Switch to List View
Image and Video Gallery
This is a searchable collection of scientific photos, illustrations, and videos. The images and videos in this gallery are licensed under Creative Commons Attribution Non-Commercial ShareAlike 3.0. This license lets you remix, tweak, and build upon this work non-commercially, as long as you credit and license your new creations under identical terms.
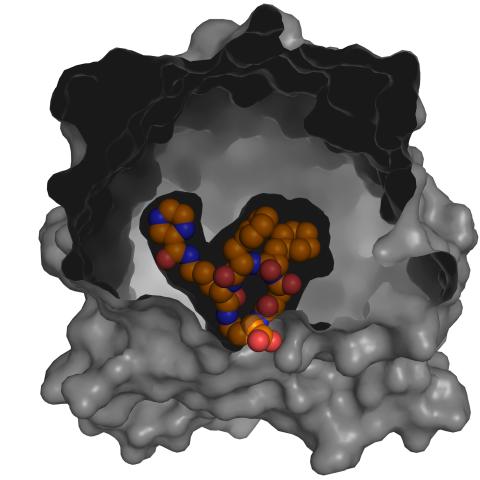
3417: X-ray co-crystal structure of Src kinase bound to a DNA-templated macrocycle inhibitor 5
3417: X-ray co-crystal structure of Src kinase bound to a DNA-templated macrocycle inhibitor 5
X-ray co-crystal structure of Src kinase bound to a DNA-templated macrocycle inhibitor. Related to images 3413, 3414, 3415, 3416, 3418, and 3419.
Markus A. Seeliger, Stony Brook University Medical School and David R. Liu, Harvard University
View Media
6850: Himastatin and bacteria
6850: Himastatin and bacteria
A model of the molecule himastatin overlaid on an image of Bacillus subtilis bacteria. Scientists first isolated himastatin from the bacterium Streptomyces himastatinicus, and the molecule shows antibiotic activity. The researchers who created this image developed a new, more concise way to synthesize himastatin so it can be studied more easily. They also tested the effects of himastatin and derivatives of the molecule on B. subtilis.
More information about the research that produced this image can be found in the Science paper “Total synthesis of himastatin” by D’Angelo et al.
Related to image 6848 and video 6851.
More information about the research that produced this image can be found in the Science paper “Total synthesis of himastatin” by D’Angelo et al.
Related to image 6848 and video 6851.
Mohammad Movassaghi, Massachusetts Institute of Technology.
View Media
2796: Anti-tumor drug ecteinascidin 743 (ET-743), structure without hydrogens 03
2796: Anti-tumor drug ecteinascidin 743 (ET-743), structure without hydrogens 03
Ecteinascidin 743 (ET-743, brand name Yondelis), was discovered and isolated from a sea squirt, Ecteinascidia turbinata, by NIGMS grantee Kenneth Rinehart at the University of Illinois. It was synthesized by NIGMS grantees E.J. Corey and later by Samuel Danishefsky. Multiple versions of this structure are available as entries 2790-2797.
Timothy Jamison, Massachusetts Institute of Technology
View Media
2526: Activation energy (with labels)
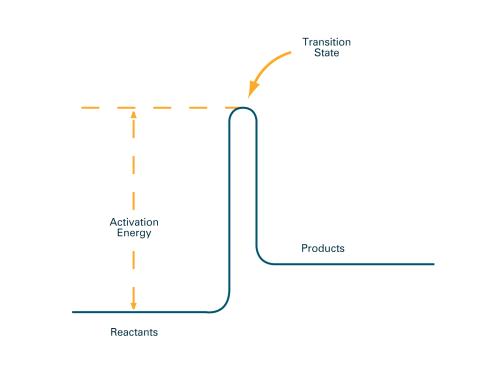
2526: Activation energy (with labels)
To become products, reactants must overcome an energy hill. See image 2525 for an unlabeled version of this illustration. Featured in The Chemistry of Health.
Crabtree + Company
View Media

3718: A Bacillus subtilis biofilm grown in a Petri dish
3718: A Bacillus subtilis biofilm grown in a Petri dish
Bacterial biofilms are tightly knit communities of bacterial cells growing on, for example, solid surfaces, such as in water pipes or on teeth. Here, cells of the bacterium Bacillus subtilis have formed a biofilm in a laboratory culture. Researchers have discovered that the bacterial cells in a biofilm communicate with each other through electrical signals via specialized potassium ion channels to share resources, such as nutrients, with each other. This insight may help scientists to improve sanitation systems to prevent biofilms, which often resist common treatments, from forming and to develop better medicines to combat bacterial infections. See the Biomedical Beat blog post Bacterial Biofilms: A Charged Environment for more information.
