolerant of the antibiotic light up in green and those that are susceptible in violet, enabling the team to monitor the cells' responses in real time.
To learn more about how the researchers studied antibiotic resistance in mycobacteria, see this news release from Tufts University. Related to video 5752.
Bree Aldridge, Tufts University
View Media
6983: Genetic mosaicism in fruit flies
6983: Genetic mosaicism in fruit flies
Fat tissue from the abdomen of a genetically mosaic adult fruit fly. Genetic mosaicism means that the fly has cells with different genotypes even though it formed from a single zygote. This specific mosaicism results in accumulation of a critical fly adipokine (blue-green) within the fat tissue cells that have reduced expression a key nutrient sensing gene (in left panel). The dotted line shows the cells lacking the gene that is present and functioning in the rest of the cells. Nuclei are labelled in magenta. This image was captured using a confocal microscope and shows a maximum intensity projection of many slices.
Related to images 6982, 6984, and 6985.
Related to images 6982, 6984, and 6985.
Akhila Rajan, Fred Hutchinson Cancer Center
View Media

3424: White Poppy
3424: White Poppy
A white poppy. View cropped image of a poppy here 3423.
Judy Coyle, Donald Danforth Plant Science Center
View Media
3547: Master clock of the mouse brain
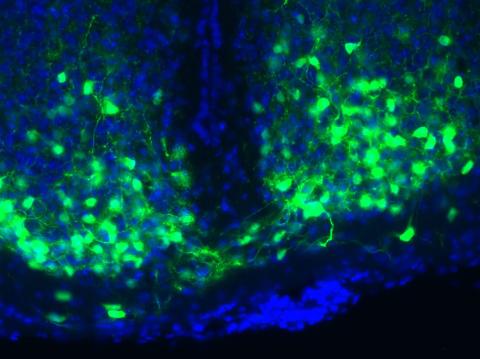
3547: Master clock of the mouse brain
An image of the area of the mouse brain that serves as the 'master clock,' which houses the brain's time-keeping neurons. The nuclei of the clock cells are shown in blue. A small molecule called VIP, shown in green, enables neurons in the central clock in the mammalian brain to synchronize.
Erik Herzog, Washington University in St. Louis
View Media
6850: Himastatin and bacteria
6850: Himastatin and bacteria
A model of the molecule himastatin overlaid on an image of Bacillus subtilis bacteria. Scientists first isolated himastatin from the bacterium Streptomyces himastatinicus, and the molecule shows antibiotic activity. The researchers who created this image developed a new, more concise way to synthesize himastatin so it can be studied more easily. They also tested the effects of himastatin and derivatives of the molecule on B. subtilis.
More information about the research that produced this image can be found in the Science paper “Total synthesis of himastatin” by D’Angelo et al.
Related to image 6848 and video 6851.
More information about the research that produced this image can be found in the Science paper “Total synthesis of himastatin” by D’Angelo et al.
Related to image 6848 and video 6851.
Mohammad Movassaghi, Massachusetts Institute of Technology.
View Media
3727: Zinc levels in a plant leaf
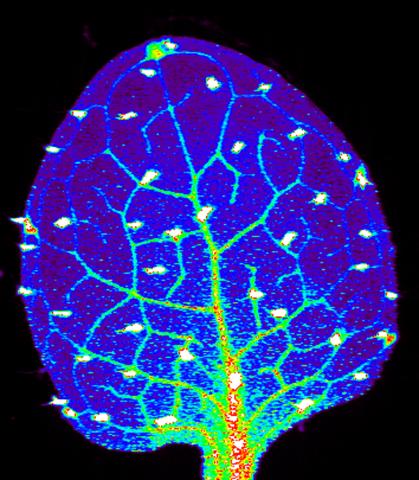
3727: Zinc levels in a plant leaf
Zinc is required for the function of more than 300 enzymes, including those that help regulate gene expression, in various organisms including humans. Researchers study how plants acquire, sequester and distribute zinc to find ways to increase the zinc content of crops to improve human health. Using synchrotron X-ray fluorescence technology, they created this heat map of zinc levels in an Arabidopsis thaliana plant leaf. This image is a winner of the 2015 FASEB Bioart contest and was featured in the NIH Director's blog.
Suzana Car, Dartmouth College
View Media
6772: Yeast cells responding to a glucose shortage
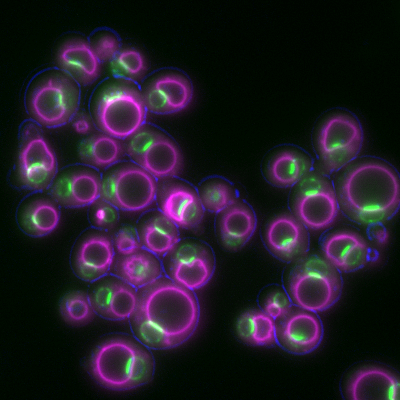
6772: Yeast cells responding to a glucose shortage
These yeast cells were exposed to a glucose (sugar) shortage. This caused the cells to compartmentalize HMGCR (green)—an enzyme involved in making cholesterol—to a patch on the nuclear envelope next to the vacuole/lysosome (purple). This process enhanced HMGCR activity and helped the yeast adapt to the glucose shortage. Researchers hope that understanding how yeast regulate cholesterol could ultimately lead to new ways to treat high cholesterol in people. This image was captured using a fluorescence microscope.
Mike Henne, University of Texas Southwestern Medical Center.
View Media
6601: Atomic-level structure of the HIV capsid
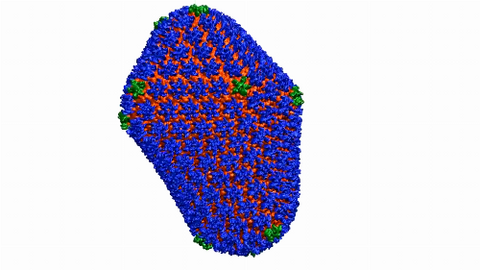
6601: Atomic-level structure of the HIV capsid
This animation shows atoms of the HIV capsid, the shell that encloses the virus's genetic material. Scientists determined the exact structure of the capsid using a variety of imaging techniques and analyses. They then entered this data into a supercomputer to produce this image. Related to image 3477.
