Switch to List View
Image and Video Gallery
This is a searchable collection of scientific photos, illustrations, and videos. The images and videos in this gallery are licensed under Creative Commons Attribution Non-Commercial ShareAlike 3.0. This license lets you remix, tweak, and build upon this work non-commercially, as long as you credit and license your new creations under identical terms.
1332: Mitosis - telophase
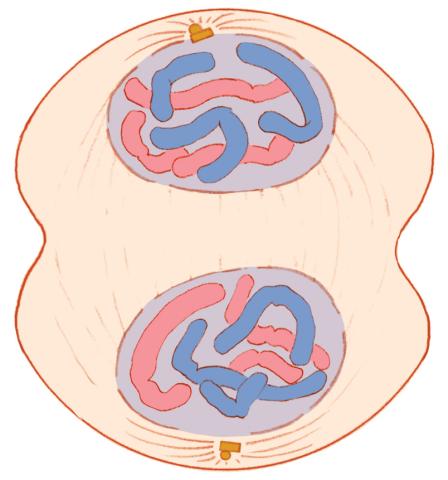
1332: Mitosis - telophase
Telophase during mitosis: Nuclear membranes form around each of the two sets of chromosomes, the chromosomes begin to spread out, and the spindle begins to break down. Mitosis is responsible for growth and development, as well as for replacing injured or worn out cells throughout the body. For simplicity, mitosis is illustrated here with only six chromosomes.
Judith Stoffer
View Media
2569: Circadian rhythm (with labels)
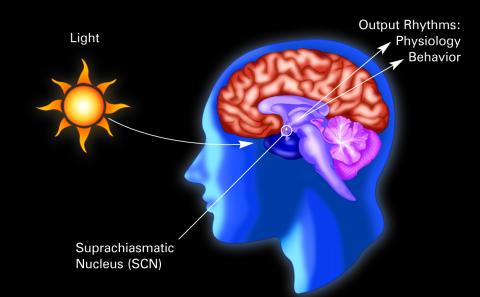
2569: Circadian rhythm (with labels)
The human body keeps time with a master clock called the suprachiasmatic nucleus or SCN. Situated inside the brain, it's a tiny sliver of tissue about the size of a grain of rice, located behind the eyes. It sits quite close to the optic nerve, which controls vision, and this means that the SCN "clock" can keep track of day and night. The SCN helps control sleep and maintains our circadian rhythm--the regular, 24-hour (or so) cycle of ups and downs in our bodily processes such as hormone levels, blood pressure, and sleepiness. The SCN regulates our circadian rhythm by coordinating the actions of billions of miniature "clocks" throughout the body. These aren't actually clocks, but rather are ensembles of genes inside clusters of cells that switch on and off in a regular, 24-hour (or so) cycle in our physiological day.
Crabtree + Company
View Media

7012: Adult Hawaiian bobtail squid burying in the sand
7012: Adult Hawaiian bobtail squid burying in the sand
Each morning, the nocturnal Hawaiian bobtail squid, Euprymna scolopes, hides from predators by digging into the sand. At dusk, it leaves the sand again to hunt.
Related to image 7010 and 7011.
Related to image 7010 and 7011.
Margaret J. McFall-Ngai, Carnegie Institution for Science/California Institute of Technology, and Edward G. Ruby, California Institute of Technology.
View Media
1017: Lily mitosis 07
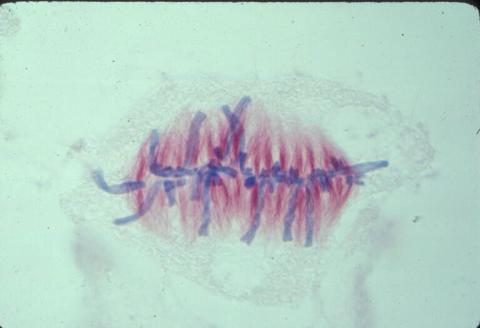
1017: Lily mitosis 07
A light microscope image of a cell from the endosperm of an African globe lily (Scadoxus katherinae). This is one frame of a time-lapse sequence that shows cell division in action. The lily is considered a good organism for studying cell division because its chromosomes are much thicker and easier to see than human ones. Staining shows microtubules in red and chromosomes in blue. Here, condensed chromosomes are clearly visible and have lined up in the middle of the dividing cell.
Related to images 1010, 1011, 1012, 1013, 1014, 1015, 1016, 1018, 1019, and 1021.
Related to images 1010, 1011, 1012, 1013, 1014, 1015, 1016, 1018, 1019, and 1021.
Andrew S. Bajer, University of Oregon, Eugene
View Media
3419: X-ray co-crystal structure of Src kinase bound to a DNA-templated macrocycle inhibitor 7
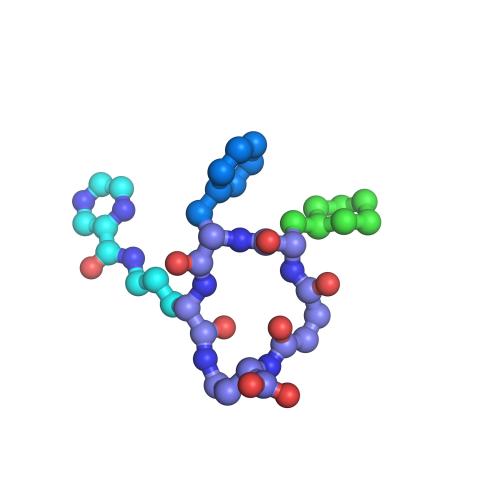
3419: X-ray co-crystal structure of Src kinase bound to a DNA-templated macrocycle inhibitor 7
X-ray co-crystal structure of Src kinase bound to a DNA-templated macrocycle inhibitor. Related to images 3413, 3414, 3415, 3416, 3417, and 3418.
Markus A. Seeliger, Stony Brook University Medical School and David R. Liu, Harvard University
View Media
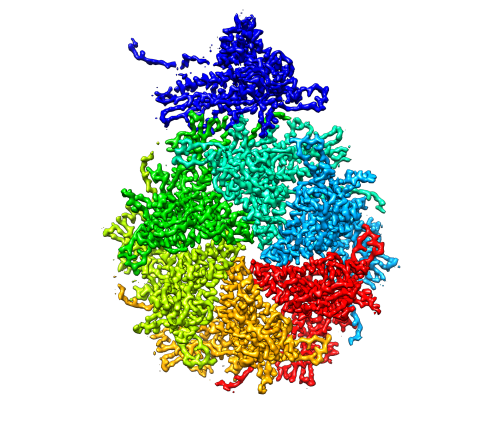
5875: Bacteriophage P22 capsid, detail
5875: Bacteriophage P22 capsid, detail
Detail of a subunit of the capsid, or outer cover, of bacteriophage P22, a virus that infects the Salmonella bacteria. Cryo-electron microscopy (cryo-EM) was used to capture details of the capsid proteins, each shown here in a separate color. Thousands of cryo-EM scans capture the structure and shape of all the individual proteins in the capsid and their position relative to other proteins. A computer model combines these scans into the image shown here. Related to image 5874.