Gürol Süel, UCSD
View Media
2793: Anti-tumor drug ecteinascidin 743 (ET-743) with hydrogens 04
2793: Anti-tumor drug ecteinascidin 743 (ET-743) with hydrogens 04
Ecteinascidin 743 (ET-743, brand name Yondelis), was discovered and isolated from a sea squirt, Ecteinascidia turbinata, by NIGMS grantee Kenneth Rinehart at the University of Illinois. It was synthesized by NIGMS grantees E.J. Corey and later by Samuel Danishefsky. Multiple versions of this structure are available as entries 2790-2797.
Timothy Jamison, Massachusetts Institute of Technology
View Media
2795: Anti-tumor drug ecteinascidin 743 (ET-743), structure without hydrogens 02
2795: Anti-tumor drug ecteinascidin 743 (ET-743), structure without hydrogens 02
Ecteinascidin 743 (ET-743, brand name Yondelis), was discovered and isolated from a sea squirt, Ecteinascidia turbinata, by NIGMS grantee Kenneth Rinehart at the University of Illinois. It was synthesized by NIGMS grantees E.J. Corey and later by Samuel Danishefsky. Multiple versions of this structure are available as entries 2790-2797.
Timothy Jamison, Massachusetts Institute of Technology
View Media
6851: Himastatin, 360-degree view
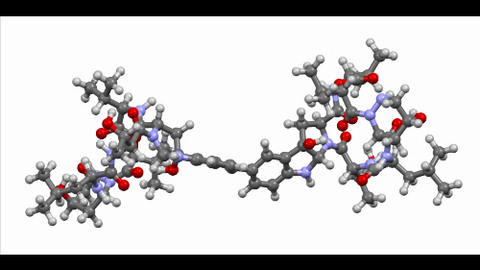
6851: Himastatin, 360-degree view
A 360-degree view of the molecule himastatin, which was first isolated from the bacterium Streptomyces himastatinicus. Himastatin shows antibiotic activity. The researchers who created this video developed a new, more concise way to synthesize himastatin so it can be studied more easily.
More information about the research that produced this video can be found in the Science paper “Total synthesis of himastatin” by D’Angelo et al.
Related to images 6848 and 6850.
More information about the research that produced this video can be found in the Science paper “Total synthesis of himastatin” by D’Angelo et al.
Related to images 6848 and 6850.
Mohammad Movassaghi, Massachusetts Institute of Technology.
View Media
2524: Plasma membrane (with labels)
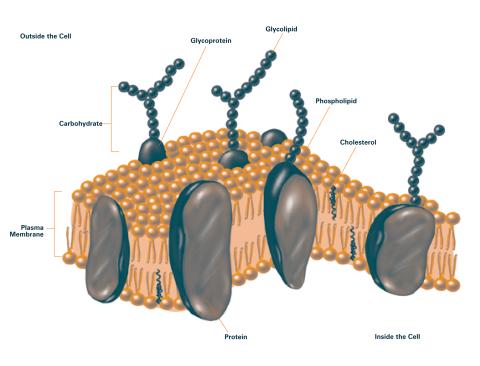
2524: Plasma membrane (with labels)
The plasma membrane is a cell's protective barrier. See image 2523 for an unlabeled version of this illustration. Featured in The Chemistry of Health.
Crabtree + Company
View Media
2746: Active site of sulfite oxidase
2746: Active site of sulfite oxidase
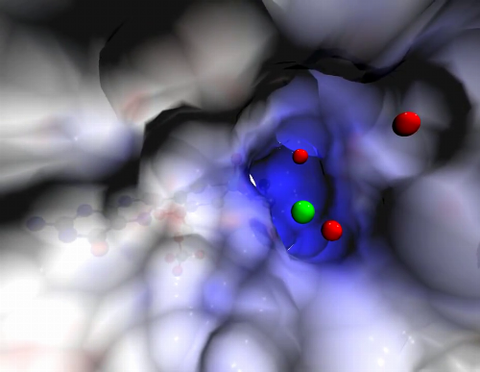
Sulfite oxidase is an enzyme that is essential for normal neurological development in children. This video shows the active site of the enzyme and its molybdenum cofactor visible as a faint ball-and-stick representation buried within the protein. The positively charged channel (blue) at the active site contains a chloride ion (green) and three water molecules (red). As the protein oscillates, one can see directly down the positively charged channel. At the bottom is the molybdenum atom of the active site (light blue) and its oxo group (red) that is transferred to sulfite to form sulfate in the catalytic reaction.