Juan R. Perilla and the Theoretical and Computational Biophysics Group, University of Illinois at Urbana-Champaign
View Media
6604: Enzyme reaction
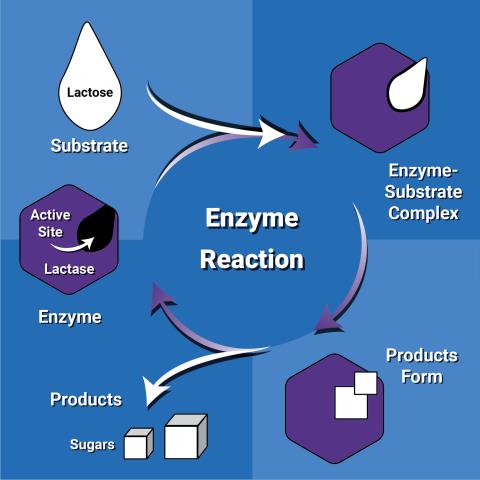
6604: Enzyme reaction
Enzymes speed up chemical reactions by reducing the amount of energy needed for the reactions. The substrate (lactose) binds to the active site of the enzyme (lactase) and is converted into products (sugars).
NIGMS
View Media

3425: Red Poppy

2750: Antibodies in silica honeycomb
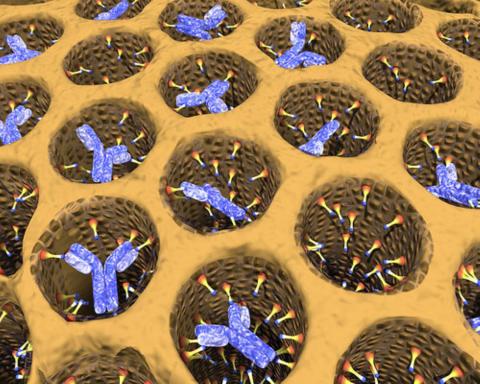
2750: Antibodies in silica honeycomb
Antibodies are among the most promising therapies for certain forms of cancer, but patients must take them intravenously, exposing healthy tissues to the drug and increasing the risk of side effects. A team of biochemists packed the anticancer antibodies into porous silica particles to deliver a heavy dose directly to tumors in mice.
Chenghong Lei, Pacific Northwest National Laboratory & Karl Erik Hellstrom, University of Washington
View Media
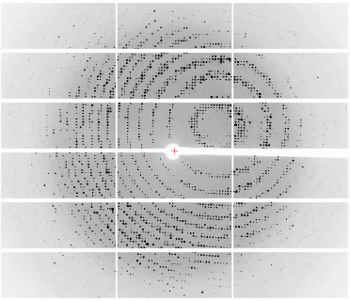
6765: X-ray diffraction pattern from a crystallized cefotaxime-CCD-1 complex
6765: X-ray diffraction pattern from a crystallized cefotaxime-CCD-1 complex
CCD-1 is an enzyme produced by the bacterium Clostridioides difficile that helps it resist antibiotics. Researchers crystallized complexes where a CCD-1 molecule and a molecule of the antibiotic cefotaxime were bound together. Then, they shot X-rays at the complexes to determine their structure—a process known as X-ray crystallography. This image shows the X-ray diffraction pattern of a complex.
Related to images 6764, 6766, and 6767.
Related to images 6764, 6766, and 6767.
Keith Hodgson, Stanford University.
View Media
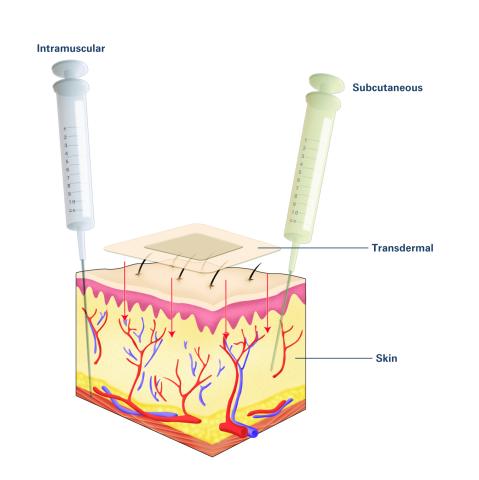
2532: Drugs enter skin (with labels)
2532: Drugs enter skin (with labels)
Drugs enter different layers of skin via intramuscular, subcutaneous, or transdermal delivery methods. See image 2531 for an unlabeled version of this illustration. Featured in Medicines By Design.
Crabtree + Company
View Media
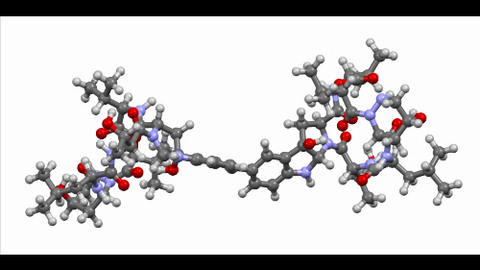
6851: Himastatin, 360-degree view
6851: Himastatin, 360-degree view
A 360-degree view of the molecule himastatin, which was first isolated from the bacterium Streptomyces himastatinicus. Himastatin shows antibiotic activity. The researchers who created this video developed a new, more concise way to synthesize himastatin so it can be studied more easily.
More information about the research that produced this video can be found in the Science paper “Total synthesis of himastatin” by D’Angelo et al.
Related to images 6848 and 6850.
More information about the research that produced this video can be found in the Science paper “Total synthesis of himastatin” by D’Angelo et al.
Related to images 6848 and 6850.
Mohammad Movassaghi, Massachusetts Institute of Technology.
View Media
2790: Anti-tumor drug ecteinascidin 743 (ET-743) with hydrogens 01
2790: Anti-tumor drug ecteinascidin 743 (ET-743) with hydrogens 01
Ecteinascidin 743 (ET-743, brand name Yondelis), was discovered and isolated from a sea squirt, Ecteinascidia turbinata, by NIGMS grantee Kenneth Rinehart at the University of Illinois. It was synthesized by NIGMS grantees E.J. Corey and later by Samuel Danishefsky. Multiple versions of this structure are available as entries 2790-2797.