Dr. Wah Chiu, Baylor College of Medicine
View Media
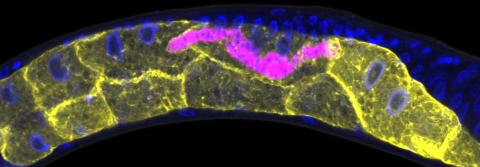
5777: Microsporidia in roundworm 1
5777: Microsporidia in roundworm 1
Many disease-causing microbes manipulate their host’s metabolism and cells for their own ends. Microsporidia—which are parasites closely related to fungi—infect and multiply inside animal cells, and take the rearranging of cells’ interiors to a new level. They reprogram animal cells such that the cells start to fuse, causing them to form long, continuous tubes. As shown in this image of the roundworm Caenorhabditis elegans, microsporidia (shown in magenta) have invaded the worm’s gut cells (shown in yellow; the cells’ nuclei are shown in blue) and have instructed the cells to merge. The cell fusion enables the microsporidia to thrive and propagate in the expanded space. Scientists study microsporidia in worms to gain more insight into how these parasites manipulate their host cells. This knowledge might help researchers devise strategies to prevent or treat infections with microsporidia. For more on the research into microsporidia, see this news release from the University of California San Diego. Related to images 5778 and 5779.
Keir Balla and Emily Troemel, University of California San Diego
View Media
6850: Himastatin and bacteria
6850: Himastatin and bacteria
A model of the molecule himastatin overlaid on an image of Bacillus subtilis bacteria. Scientists first isolated himastatin from the bacterium Streptomyces himastatinicus, and the molecule shows antibiotic activity. The researchers who created this image developed a new, more concise way to synthesize himastatin so it can be studied more easily. They also tested the effects of himastatin and derivatives of the molecule on B. subtilis.
More information about the research that produced this image can be found in the Science paper “Total synthesis of himastatin” by D’Angelo et al.
Related to image 6848 and video 6851.
More information about the research that produced this image can be found in the Science paper “Total synthesis of himastatin” by D’Angelo et al.
Related to image 6848 and video 6851.
Mohammad Movassaghi, Massachusetts Institute of Technology.
View Media
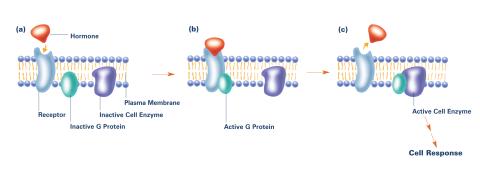
2538: G switch (with labels and stages)
2538: G switch (with labels and stages)
The G switch allows our bodies to respond rapidly to hormones. G proteins act like relay batons to pass messages from circulating hormones into cells. A hormone (red) encounters a receptor (blue) in the membrane of a cell. Next, a G protein (green) becomes activated and makes contact with the receptor to which the hormone is attached. Finally, the G protein passes the hormone's message to the cell by switching on a cell enzyme (purple) that triggers a response. See image 2536 and 2537 for other versions of this image. Featured in Medicines By Design.
Crabtree + Company
View Media
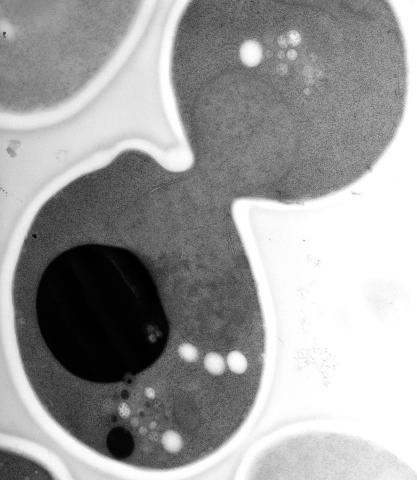
5770: EM of yeast cell division
5770: EM of yeast cell division
Cell division is an incredibly coordinated process. It not only ensures that the new cells formed during this event have a full set of chromosomes, but also that they are endowed with all the cellular materials, including proteins, lipids and small functional compartments called organelles, that are required for normal cell activity. This proper apportioning of essential cell ingredients helps each cell get off to a running start.
This image shows an electron microscopy (EM) thin section taken at 10,000x magnification of a dividing yeast cell over-expressing the protein ubiquitin, which is involved in protein degradation and recycling. The picture features mother and daughter endosome accumulations (small organelles with internal vesicles), a darkly stained vacuole and a dividing nucleus in close contact with a cadre of lipid droplets (unstained spherical bodies). Other dynamic events are also visible, such as spindle microtubules in the nucleus and endocytic pits at the plasma membrane.
These extensive details were revealed thanks to a preservation method involving high-pressure freezing, freeze-substitution and Lowicryl HM20 embedding.
This image shows an electron microscopy (EM) thin section taken at 10,000x magnification of a dividing yeast cell over-expressing the protein ubiquitin, which is involved in protein degradation and recycling. The picture features mother and daughter endosome accumulations (small organelles with internal vesicles), a darkly stained vacuole and a dividing nucleus in close contact with a cadre of lipid droplets (unstained spherical bodies). Other dynamic events are also visible, such as spindle microtubules in the nucleus and endocytic pits at the plasma membrane.
These extensive details were revealed thanks to a preservation method involving high-pressure freezing, freeze-substitution and Lowicryl HM20 embedding.
Matthew West and Greg Odorizzi, University of Colorado
View Media

5758: Migrating pigment cells
5758: Migrating pigment cells
Pigment cells are cells that give skin its color. In fishes and amphibians, like frogs and salamanders, pigment cells are responsible for the characteristic skin patterns that help these organisms to blend into their surroundings or attract mates. The pigment cells are derived from neural crest cells, which are cells originating from the neural tube in the early embryo. This image shows neural crest cell-derived, migrating pigment cells in a salamander. Investigating pigment cell formation and migration in animals helps answer important fundamental questions about the factors that control pigmentation in the skin of animals, including humans. Related to images 5754, 5755, 5756 and 5757.
David Parichy, University of Washington
View Media

3344: Artificial cilia exhibit spontaneous beating
3344: Artificial cilia exhibit spontaneous beating
Researchers have created artificial cilia that wave like the real thing. Zvonimir Dogic and his Brandeis University colleagues combined just a few cilia proteins to create cilia that are able to wave and sweep material around--although more slowly and simply than real ones. The researchers are using the lab-made cilia to study how the structures coordinate their movements and what happens when they don't move properly. Featured in the August 18, 2011, issue of Biomedical Beat.
Zvonimir Dogic
View Media

2728: Sponge
2728: Sponge
Many of today's medicines come from products found in nature, such as this sponge found off the coast of Palau in the Pacific Ocean. Chemists have synthesized a compound called Palau'amine, which appears to act against cancer, bacteria and fungi. In doing so, they invented a new chemical technique that will empower the synthesis of other challenging molecules.
Phil Baran, Scripps Research Institute
View Media
6776: Tracking cells in a gastrulating zebrafish embryo
6776: Tracking cells in a gastrulating zebrafish embryo
During development, a zebrafish embryo is transformed from a ball of cells into a recognizable body plan by sweeping convergence and extension cell movements. This process is called gastrulation. Each line in this video represents the movement of a single zebrafish embryo cell over the course of 3 hours. The video was created using time-lapse confocal microscopy. Related to image 6775.