John Enemark, University of Arizona
View Media
2490: Cascade reaction promoted by water
2490: Cascade reaction promoted by water
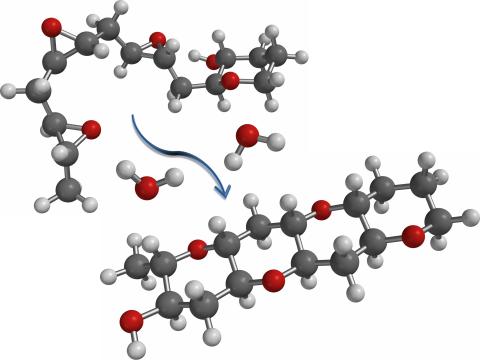
This illustration of an epoxide-opening cascade promoted by water emulates the proposed biosynthesis of some of the Red Tide toxins.
Tim Jamison, Massachusetts Institute of Technology
View Media
3429: Enzyme transition states
3429: Enzyme transition states
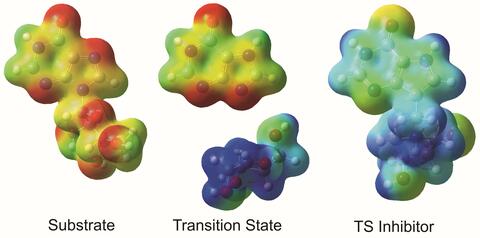
The molecule on the left is an electrostatic potential map of the van der Waals surface of the transition state for human purine nucleoside phosphorylase. The colors indicate the electron density at any position of the molecule. Red indicates electron-rich regions with negative charge and blue indicates electron-poor regions with positive charge. The molecule on the right is called DADMe-ImmH. It is a chemically stable analogue of the transition state on the left. It binds to the enzyme millions of times tighter than the substrate. This inhibitor is in human clinical trials for treating patients with gout. This image appears in Figure 4, Schramm, V.L. (2011) Annu. Rev. Biochem. 80:703-732.
Vern Schramm, Albert Einstein College of Medicine of Yeshiva University
View Media
2687: Serratezomine A
2687: Serratezomine A
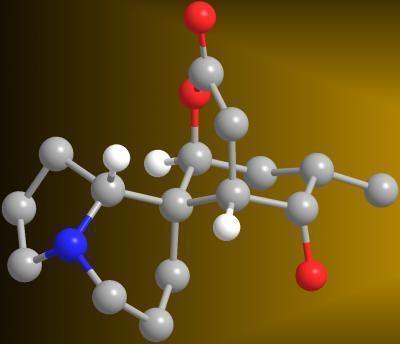
A 3-D model of the alkaloid serratezomine A shows the molecule's complex ring structure.
View Media
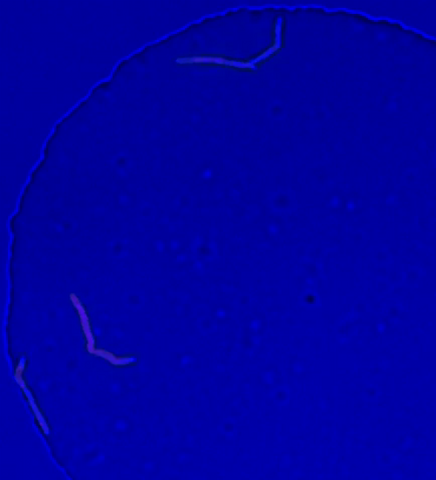

5752: Genetically identical mycobacteria respond differently to antibiotic 2
5752: Genetically identical mycobacteria respond differently to antibiotic 2
Antibiotic resistance in microbes is a serious health concern. So researchers have turned their attention to how bacteria undo the action of some antibiotics. Here, scientists set out to find the conditions that help individual bacterial cells survive in the presence of the antibiotic rifampicin. The research team used Mycobacterium smegmatis, a more harmless relative of Mycobacterium tuberculosis, which infects the lung and other organs to cause serious disease.