Timothy Jamison, Massachusetts Institute of Technology
View Media
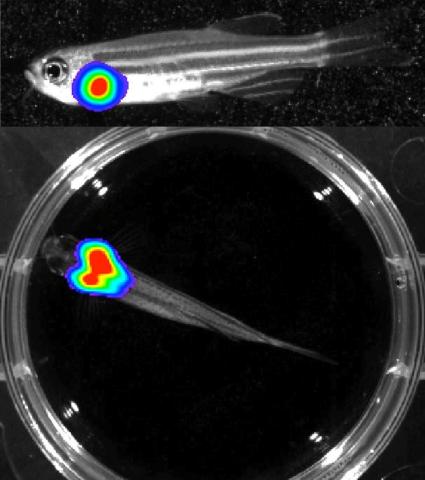
3556: Bioluminescent imaging in adult zebrafish - lateral and overhead view
3556: Bioluminescent imaging in adult zebrafish - lateral and overhead view
Luciferase-based imaging enables visualization and quantification of internal organs and transplanted cells in live adult zebrafish. In this image, a cardiac muscle-restricted promoter drives firefly luciferase expression. This is the lateral and overhead (Bottom) view.
For imagery of the overhead view go to 3557.
For imagery of the lateral view go to 3558.
For more information about the illumated area go to 3559.
For imagery of the overhead view go to 3557.
For imagery of the lateral view go to 3558.
For more information about the illumated area go to 3559.
Kenneth Poss, Duke University
View Media
3787: In vitro assembly of a cell-signaling pathway
3787: In vitro assembly of a cell-signaling pathway
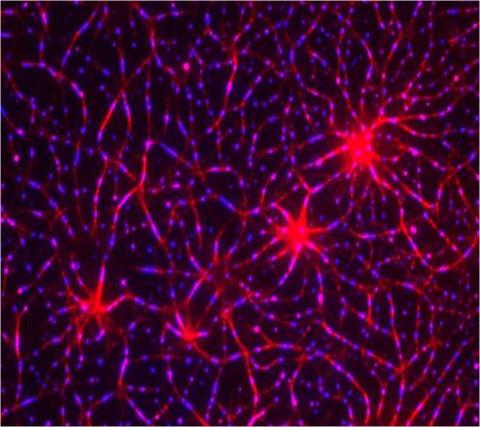
T cells are white blood cells that are important in defending the body against bacteria, viruses and other pathogens. Each T cell carries proteins, called T-cell receptors, on its surface that are activated when they come in contact with an invader. This activation sets in motion a cascade of biochemical changes inside the T cell to mount a defense against the invasion. Scientists have been interested for some time what happens after a T-cell receptor is activated. One obstacle has been to study how this signaling cascade, or pathway, proceeds inside T cells.
In this image, researchers have created a T-cell receptor pathway consisting of 12 proteins outside the cell on an artificial membrane. The image shows two key steps during the signaling process: clustering of a protein called linker for activation of T cells (LAT) (blue) and polymerization of the cytoskeleton protein actin (red). The findings show that the T-cell receptor signaling proteins self-organize into separate physical and biochemical compartments. This new system of studying molecular pathways outside the cells will enable scientists to better understand how the immune system combats microbes or other agents that cause infection.
To learn more how researchers assembled this T-cell receptor pathway, see this press release from HHMI's Marine Biological Laboratory Whitman Center. Related to video 3786.
In this image, researchers have created a T-cell receptor pathway consisting of 12 proteins outside the cell on an artificial membrane. The image shows two key steps during the signaling process: clustering of a protein called linker for activation of T cells (LAT) (blue) and polymerization of the cytoskeleton protein actin (red). The findings show that the T-cell receptor signaling proteins self-organize into separate physical and biochemical compartments. This new system of studying molecular pathways outside the cells will enable scientists to better understand how the immune system combats microbes or other agents that cause infection.
To learn more how researchers assembled this T-cell receptor pathway, see this press release from HHMI's Marine Biological Laboratory Whitman Center. Related to video 3786.
Xiaolei Su, HHMI Whitman Center of the Marine Biological Laboratory
View Media

5895: Bioluminescence in a Tube
5895: Bioluminescence in a Tube
Details about the basic biology and chemistry of the ingredients that produce bioluminescence are allowing scientists to harness it as an imaging tool. Credit: Nathan Shaner, Scintillon Institute.
From Biomedical Beat article July 2017: Chasing Fireflies—and Better Cellular Imaging Techniques
From Biomedical Beat article July 2017: Chasing Fireflies—and Better Cellular Imaging Techniques
Nathan Shaner, Scintillon Institute
View Media
3557: Bioluminescent imaging in adult zebrafish - overhead view
3557: Bioluminescent imaging in adult zebrafish - overhead view
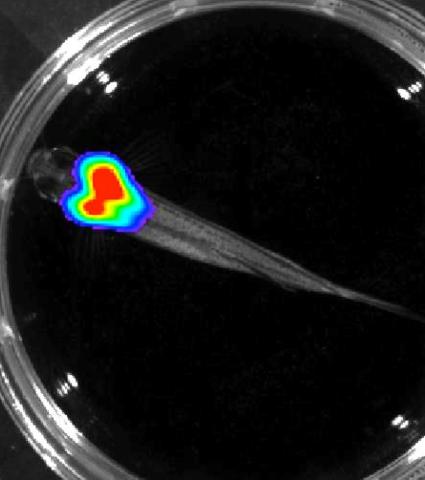
Luciferase-based imaging enables visualization and quantification of internal organs and transplanted cells in live adult zebrafish. In this image, a cardiac muscle-restricted promoter drives firefly luciferase expression.
For imagery of both the lateral and overhead view go to 3556.
For imagery of the lateral view go to 3558.
For more information about the illumated area go to 3559.
For imagery of both the lateral and overhead view go to 3556.
For imagery of the lateral view go to 3558.
For more information about the illumated area go to 3559.
Kenneth Poss, Duke University
View Media
3412: Active Site of E. coli response regulator PhoB
3412: Active Site of E. coli response regulator PhoB
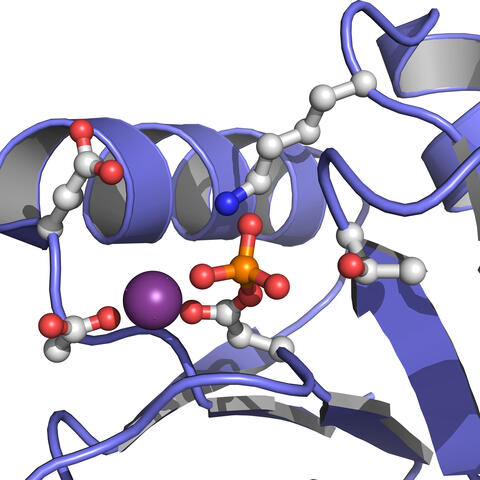
Active site of E. coli response regulator PhoB.