Liliana Solnica-Krezel, Washington University School of Medicine in St. Louis.
View Media

6902: Arachnoidiscus diatom
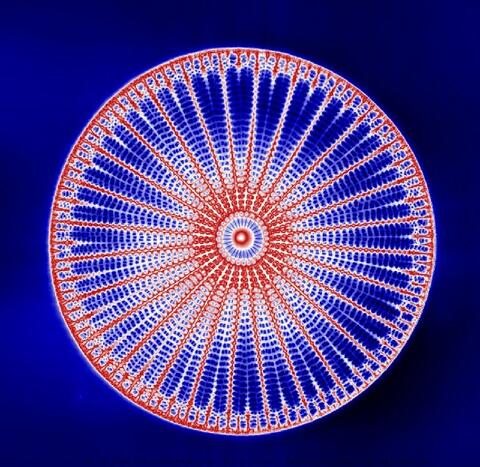
6902: Arachnoidiscus diatom
An Arachnoidiscus diatom with a diameter of 190µm. Diatoms are microscopic algae that have cell walls made of silica, which is the strongest known biological material relative to its density. In Arachnoidiscus, the cell wall is a radially symmetric pillbox-like shell composed of overlapping halves that contain intricate and delicate patterns. Sometimes, Arachnoidiscus is called “a wheel of glass.”
This image was taken with the orientation-independent differential interference contrast microscope.
This image was taken with the orientation-independent differential interference contrast microscope.
Michael Shribak, Marine Biological Laboratory/University of Chicago.
View Media

2371: NMR spectrometer
2371: NMR spectrometer
This photo shows a Varian Unity Inova 900 MHz, 21.1 T standard bore magnet Nuclear Magnetic Resonnance (NMR) spectrometer. NMR spectroscopy provides data used to determine the structures of proteins in solution, rather than in crystal form, as in X-ray crystallography. The technique is limited to smaller proteins or protein fragments in a high throughput approach.
Center for Eukaryotic Structural Genomics
View Media
3451: Proteasome
3451: Proteasome
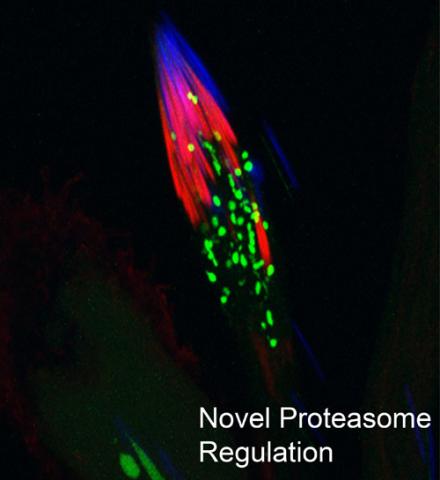
This fruit fly spermatid recycles various molecules, including malformed or damaged proteins. Actin filaments (red) in the cell draw unwanted proteins toward a barrel-shaped structure called the proteasome (green clusters), which degrades the molecules into their basic parts for re-use.
Sigi Benjamin-Hong, Rockefeller University
View Media
3332: Polarized cells- 01
3332: Polarized cells- 01
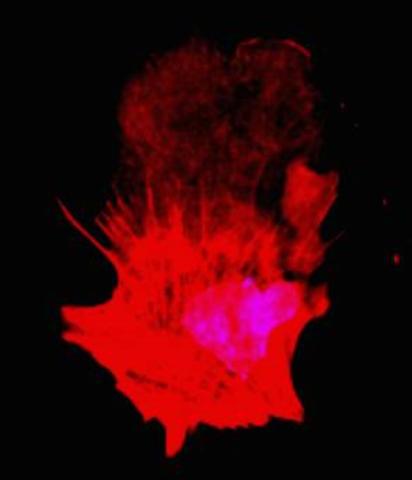
Cells move forward with lamellipodia and filopodia supported by networks and bundles of actin filaments. Proper, controlled cell movement is a complex process. Recent research has shown that an actin-polymerizing factor called the Arp2/3 complex is the key component of the actin polymerization engine that drives amoeboid cell motility. ARPC3, a component of the Arp2/3 complex, plays a critical role in actin nucleation. In this photo, the ARPC3+/+ fibroblast cells were fixed and stained with Alexa 546 phalloidin for F-actin (red) and DAPI to visualize the nucleus (blue). ARPC3+/+ fibroblast cells with lamellipodia leading edge. Related to images 3328, 3329, 3330, 3331, and 3333.
Rong Li and Praveen Suraneni, Stowers Institute for Medical Research
View Media

3658: Electrostatic map of human spermine synthase
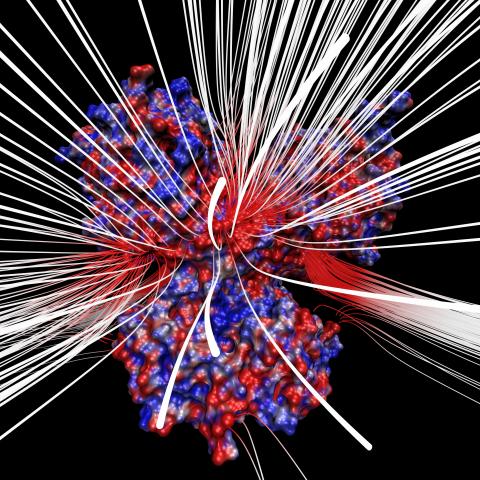
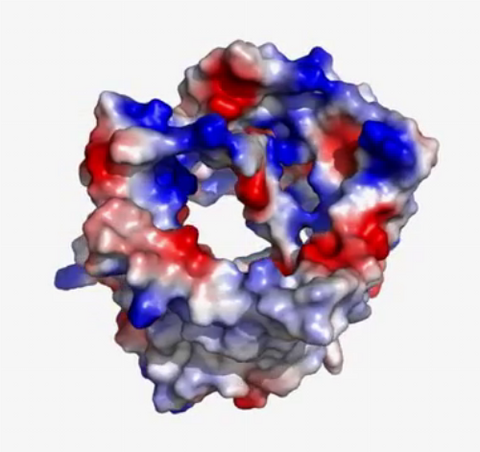
3658: Electrostatic map of human spermine synthase
From PDB entry 3c6k, Crystal structure of human spermine synthase in complex with spermidine and 5-methylthioadenosine.
Emil Alexov, Clemson University
View Media
7015: Bacterial cells migrating through the tissues of the squid light organ
7015: Bacterial cells migrating through the tissues of the squid light organ
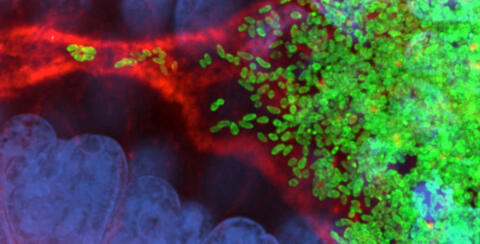
Vibrio fischeri cells (~ 2 mm), labeled with green fluorescent protein (GFP), passing through a very narrow bottleneck in the tissues (red) of the Hawaiian bobtail squid, Euprymna scolopes, on the way to the crypts where the symbiont population resides. This image was taken using a confocal fluorescence microscope.