In this video, genetically identical mycobacteria are growing in a miniature growth chamber called a microfluidic chamber. Using live imaging, the researchers found that individual mycobacteria will respond differently to the antibiotic, depending on the growth stage and other timing factors. The researchers used genetic tagging with green fluorescent protein to distinguish cells that can resist rifampicin and those that cannot. With this gene tag, cells tolerant of the antibiotic light up in green and those that are susceptible in violet, enabling the team to monitor the cells' responses in real time.
To learn more about how the researchers studied antibiotic resistance in mycobacteria, see this news release from Tufts University. Related to image 5751.
In this video, genetically identical mycobacteria are growing in a miniature growth chamber called a microfluidic chamber. Using live imaging, the researchers found that individual mycobacteria will respond differently to the antibiotic, depending on the growth stage and other timing factors. The researchers used genetic tagging with green fluorescent protein to distinguish cells that can resist rifampicin and those that cannot. With this gene tag, cells tolerant of the antibiotic light up in green and those that are susceptible in violet, enabling the team to monitor the cells' responses in real time.
To learn more about how the researchers studied antibiotic resistance in mycobacteria, see this news release from Tufts University. Related to image 5751.
Bree Aldridge, Tufts University
View Media
2535: Kinases (with labels)
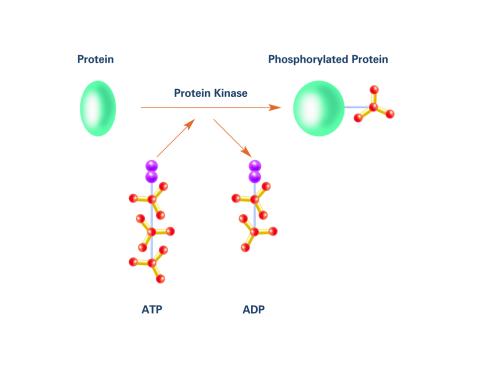
2535: Kinases (with labels)
Kinases are enzymes that add phosphate groups (red-yellow structures) to proteins (green), assigning the proteins a code. In this reaction, an intermediate molecule called ATP (adenosine triphosphate) donates a phosphate group from itself, becoming ADP (adenosine diphosphate). See image 2534 for an unlabeled version of this illustration. Featured in Medicines By Design.
Crabtree + Company
View Media
2533: Dose response curves
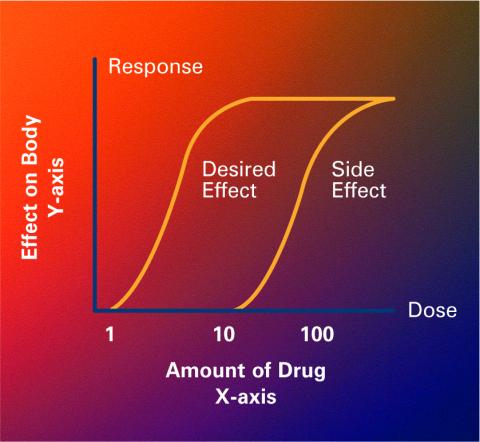
2533: Dose response curves
Dose-response curves determine how much of a drug (X-axis) causes a particular effect, or a side effect, in the body (Y-axis). Featured in Medicines By Design.
Crabtree + Company
View Media

3779: Precisely Delivering Chemical Cargo to Cells
3779: Precisely Delivering Chemical Cargo to Cells
Moving protein or other molecules to specific cells to treat or examine them has been a major biological challenge. Scientists have now developed a technique for delivering chemicals to individual cells. The approach involves gold nanowires that, for example, can carry tumor-killing proteins. The advance was possible after researchers developed electric tweezers that could manipulate gold nanowires to help deliver drugs to single cells.
This movie shows the manipulation of the nanowires for drug delivery to a single cell. To learn more about this technique, see this post in the Computing Life series.
This movie shows the manipulation of the nanowires for drug delivery to a single cell. To learn more about this technique, see this post in the Computing Life series.