Ann Stock, Rutgers University
View Media
6613: Circadian rhythms and the SCN
6613: Circadian rhythms and the SCN
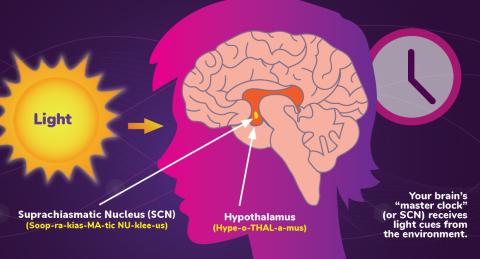
Circadian rhythms are physical, mental, and behavioral changes that follow a 24-hour cycle. Circadian rhythms are influenced by light and regulated by the brain’s suprachiasmatic nucleus (SCN), sometimes referred to as a master clock. Learn more in NIGMS’ circadian rhythms fact sheet. See 6614 for the Spanish version of this infographic.
NIGMS
View Media
6779: Brain waves of a patient anesthetized with propofol
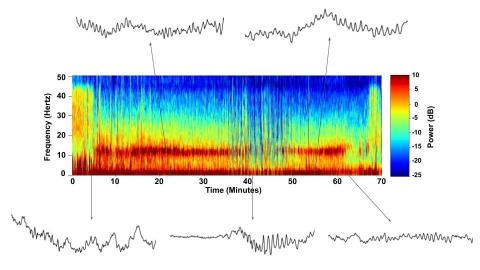
6779: Brain waves of a patient anesthetized with propofol
A representation of a patient’s brain waves after receiving the anesthetic propofol. All anesthetics create brain wave changes that vary depending on the patient’s age and the type and dose of anesthetic used. These changes are visible in raw electroencephalogram (EEG) readings, but they’re easier to interpret using a spectrogram where the signals are broken down by time (x-axis), frequency (y-axis), and power (color scale). This spectrogram shows the changes in brain waves before, during, and after propofol-induced anesthesia. The patient is unconscious from minute 5, upon propofol administration, through minute 69 (change in power and frequency). But, between minutes 35 and 48, the patient fell into a profound state of unconsciousness (disappearance of dark red oscillations between 8 to 12 Hz), which required the anesthesiologist to adjust the rate of propofol administration. The propofol was stopped at minute 62 and the patient woke up around minute 69.
Emery N. Brown, M.D., Ph.D., Massachusetts General Hospital/Harvard Medical School, Picower Institute for Learning and Memory, and Massachusetts Institute of Technology.
View Media
3480: Cancer Cells Glowing from Luciferin
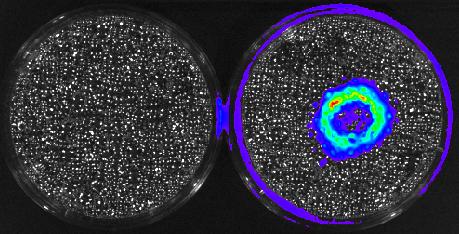
3480: Cancer Cells Glowing from Luciferin
The activator cancer cell culture, right, contains a chemical that causes the cells to emit light when in the presence of immune cells.
Mark Sellmyer, Stanford University School of Medicine
View Media

2531: Drugs enter skin
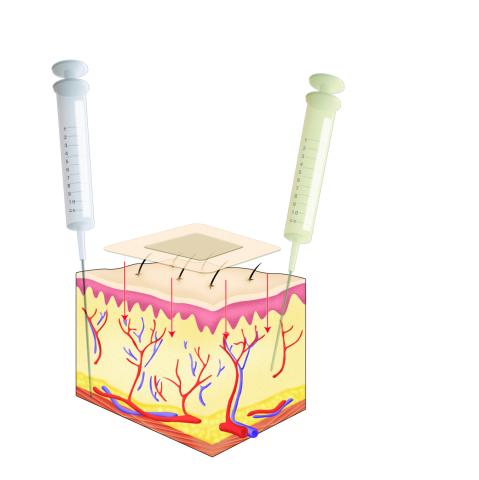
2531: Drugs enter skin
Drugs enter different layers of skin via intramuscular, subcutaneous, or transdermal delivery methods. See image 2532 for a labeled version of this illustration. Featured in Medicines By Design.
Crabtree + Company
View Media
2535: Kinases (with labels)
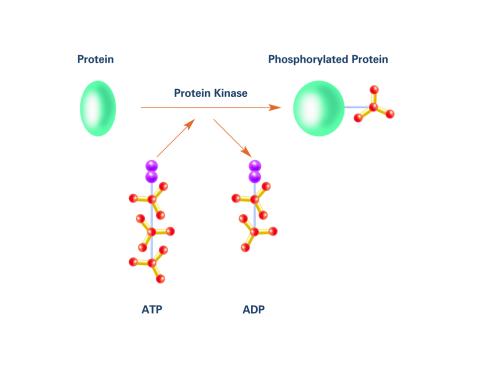
2535: Kinases (with labels)
Kinases are enzymes that add phosphate groups (red-yellow structures) to proteins (green), assigning the proteins a code. In this reaction, an intermediate molecule called ATP (adenosine triphosphate) donates a phosphate group from itself, becoming ADP (adenosine diphosphate). See image 2534 for an unlabeled version of this illustration. Featured in Medicines By Design.