Margaret J. McFall-Ngai, Carnegie Institution for Science/California Institute of Technology, and Edward G. Ruby, California Institute of Technology.
View Media
5887: Plasma-Derived Membrane Vesicles
5887: Plasma-Derived Membrane Vesicles
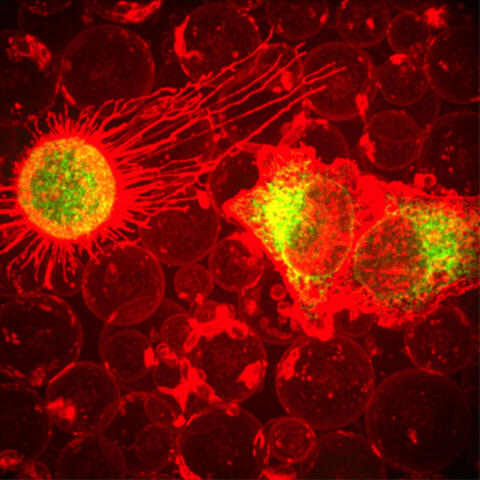
This fiery image doesn’t come from inside a bubbling volcano. Instead, it shows animal cells caught in the act of making bubbles, or blebbing. Some cells regularly pinch off parts of their membranes to produce bubbles filled with a mix of proteins and fats. The bubbles (red) are called plasma-derived membrane vesicles, or PMVs, and can travel to other parts of the body where they may aid in cell-cell communication. The University of Texas, Austin, researchers responsible for this photo are exploring ways to use PMVs to deliver medicines to precise locations in the body.
This image, entered in the Biophysical Society’s 2017 Art of Science Image contest, used two-channel spinning disk confocal fluorescence microscopy. It was also featured in the NIH Director’s Blog in May 2017.
This image, entered in the Biophysical Society’s 2017 Art of Science Image contest, used two-channel spinning disk confocal fluorescence microscopy. It was also featured in the NIH Director’s Blog in May 2017.
Jeanne Stachowiak, University of Texas at Austin
View Media
3620: Anglerfish ovary cross-section
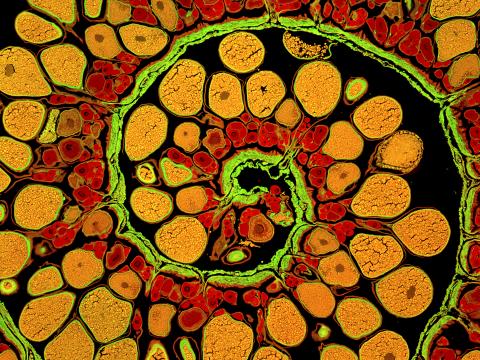
3620: Anglerfish ovary cross-section
This image captures the spiral-shaped ovary of an anglerfish in cross-section. Once matured, these eggs will be released in a gelatinous, floating mass. For some species of anglerfish, this egg mass can be up to 3 feet long and include nearly 200,000 eggs.
This image was part of the Life: Magnified exhibit that ran from June 3, 2014, to January 21, 2015, at Dulles International Airport.
This image was part of the Life: Magnified exhibit that ran from June 3, 2014, to January 21, 2015, at Dulles International Airport.
James E. Hayden, The Wistar Institute, Philadelphia, Pa.
View Media
5751: Genetically identical mycobacteria respond differently to antibiotic 1
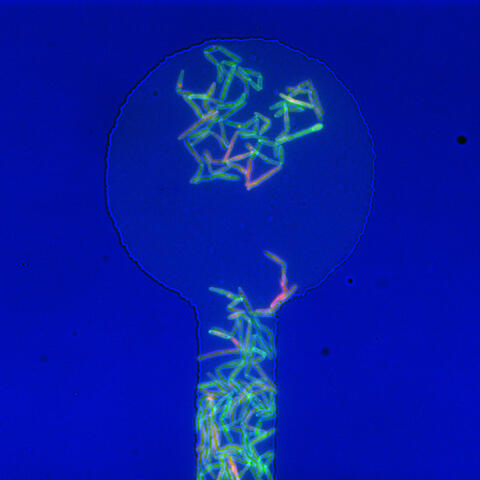
5751: Genetically identical mycobacteria respond differently to antibiotic 1
Antibiotic resistance in microbes is a serious health concern. So researchers have turned their attention to how bacteria undo the action of some antibiotics. Here, scientists set out to find the conditions that help individual bacterial cells survive in the presence of the antibiotic rifampicin. The research team used Mycobacterium smegmatis, a more harmless relative of Mycobacterium tuberculosis, which infects the lung and other organs and causes serious disease.
In this image, genetically identical mycobacteria are growing in a miniature growth chamber called a microfluidic chamber. Using live imaging, the researchers found that individual mycobacteria will respond differently to the antibiotic, depending on the growth stage and other timing factors. The researchers used genetic tagging with green fluorescent protein to distinguish cells that can resist rifampicin and those that cannot. With this gene tag, cells tolerant of the antibiotic light up in green and those that are susceptible in violet, enabling the team to monitor the cells' responses in real time.
To learn more about how the researchers studied antibiotic resistance in mycobacteria, see this news release from Tufts University. Related to video 5752.
In this image, genetically identical mycobacteria are growing in a miniature growth chamber called a microfluidic chamber. Using live imaging, the researchers found that individual mycobacteria will respond differently to the antibiotic, depending on the growth stage and other timing factors. The researchers used genetic tagging with green fluorescent protein to distinguish cells that can resist rifampicin and those that cannot. With this gene tag, cells tolerant of the antibiotic light up in green and those that are susceptible in violet, enabling the team to monitor the cells' responses in real time.
To learn more about how the researchers studied antibiotic resistance in mycobacteria, see this news release from Tufts University. Related to video 5752.
Bree Aldridge, Tufts University
View Media
3571: HIV-1 virus in the colon
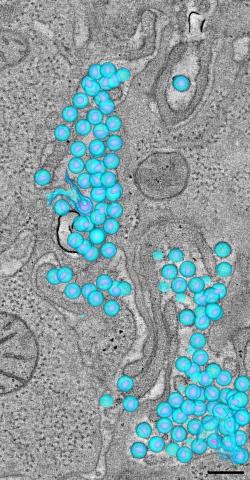
3571: HIV-1 virus in the colon
A tomographic reconstruction of the colon shows the location of large pools of HIV-1 virus particles (in blue) located in the spaces between adjacent cells. The purple objects within each sphere represent the conical cores that are one of the structural hallmarks of the HIV virus.
Mark Ladinsky, California Institute of Technology
View Media

2735: Network Map
2735: Network Map
This network map shows the overlap (green) between the long QT syndrome (yellow) and epilepsy (blue) protein-interaction neighborhoods located within the human interactome. Researchers have learned to integrate genetic, cellular and clinical information to find out why certain medicines can trigger fatal heart arrhythmias. Featured in Computing Life magazine.