Nature Nanotechnology
View Media
3789: Nucleolus subcompartments spontaneously self-assemble 1
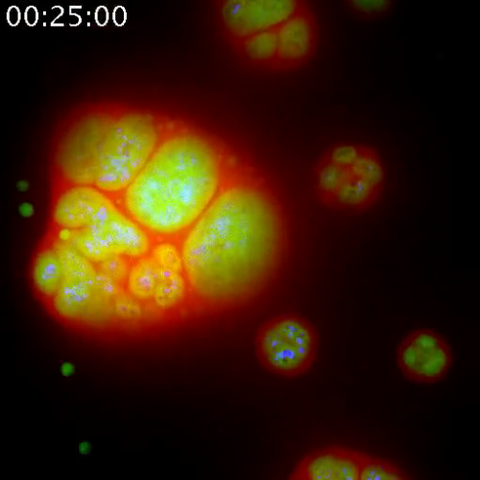
3789: Nucleolus subcompartments spontaneously self-assemble 1
The nucleolus is a small but very important protein complex located in the cell's nucleus. It forms on the chromosomes at the location where the genes for the RNAs are that make up the structure of the ribosome, the indispensable cellular machine that makes proteins from messenger RNAs.
However, how the nucleolus grows and maintains its structure has puzzled scientists for some time. It turns out that even though it looks like a simple liquid blob, it's rather well-organized, consisting of three distinct layers: the fibrillar center, where the RNA polymerase is active; the dense fibrillar component, which is enriched in the protein fibrillarin; and the granular component, which contains a protein called nucleophosmin. Researchers have now discovered that this multilayer structure of the nucleolus arises from difference in how the proteins in each compartment mix with water and with each other. These differences let them readily separate from each other into the three nucleolus compartments.
This video of nucleoli in the eggs of a commonly used lab animal, the frog Xenopus laevis, shows how each of the compartments (the granular component is shown in red, the fibrillarin in yellow-green, and the fibrillar center in blue) spontaneously fuse with each other on encounter without mixing with the other compartments. For more details on this research, see this press release from Princeton. Related to video 3791, image 3792 and image 3793.
However, how the nucleolus grows and maintains its structure has puzzled scientists for some time. It turns out that even though it looks like a simple liquid blob, it's rather well-organized, consisting of three distinct layers: the fibrillar center, where the RNA polymerase is active; the dense fibrillar component, which is enriched in the protein fibrillarin; and the granular component, which contains a protein called nucleophosmin. Researchers have now discovered that this multilayer structure of the nucleolus arises from difference in how the proteins in each compartment mix with water and with each other. These differences let them readily separate from each other into the three nucleolus compartments.
This video of nucleoli in the eggs of a commonly used lab animal, the frog Xenopus laevis, shows how each of the compartments (the granular component is shown in red, the fibrillarin in yellow-green, and the fibrillar center in blue) spontaneously fuse with each other on encounter without mixing with the other compartments. For more details on this research, see this press release from Princeton. Related to video 3791, image 3792 and image 3793.
Nilesh Vaidya, Princeton University
View Media
6762: CCP enzyme
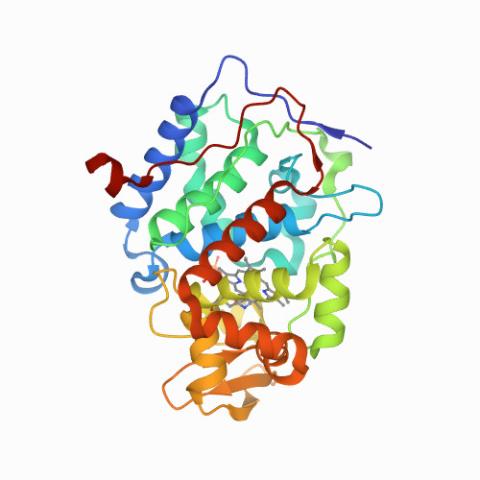
6762: CCP enzyme
The enzyme CCP is found in the mitochondria of baker’s yeast. Scientists study the chemical reactions that CCP triggers, which involve a water molecule, iron, and oxygen. This structure was determined using an X-ray free electron laser.
Protein Data Bank.
View Media
3419: X-ray co-crystal structure of Src kinase bound to a DNA-templated macrocycle inhibitor 7
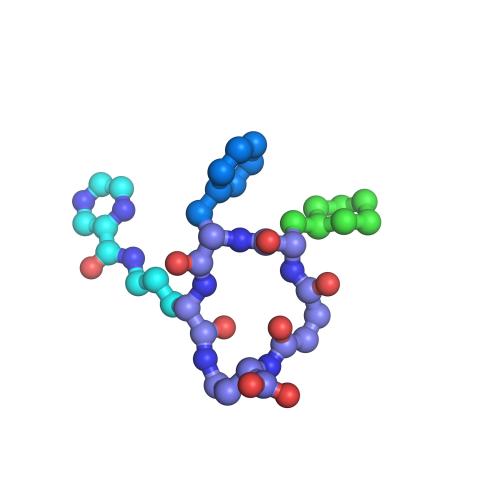
3419: X-ray co-crystal structure of Src kinase bound to a DNA-templated macrocycle inhibitor 7
X-ray co-crystal structure of Src kinase bound to a DNA-templated macrocycle inhibitor. Related to images 3413, 3414, 3415, 3416, 3417, and 3418.