Crabtree + Company
View Media
2530: Aspirin (with labels)
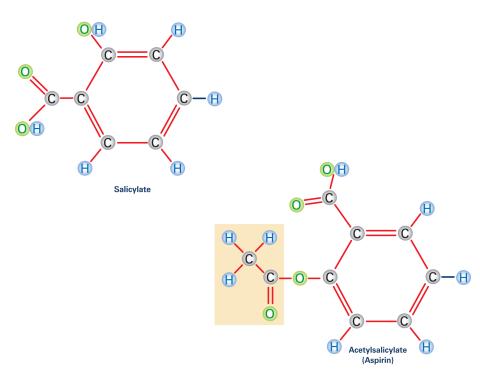
2530: Aspirin (with labels)
Acetylsalicylate (bottom) is the aspirin of today. Adding a chemical tag called an acetyl group (shaded box, bottom) to a molecule derived from willow bark (salicylate, top) makes the molecule less acidic (and easier on the lining of the digestive tract), but still effective at relieving pain. See image 2529 for an unlabeled version of this illustration. Featured in Medicines By Design.
Crabtree + Company
View Media
6602: See how immune cell acid destroys bacterial proteins
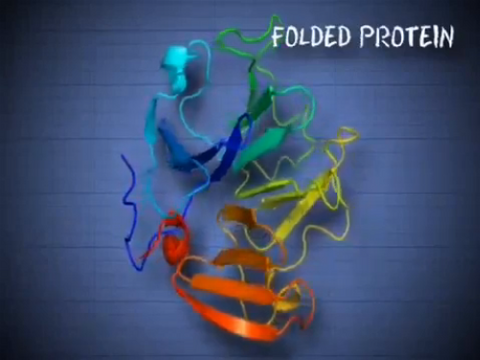
6602: See how immune cell acid destroys bacterial proteins
This animation shows the effect of exposure to hypochlorous acid, which is found in certain types of immune cells, on bacterial proteins. The proteins unfold and stick to one another, leading to cell death.
American Chemistry Council
View Media
3491: Kinesin moves cellular cargo
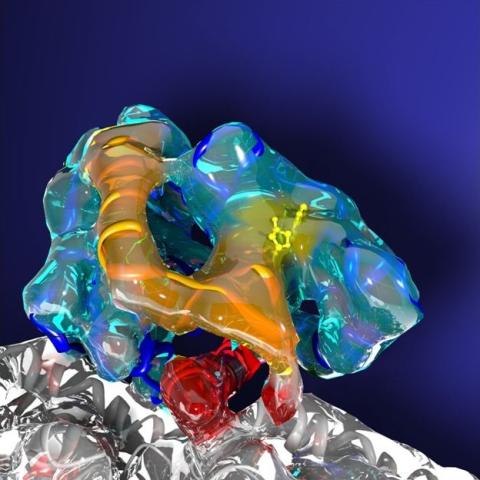
3491: Kinesin moves cellular cargo
A protein called kinesin (blue) is in charge of moving cargo around inside cells and helping them divide. It's powered by biological fuel called ATP (bright yellow) as it scoots along tube-like cellular tracks called microtubules (gray).
Charles Sindelar, Yale University
View Media
6762: CCP enzyme
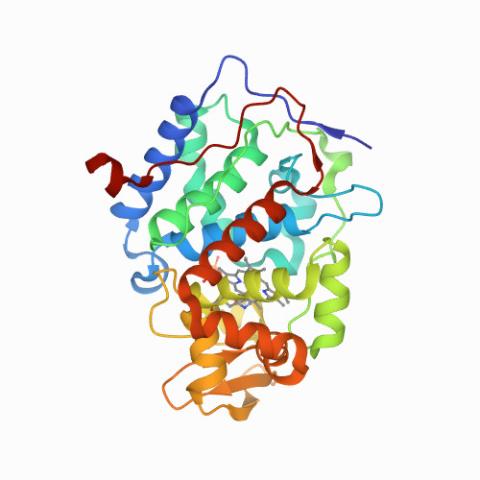
6762: CCP enzyme
The enzyme CCP is found in the mitochondria of baker’s yeast. Scientists study the chemical reactions that CCP triggers, which involve a water molecule, iron, and oxygen. This structure was determined using an X-ray free electron laser.
Protein Data Bank.
View Media

2735: Network Map
2735: Network Map
This network map shows the overlap (green) between the long QT syndrome (yellow) and epilepsy (blue) protein-interaction neighborhoods located within the human interactome. Researchers have learned to integrate genetic, cellular and clinical information to find out why certain medicines can trigger fatal heart arrhythmias. Featured in Computing Life magazine.
Seth Berger, Mount Sinai School of Medicine
View Media
2746: Active site of sulfite oxidase
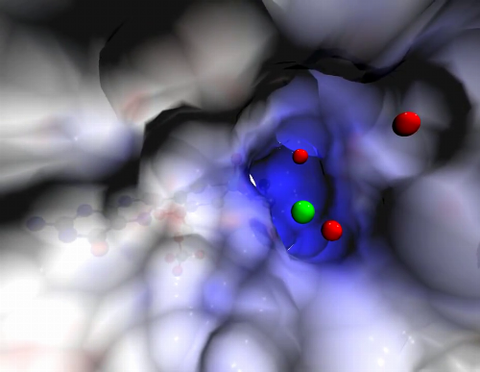
2746: Active site of sulfite oxidase
Sulfite oxidase is an enzyme that is essential for normal neurological development in children. This video shows the active site of the enzyme and its molybdenum cofactor visible as a faint ball-and-stick representation buried within the protein. The positively charged channel (blue) at the active site contains a chloride ion (green) and three water molecules (red). As the protein oscillates, one can see directly down the positively charged channel. At the bottom is the molybdenum atom of the active site (light blue) and its oxo group (red) that is transferred to sulfite to form sulfate in the catalytic reaction.
John Enemark, University of Arizona
View Media

2579: Bottles of warfarin
2579: Bottles of warfarin
In 2007, the FDA modified warfarin's label to indicate that genetic makeup may affect patient response to the drug. The widely used blood thinner is sold under the brand name Coumadin®. Scientists involved in the NIH Pharmacogenetics Research Network are investigating whether genetic information can be used to improve optimal dosage prediction for patients.