Seth Berger, Mount Sinai School of Medicine
View Media
2456: Z rings in bacterial division
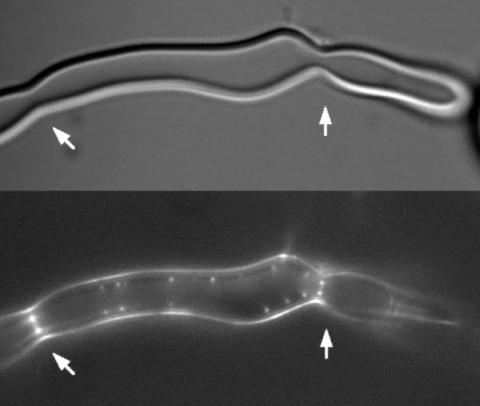
2456: Z rings in bacterial division
Lab-made liposomes contract where Z rings have gathered together and the constriction forces are greatest (arrows). The top picture shows a liposome, and the bottom picture shows fluorescence from Z rings (arrows) inside the same liposome simultaneously.
Masaki Osawa, Duke University
View Media
3255: Centromeres on human chromosomes
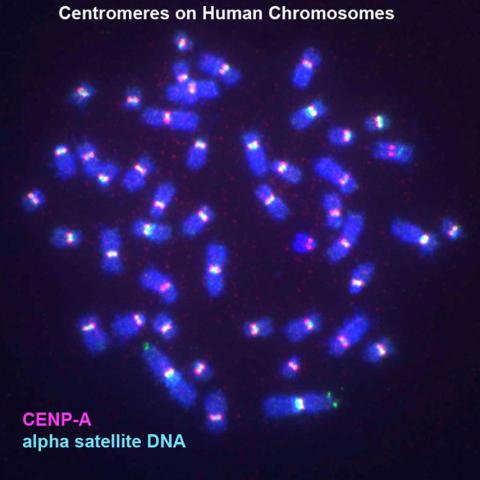
3255: Centromeres on human chromosomes
Human metaphase chromosomes are visible with fluorescence in vitro hybridization (FISH). Centromeric alpha satellite DNA (green) are found in the heterochromatin at each centromere. Immunofluorescence with CENP-A (red) shows the centromere-specific histone H3 variant that specifies the kinetochore.
Peter Warburton, Mount Sinai School of Medicine
View Media
6765: X-ray diffraction pattern from a crystallized cefotaxime-CCD-1 complex
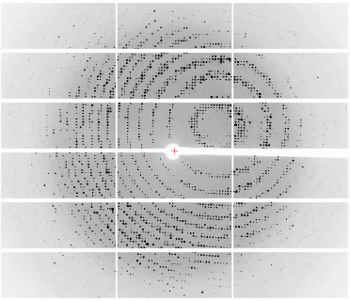
6765: X-ray diffraction pattern from a crystallized cefotaxime-CCD-1 complex
CCD-1 is an enzyme produced by the bacterium Clostridioides difficile that helps it resist antibiotics. Researchers crystallized complexes where a CCD-1 molecule and a molecule of the antibiotic cefotaxime were bound together. Then, they shot X-rays at the complexes to determine their structure—a process known as X-ray crystallography. This image shows the X-ray diffraction pattern of a complex.
Related to images 6764, 6766, and 6767.
Related to images 6764, 6766, and 6767.
Keith Hodgson, Stanford University.
View Media
3584: Rotavirus structure
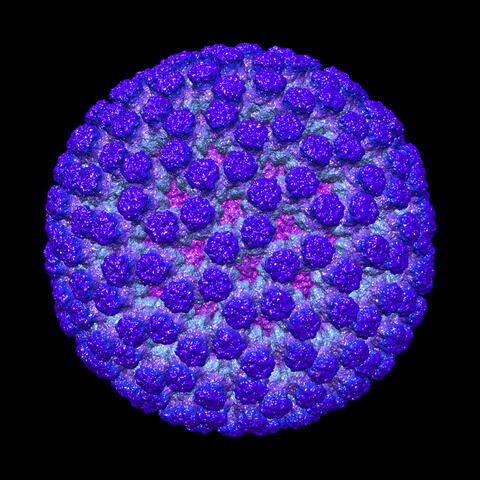
3584: Rotavirus structure
This image shows a computer-generated, three-dimensional map of the rotavirus structure. This virus infects humans and other animals and causes severe diarrhea in infants and young children. By the age of five, almost every child in the world has been infected with this virus at least once. Scientists have found a vaccine against rotavirus, so in the United States there are very few fatalities, but in developing countries and in places where the vaccine is unavailable, this virus is responsible for more than 200,000 deaths each year.
The rotavirus comprises three layers: the outer, middle and inner layers. On infection, the outer layer is removed, leaving behind a "double-layered particle." Researchers have studied the structure of this double-layered particle with a transmission electron microscope. Many images of the virus at a magnification of ~50,000x were acquired, and computational analysis was used to combine the individual particle images into a three-dimensional reconstruction.
The image was rendered by Melody Campbell (PhD student at TSRI). Work that led to the 3D map was published in Campbell et al. Movies of ice-embedded particles enhance resolution in electron cryo-microscopy. Structure. 2012;20(11):1823-8. PMCID: PMC3510009.
This image was part of the Life: Magnified exhibit that ran from June 3, 2014, to January 21, 2015, at Dulles International Airport.
The rotavirus comprises three layers: the outer, middle and inner layers. On infection, the outer layer is removed, leaving behind a "double-layered particle." Researchers have studied the structure of this double-layered particle with a transmission electron microscope. Many images of the virus at a magnification of ~50,000x were acquired, and computational analysis was used to combine the individual particle images into a three-dimensional reconstruction.
The image was rendered by Melody Campbell (PhD student at TSRI). Work that led to the 3D map was published in Campbell et al. Movies of ice-embedded particles enhance resolution in electron cryo-microscopy. Structure. 2012;20(11):1823-8. PMCID: PMC3510009.
This image was part of the Life: Magnified exhibit that ran from June 3, 2014, to January 21, 2015, at Dulles International Airport.
Bridget Carragher, The Scripps Research Institute, La Jolla, CA
View Media
2430: Fruit fly retina 01
2430: Fruit fly retina 01
Image showing rhabdomeres (red), the light-sensitive structures in the fruit fly retina, and rhodopsin-4 (blue), a light-sensing molecule.
Hermann Steller, Rockefeller University
View Media
5883: Beta-galactosidase montage showing cryo-EM improvement--gradient background
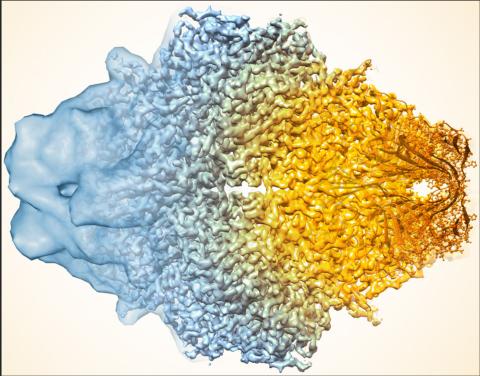
5883: Beta-galactosidase montage showing cryo-EM improvement--gradient background
Composite image of beta-galactosidase showing how cryo-EM’s resolution has improved dramatically in recent years. Older images to the left, more recent to the right. Related to image 5882. NIH Director Francis Collins featured this on his blog on January 14, 2016.