Markus A. Seeliger, Stony Brook University Medical School and David R. Liu, Harvard University
View Media
3481: Bacillus anthracis being killed
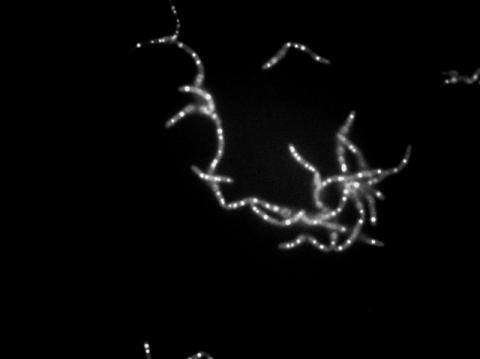
3481: Bacillus anthracis being killed
Bacillus anthracis (anthrax) cells being killed by a fluorescent trans-translation inhibitor, which disrupts bacterial protein synthesis. The inhibitor is naturally fluorescent and looks blue when it is excited by ultraviolet light in the microscope. This is a black-and-white version of Image 3525.
John Alumasa, Keiler Laboratory, Pennsylvania State University
View Media
3422: Atomic Structure of Poppy Enzyme
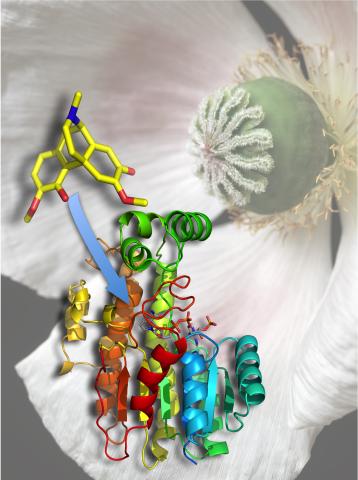
3422: Atomic Structure of Poppy Enzyme
The atomic structure of the morphine biosynthetic enzyme salutaridine reductase bound to the cofactor NADPH. The substrate salutaridine is shown entering the active site.
Judy Coyle, Donald Danforth Plant Science Center
View Media
2507: Carbon building blocks (with examples)
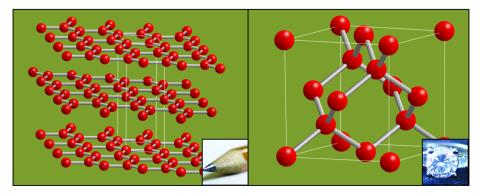
2507: Carbon building blocks (with examples)
The arrangement of identical molecular components can make a dramatic difference. For example, carbon atoms can be arranged into dull graphite (left) or sparkly diamonds (right). See image 2506 for an illustration without examples.
Crabtree + Company
View Media
6983: Genetic mosaicism in fruit flies
6983: Genetic mosaicism in fruit flies
Fat tissue from the abdomen of a genetically mosaic adult fruit fly. Genetic mosaicism means that the fly has cells with different genotypes even though it formed from a single zygote. This specific mosaicism results in accumulation of a critical fly adipokine (blue-green) within the fat tissue cells that have reduced expression a key nutrient sensing gene (in left panel). The dotted line shows the cells lacking the gene that is present and functioning in the rest of the cells. Nuclei are labelled in magenta. This image was captured using a confocal microscope and shows a maximum intensity projection of many slices.
Related to images 6982, 6984, and 6985.
Related to images 6982, 6984, and 6985.