Alisa Machalek, NIGMS/NIH
View Media
2518: ATP synthase (with labels)
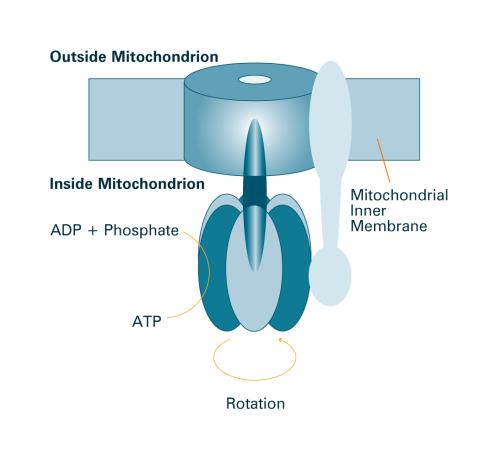
2518: ATP synthase (with labels)
The world's smallest motor, ATP synthase, generates energy for the cell. See image 2517 for an unlabeled version of this illustration. Featured in The Chemistry of Health.
Crabtree + Company
View Media
2520: Bond types (with labels)
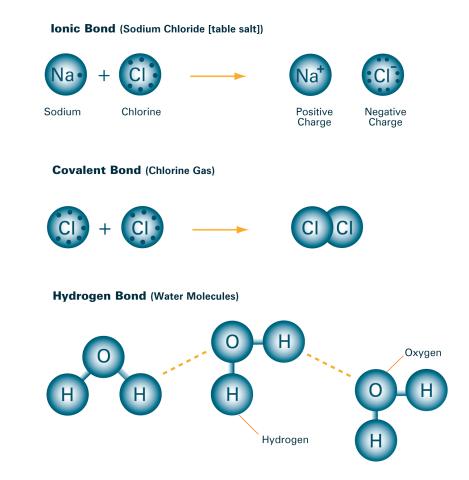
2520: Bond types (with labels)
Ionic and covalent bonds hold molecules, like sodium chloride and chlorine gas, together. Hydrogen bonds among molecules, notably involving water, also play an important role in biology. See image 2519 for an unlabeled version of this illustration. Featured in The Chemistry of Health.
Crabtree + Company
View Media
6802: Antibiotic-surviving bacteria
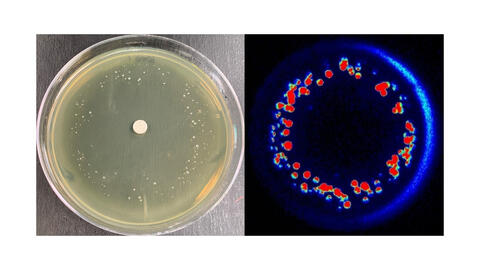
6802: Antibiotic-surviving bacteria
Colonies of bacteria growing despite high concentrations of antibiotics. These colonies are visible both by eye, as seen on the left, and by bioluminescence imaging, as seen on the right. The bioluminescent color indicates the metabolic activity of these bacteria, with their red centers indicating high metabolism.
More information about the research that produced this image can be found in the Antimicrobial Agents and Chemotherapy paper “Novel aminoglycoside-tolerant phoenix colony variants of Pseudomonas aeruginosa” by Sindeldecker et al.
More information about the research that produced this image can be found in the Antimicrobial Agents and Chemotherapy paper “Novel aminoglycoside-tolerant phoenix colony variants of Pseudomonas aeruginosa” by Sindeldecker et al.
Paul Stoodley, The Ohio State University.
View Media

3786: Movie of in vitro assembly of a cell-signaling pathway
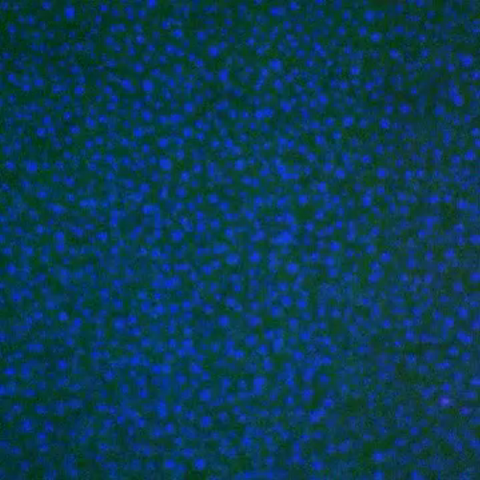
3786: Movie of in vitro assembly of a cell-signaling pathway
T cells are white blood cells that are important in defending the body against bacteria, viruses and other pathogens. Each T cell carries proteins, called T-cell receptors, on its surface that are activated when they come in contact with an invader. This activation sets in motion a cascade of biochemical changes inside the T cell to mount a defense against the invasion. Scientists have been interested for some time what happens after a T-cell receptor is activated. One obstacle has been to study how this signaling cascade, or pathway, proceeds inside T cells.
In this video, researchers have created a T-cell receptor pathway consisting of 12 proteins outside the cell on an artificial membrane. The video shows three key steps during the signaling process: phosphorylation of the T-cell receptor (green), clustering of a protein called linker for activation of T cells (LAT) (blue) and polymerization of the cytoskeleton protein actin (red). The findings show that the T-cell receptor signaling proteins self-organize into separate physical and biochemical compartments. This new system of studying molecular pathways outside the cells will enable scientists to better understand how the immune system combats microbes or other agents that cause infection.
To learn more how researchers assembled this T-cell receptor pathway, see this press release from HHMI's Marine Biological Laboratory Whitman Center. Related to image 3787.
In this video, researchers have created a T-cell receptor pathway consisting of 12 proteins outside the cell on an artificial membrane. The video shows three key steps during the signaling process: phosphorylation of the T-cell receptor (green), clustering of a protein called linker for activation of T cells (LAT) (blue) and polymerization of the cytoskeleton protein actin (red). The findings show that the T-cell receptor signaling proteins self-organize into separate physical and biochemical compartments. This new system of studying molecular pathways outside the cells will enable scientists to better understand how the immune system combats microbes or other agents that cause infection.
To learn more how researchers assembled this T-cell receptor pathway, see this press release from HHMI's Marine Biological Laboratory Whitman Center. Related to image 3787.
Xiaolei Su, HHMI Whitman Center of the Marine Biological Laboratory
View Media
2525: Activation energy
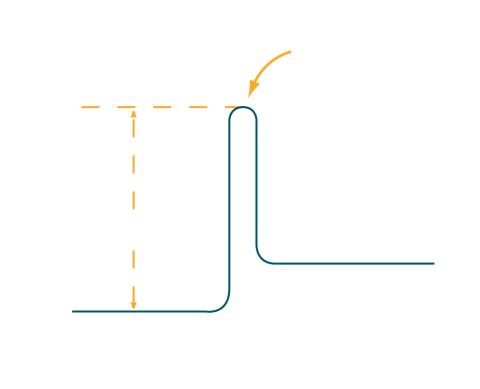
2525: Activation energy
To become products, reactants must overcome an energy hill. See image 2526 for a labeled version of this illustration. Featured in The Chemistry of Health.