Veronica Falconieri, Sriram Subramaniam Lab, National Cancer Institute
View Media
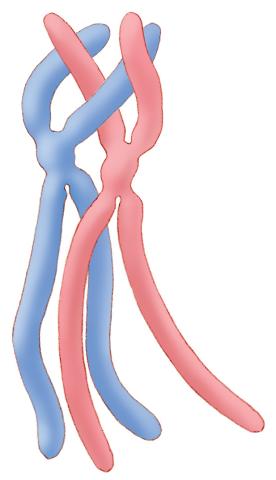
1315: Chromosomes before crossing over
1315: Chromosomes before crossing over
Duplicated pair of chromosomes lined up and ready to cross over.
Judith Stoffer
View Media
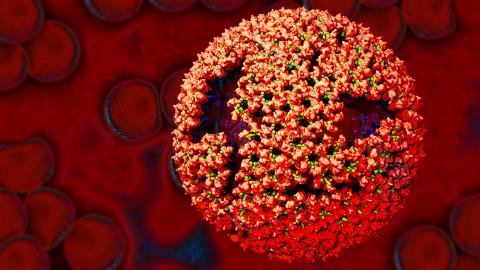
3771: Molecular model of freshly made Rous sarcoma virus (RSV)
3771: Molecular model of freshly made Rous sarcoma virus (RSV)
Viruses have been the foes of animals and other organisms for time immemorial. For almost as long, they've stayed well hidden from view because they are so tiny (they aren't even cells, so scientists call the individual virus a "particle"). This image shows a molecular model of a particle of the Rous sarcoma virus (RSV), a virus that infects and sometimes causes cancer in chickens. In the background is a photo of red blood cells. The particle shown is "immature" (not yet capable of infecting new cells) because it has just budded from an infected chicken cell and entered the bird's bloodstream. The outer shell of the immature virus is made up of a regular assembly of large proteins (shown in red) that are linked together with short protein molecules called peptides (green). This outer shell covers and protects the proteins (blue) that form the inner shell of the particle. But as you can see, the protective armor of the immature virus contains gaping holes. As the particle matures, the short peptides are removed and the large proteins rearrange, fusing together into a solid sphere capable of infecting new cells. While still immature, the particle is vulnerable to drugs that block its development. Knowing the structure of the immature particle may help scientists develop better medications against RSV and similar viruses in humans. Scientists used sophisticated computational tools to reconstruct the RSV atomic structure by crunching various data on the RSV proteins to simulate the entire structure of immature RSV.
Boon Chong Goh, University of Illinois at Urbana-Champaign
View Media
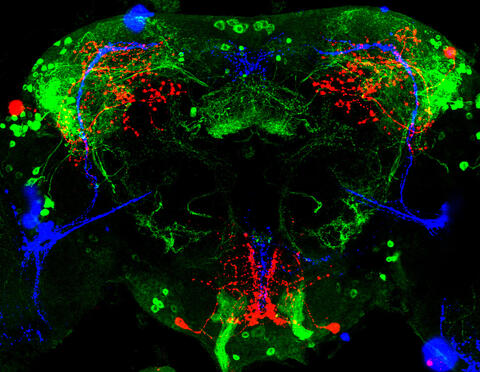
3754: Circadian rhythm neurons in the fruit fly brain
3754: Circadian rhythm neurons in the fruit fly brain
Some nerve cells (neurons) in the brain keep track of the daily cycle. This time-keeping mechanism, called the circadian clock, is found in all animals including us. The circadian clock controls our daily activities such as sleep and wakefulness. Researchers are interested in finding the neuron circuits involved in this time keeping and how the information about daily time in the brain is relayed to the rest of the body. In this image of a brain of the fruit fly Drosophila the time-of-day information flowing through the brain has been visualized by staining the neurons involved: clock neurons (shown in blue) function as "pacemakers" by communicating with neurons that produce a short protein called leucokinin (LK) (red), which, in turn, relays the time signal to other neurons, called LK-R neurons (green). This signaling cascade set in motion by the pacemaker neurons helps synchronize the fly's daily activity with the 24-hour cycle. To learn more about what scientists have found out about circadian pacemaker neurons in the fruit fly see this news release by New York University. This work was featured in the Biomedical Beat blog post Cool Image: A Circadian Circuit.
Justin Blau, New York University
View Media
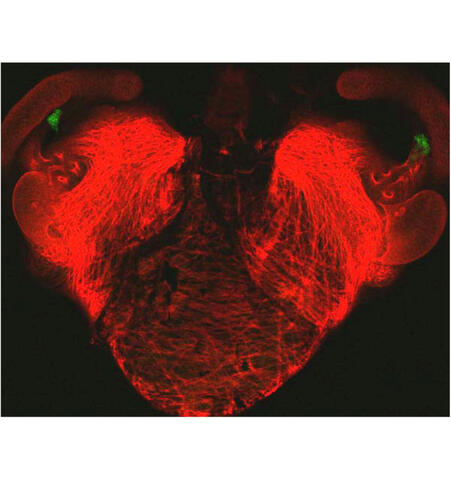
7018: Bacterial cells aggregating above the light organ of the Hawaiian bobtail squid
7018: Bacterial cells aggregating above the light organ of the Hawaiian bobtail squid
A light organ (~0.5 mm across) of a juvenile Hawaiian bobtail squid, Euprymna scolopes. Movement of cilia on the surface of the organ aggregates bacterial symbionts (green) into two areas above sets of pores that lead to interior crypts. This image was taken using a confocal fluorescence microscope.
Related to images 7016, 7017, 7019, and 7020.
Related to images 7016, 7017, 7019, and 7020.
Margaret J. McFall-Ngai, Carnegie Institution for Science/California Institute of Technology, and Edward G. Ruby, California Institute of Technology.
View Media

3475: Automated Worm Sorter - 4
3475: Automated Worm Sorter - 4
Georgia Tech associate professor Hang Lu holds a microfluidic chip that is part of a system that uses artificial intelligence and cutting-edge image processing to automatically examine large number of nematodes used for genetic research.
Georgia Tech/Gary Meek
View Media

2410: DNase
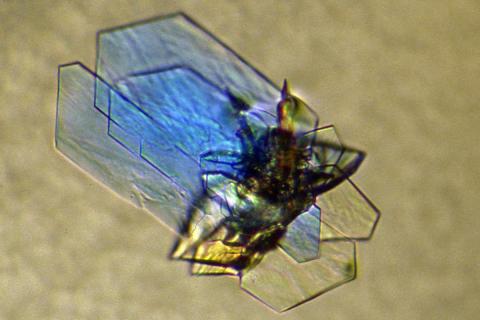
2410: DNase
Crystals of DNase protein created for X-ray crystallography, which can reveal detailed, three-dimensional protein structures.
Alex McPherson, University of California, Irvine
View Media
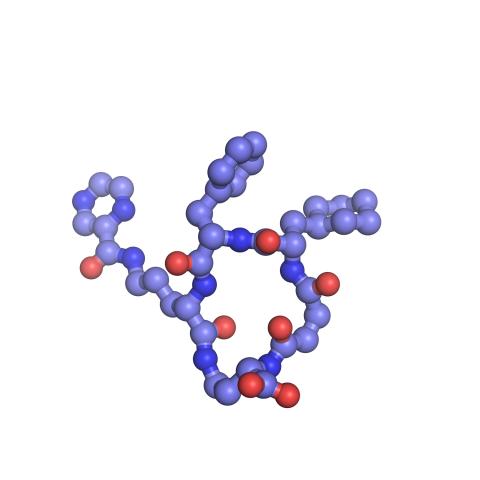
3418: X-ray co-crystal structure of Src kinase bound to a DNA-templated macrocycle inhibitor 6
3418: X-ray co-crystal structure of Src kinase bound to a DNA-templated macrocycle inhibitor 6
X-ray co-crystal structure of Src kinase bound to a DNA-templated macrocycle inhibitor. Related to images 3413, 3414, 3415, 3416, 3417, and 3419.
Markus A. Seeliger, Stony Brook University Medical School and David R. Liu, Harvard University
View Media
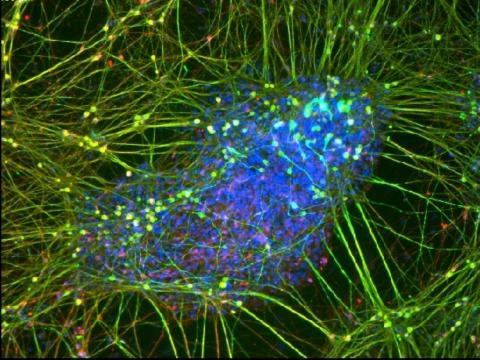
3264: Peripheral nerve cell derived from ES cells
3264: Peripheral nerve cell derived from ES cells
A peripheral nerve cell made from human embryonic stem cell-derived neural crest stem cells. The nucleus is shown in blue, and nerve cell proteins peripherin and beta-tubulin (Tuj1) are shown in green and red, respectively. Related to image 3263.
Stephen Dalton, University of Georgia
View Media
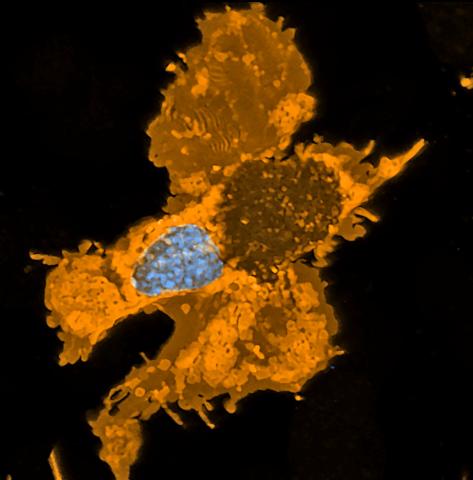
5754: Zebrafish pigment cell
5754: Zebrafish pigment cell
Pigment cells are cells that give skin its color. In fishes and amphibians, like frogs and salamanders, pigment cells are responsible for the characteristic skin patterns that help these organisms to blend into their surroundings or attract mates. The pigment cells are derived from neural crest cells, which are cells originating from the neural tube in the early embryo. Investigating pigment cell formation and migration in animals helps answer important fundamental questions about the factors that control pigmentation in the skin of animals, including humans. This image shows a pigment cell from zebrafish at high resolution. Related to images 5755, 5756, 5757 and 5758.
David Parichy, University of Washington
View Media
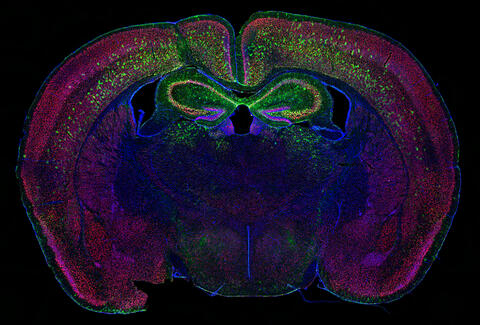
6780: Calling Cards in a mouse brain
6780: Calling Cards in a mouse brain
The green spots in this mouse brain are cells labeled with Calling Cards, a technology that records molecular events in brain cells as they mature. Understanding these processes during healthy development can guide further research into what goes wrong in cases of neuropsychiatric disorders. Also fluorescently labeled in this image are neurons (red) and nuclei (blue). Calling Cards and its application are described in the Cell paper “Self-Reporting Transposons Enable Simultaneous Readout of Gene Expression and Transcription Factor Binding in Single Cells” by Moudgil et al.; and the Proceedings of the National Academy of Sciences paper “A viral toolkit for recording transcription factor–DNA interactions in live mouse tissues” by Cammack et al. The technology was also featured in the NIH Director’s Blog post The Amazing Brain: Tracking Molecular Events with Calling Cards.
Related to video
Related to video
Allen Yen, Lab of Joseph Dougherty, Washington University School of Medicine in St. Louis.
View Media
3664: Mitochondria from rat heart muscle cell_2
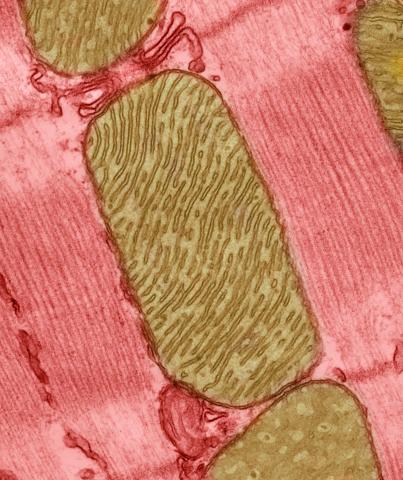
3664: Mitochondria from rat heart muscle cell_2
These mitochondria (brown) are from the heart muscle cell of a rat. Mitochondria have an inner membrane that folds in many places (and that appears here as striations). This folding vastly increases the surface area for energy production. Nearly all our cells have mitochondria. Related to image 3661.
National Center for Microscopy and Imaging Research
View Media
3720: Cas4 nuclease protein structure
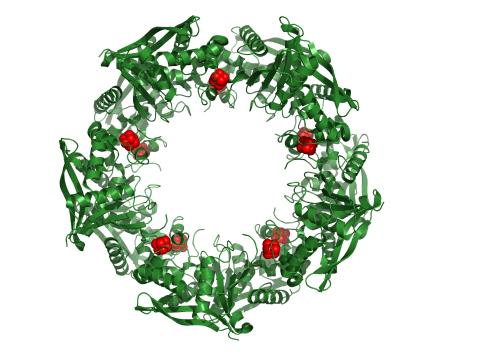
3720: Cas4 nuclease protein structure
This wreath represents the molecular structure of a protein, Cas4, which is part of a system, known as CRISPR, that bacteria use to protect themselves against viral invaders. The green ribbons show the protein's structure, and the red balls show the location of iron and sulfur molecules important for the protein's function. Scientists harnessed Cas9, a different protein in the bacterial CRISPR system, to create a gene-editing tool known as CRISPR-Cas9. Using this tool, researchers are able to study a range of cellular processes and human diseases more easily, cheaply and precisely. In December, 2015, Science magazine recognized the CRISPR-Cas9 gene-editing tool as the "breakthrough of the year." Read more about Cas4 in the December 2015 Biomedical Beat post A Holiday-Themed Image Collection.