Crabtree + Company
View Media
2797: Anti-tumor drug ecteinascidin 743 (ET-743), structure without hydrogens 04
2797: Anti-tumor drug ecteinascidin 743 (ET-743), structure without hydrogens 04
Ecteinascidin 743 (ET-743, brand name Yondelis), was discovered and isolated from a sea squirt, Ecteinascidia turbinata, by NIGMS grantee Kenneth Rinehart at the University of Illinois. It was synthesized by NIGMS grantees E.J. Corey and later by Samuel Danishefsky. Multiple versions of this structure are available as entries 2790-2797.
Timothy Jamison, Massachusetts Institute of Technology
View Media
2519: Bond types
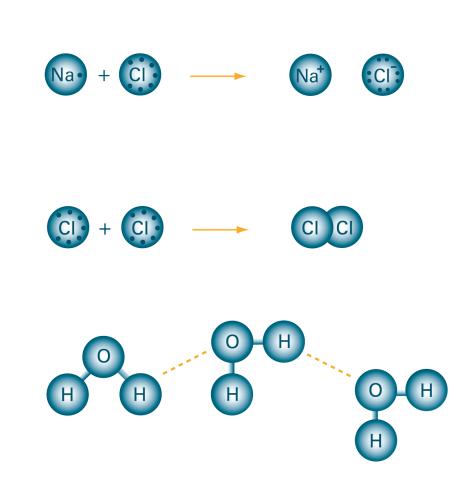
2519: Bond types
Ionic and covalent bonds hold molecules, like sodium chloride and chlorine gas, together. Hydrogen bonds among molecules, notably involving water, also play an important role in biology. See image 2520 for a labeled version of this illustration. Featured in The Chemistry of Health.
Crabtree + Company
View Media
3720: Cas4 nuclease protein structure
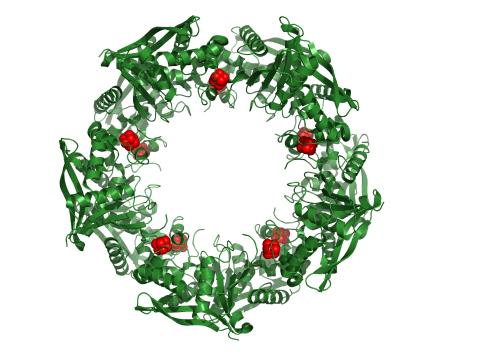
3720: Cas4 nuclease protein structure
This wreath represents the molecular structure of a protein, Cas4, which is part of a system, known as CRISPR, that bacteria use to protect themselves against viral invaders. The green ribbons show the protein's structure, and the red balls show the location of iron and sulfur molecules important for the protein's function. Scientists harnessed Cas9, a different protein in the bacterial CRISPR system, to create a gene-editing tool known as CRISPR-Cas9. Using this tool, researchers are able to study a range of cellular processes and human diseases more easily, cheaply and precisely. In December, 2015, Science magazine recognized the CRISPR-Cas9 gene-editing tool as the "breakthrough of the year." Read more about Cas4 in the December 2015 Biomedical Beat post A Holiday-Themed Image Collection.
Fred Dyda, NIDDK
View Media
3559: Bioluminescent imaging in adult zebrafish 04
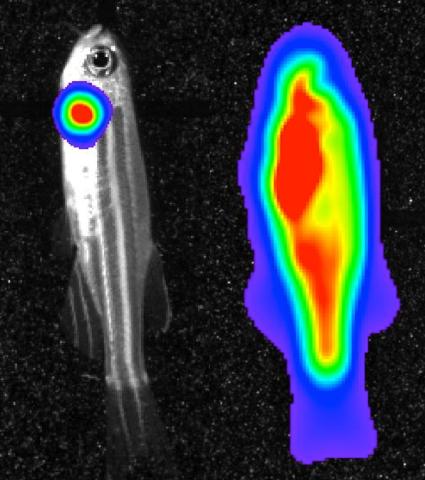
3559: Bioluminescent imaging in adult zebrafish 04
Luciferase-based imaging enables visualization and quantification of internal organs and transplanted cells in live adult zebrafish. This image shows how luciferase-based imaging could be used to visualize the heart for regeneration studies (left), or label all tissues for stem cell transplantation (right).
For imagery of both the lateral and overhead view go to 3556.
For imagery of the overhead view go to 3557.
For imagery of the lateral view go to 3558.
View Media
For imagery of both the lateral and overhead view go to 3556.
For imagery of the overhead view go to 3557.
For imagery of the lateral view go to 3558.
3314: Human opioid receptor structure superimposed on poppy
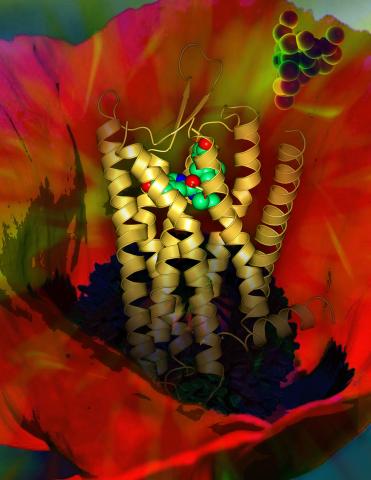
3314: Human opioid receptor structure superimposed on poppy
Opioid receptors on the surfaces of brain cells are involved in pleasure, pain, addiction, depression, psychosis, and other conditions. The receptors bind to both innate opioids and drugs ranging from hospital anesthetics to opium. Researchers at The Scripps Research Institute, supported by the NIGMS Protein Structure Initiative, determined the first three-dimensional structure of a human opioid receptor, a kappa-opioid receptor. In this illustration, the submicroscopic receptor structure is shown while bound to an agonist (or activator). The structure is superimposed on a poppy flower, the source of opium.
Raymond Stevens, The Scripps Research Institute
View Media
5888: Independence Day
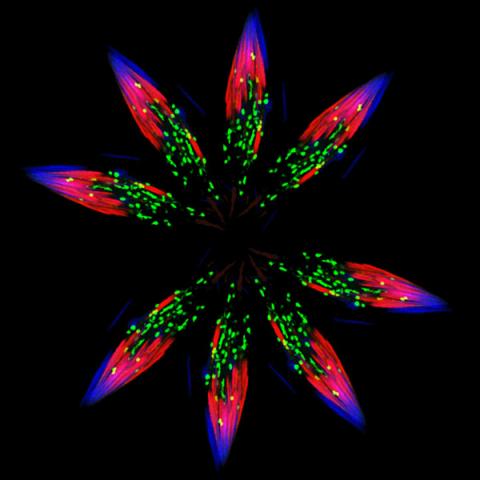
5888: Independence Day
This graphic that resembles a firework was created from a picture of a fruit fly spermatid. This fruit fly spermatid recycles various molecules, including malformed or damaged proteins. Actin filaments (red) in the cell draw unwanted proteins toward a barrel-shaped structure called the proteasome (green clusters), which degrades the molecules into their basic parts for re-use.
Sigi Benjamin-Hong, Rockefeller University
View Media
2534: Kinases
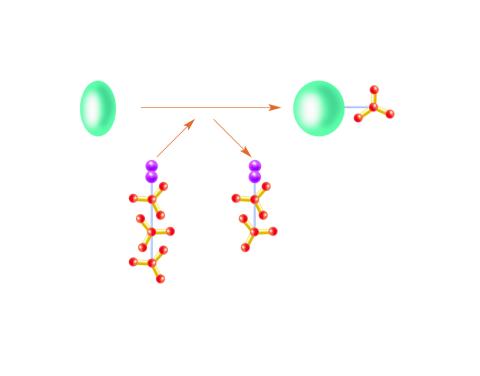
2534: Kinases
Kinases are enzymes that add phosphate groups (red-yellow structures) to proteins (green), assigning the proteins a code. In this reaction, an intermediate molecule called ATP (adenosine triphosphate) donates a phosphate group from itself, becoming ADP (adenosine diphosphate). See image 2535 for a labeled version of this illustration. Featured in Medicines By Design.
Crabtree + Company
View Media
2507: Carbon building blocks (with examples)
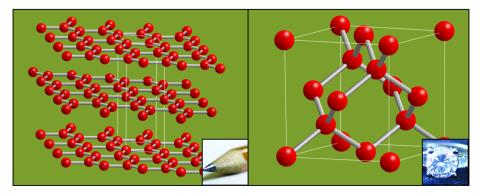
2507: Carbon building blocks (with examples)
The arrangement of identical molecular components can make a dramatic difference. For example, carbon atoms can be arranged into dull graphite (left) or sparkly diamonds (right). See image 2506 for an illustration without examples.
Crabtree + Company
View Media
3326: Cytochrome structure with anticancer drug
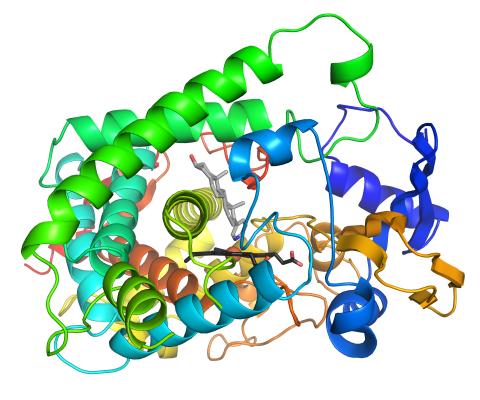
3326: Cytochrome structure with anticancer drug
This image shows the structure of the CYP17A1 enzyme (ribbons colored from blue N-terminus to red C-terminus), with the associated heme colored black. The prostate cancer drug abiraterone is colored gray. Cytochrome P450 enzymes bind to and metabolize a variety of chemicals, including drugs. Cytochrome P450 17A1 also helps create steroid hormones. Emily Scott's lab is studying how CYP17A1 could be selectively inhibited to treat prostate cancer. She and graduate student Natasha DeVore elucidated the structure shown using X-ray crystallography. Dr. Scott created the image (both white bg and transparent bg) for the NIGMS image gallery. See the "Medium-Resolution Image" for a PNG version of the image that is transparent.
Emily Scott, University of Kansas
View Media
2528: A drug's life in the body (with labels)
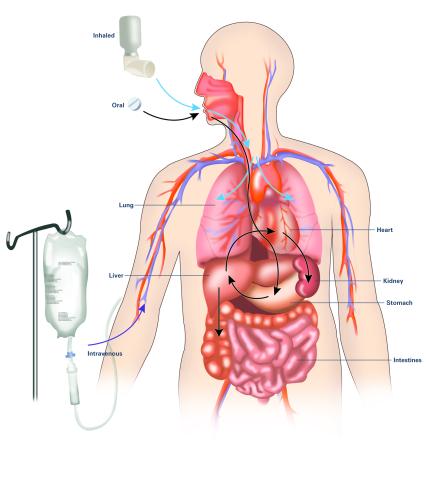
2528: A drug's life in the body (with labels)
A drug's life in the body. Medicines taken by mouth (oral) pass through the liver before they are absorbed into the bloodstream. Other forms of drug administration bypass the liver, entering the blood directly. See 2527 for an unlabeled version of this illustration. Featured in Medicines By Design.
Crabtree + Company
View Media
2526: Activation energy (with labels)
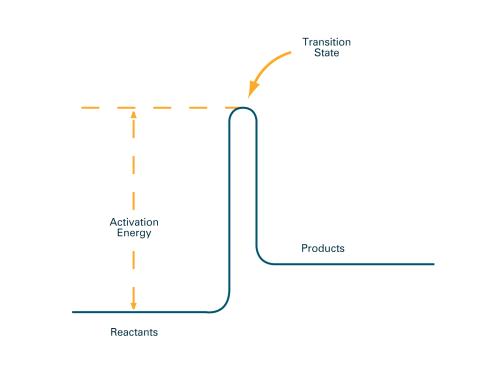
2526: Activation energy (with labels)
To become products, reactants must overcome an energy hill. See image 2525 for an unlabeled version of this illustration. Featured in The Chemistry of Health.
Crabtree + Company
View Media