Fred Dyda, NIDDK
View Media
2841: Circadian rhythm
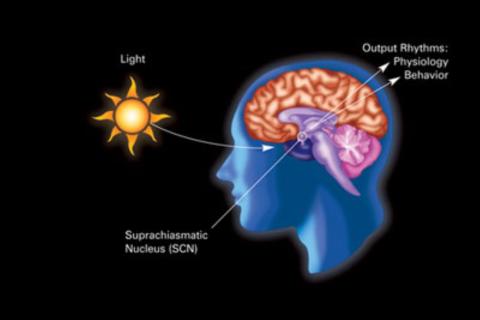
2841: Circadian rhythm
The human body keeps time with a master clock called the suprachiasmatic nucleus or SCN. Situated inside the brain, it's a tiny sliver of tissue about the size of a grain of rice, located behind the eyes. It sits quite close to the optic nerve, which controls vision, and this means that the SCN "clock" can keep track of day and night. The SCN helps control sleep by coordinating the actions of billions of miniature "clocks" throughout the body. These aren't actually clocks, but rather are ensembles of genes inside clusters of cells that switch on and off in a regular, 24-hour cycle in our physiological day.
Crabtree + Company
View Media
6750: C. elegans with blue and yellow lights in the background
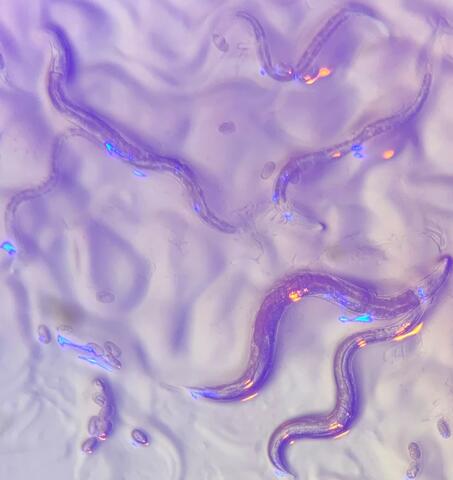
6750: C. elegans with blue and yellow lights in the background
These microscopic roundworms, called Caenorhabditis elegans, lack eyes and the opsin proteins used by visual systems to detect colors. However, researchers found that the worms can still sense the color of light in a way that enables them to avoid pigmented toxins made by bacteria. This image was captured using a stereo microscope.
H. Robert Horvitz and Dipon Ghosh, Massachusetts Institute of Technology.
View Media
2571: VDAC video 02
2571: VDAC video 02
This video shows the structure of the pore-forming protein VDAC-1 from humans. This molecule mediates the flow of products needed for metabolism--in particular the export of ATP--across the outer membrane of mitochondria, the power plants for eukaryotic cells. VDAC-1 is involved in metabolism and the self-destruction of cells--two biological processes central to health.
Related to videos 2570 and 2572.
Related to videos 2570 and 2572.
Gerhard Wagner, Harvard Medical School
View Media
6889: Lysosomes and microtubules
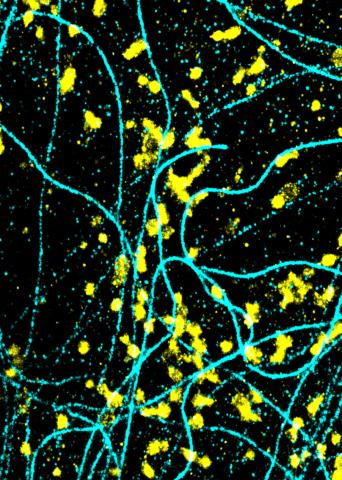
6889: Lysosomes and microtubules
Lysosomes (yellow) and detyrosinated microtubules (light blue). Lysosomes are bubblelike organelles that take in molecules and use enzymes to break them down. Microtubules are strong, hollow fibers that provide structural support to cells. The researchers who took this image found that in epithelial cells, detyrosinated microtubules are a small subset of fibers, and they concentrate lysosomes around themselves. This image was captured using Stochastic Optical Reconstruction Microscopy (STORM).
Related to images 6890, 6891, and 6892.
Related to images 6890, 6891, and 6892.
Melike Lakadamyali, Perelman School of Medicine at the University of Pennsylvania.
View Media
6998: Zika virus
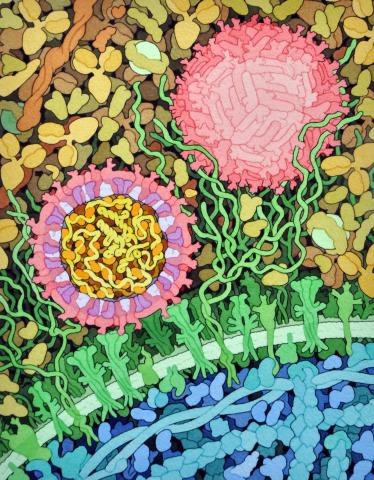
6998: Zika virus
Zika virus is shown in cross section at center left. On the outside, it includes envelope protein (red) and membrane protein (magenta) embedded in a lipid membrane (light purple). Inside, the RNA genome (yellow) is associated with capsid proteins (orange). The viruses are shown interacting with receptors on the cell surface (green) and are surrounded by blood plasma molecules at the top.
Amy Wu and Christine Zardecki, RCSB Protein Data Bank.
View Media
2739: Tetrapolar mitosis
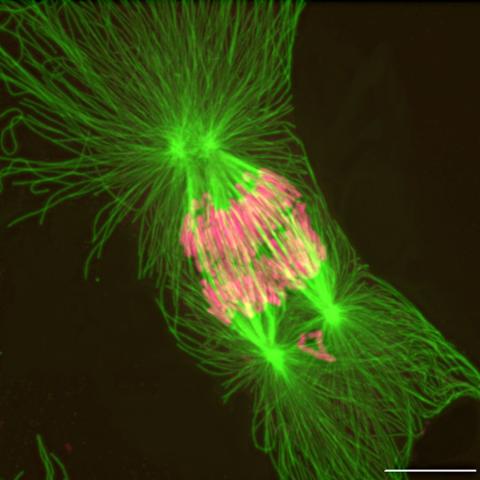
2739: Tetrapolar mitosis
This image shows an abnormal, tetrapolar mitosis. Chromosomes are highlighted pink. The cells shown are S3 tissue cultured cells from Xenopus laevis, African clawed frog.
Gary Gorbsky, Oklahoma Medical Research Foundation
View Media