Akhila Rajan, Fred Hutchinson Cancer Center
View Media
6848: Himastatin
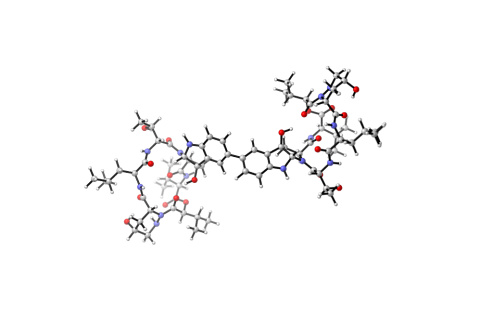
6848: Himastatin
A model of the molecule himastatin, which was first isolated from the bacterium Streptomyces himastatinicus. Himastatin shows antibiotic activity. The researchers who created this image developed a new, more concise way to synthesize himastatin so it can be studied more easily.
More information about the research that produced this image can be found in the Science paper “Total synthesis of himastatin” by D’Angelo et al.
Related to image 6850 and video 6851.
More information about the research that produced this image can be found in the Science paper “Total synthesis of himastatin” by D’Angelo et al.
Related to image 6850 and video 6851.
Mohammad Movassaghi, Massachusetts Institute of Technology.
View Media
2518: ATP synthase (with labels)
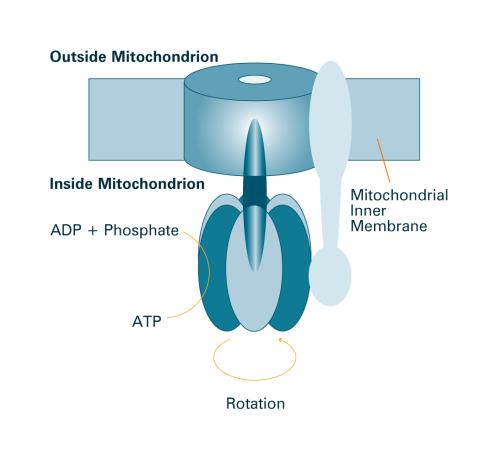
2518: ATP synthase (with labels)
The world's smallest motor, ATP synthase, generates energy for the cell. See image 2517 for an unlabeled version of this illustration. Featured in The Chemistry of Health.
Crabtree + Company
View Media
6779: Brain waves of a patient anesthetized with propofol
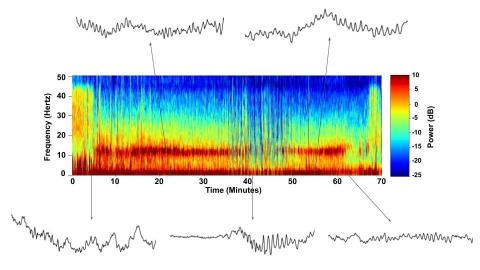
6779: Brain waves of a patient anesthetized with propofol
A representation of a patient’s brain waves after receiving the anesthetic propofol. All anesthetics create brain wave changes that vary depending on the patient’s age and the type and dose of anesthetic used. These changes are visible in raw electroencephalogram (EEG) readings, but they’re easier to interpret using a spectrogram where the signals are broken down by time (x-axis), frequency (y-axis), and power (color scale). This spectrogram shows the changes in brain waves before, during, and after propofol-induced anesthesia. The patient is unconscious from minute 5, upon propofol administration, through minute 69 (change in power and frequency). But, between minutes 35 and 48, the patient fell into a profound state of unconsciousness (disappearance of dark red oscillations between 8 to 12 Hz), which required the anesthesiologist to adjust the rate of propofol administration. The propofol was stopped at minute 62 and the patient woke up around minute 69.
Emery N. Brown, M.D., Ph.D., Massachusetts General Hospital/Harvard Medical School, Picower Institute for Learning and Memory, and Massachusetts Institute of Technology.
View Media
3525: Bacillus anthracis being killed
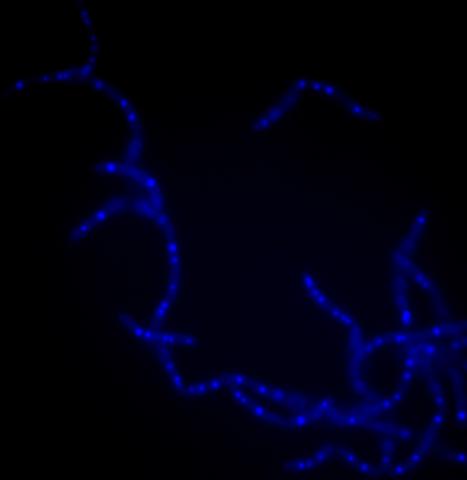
3525: Bacillus anthracis being killed
Bacillus anthracis (anthrax) cells being killed by a fluorescent trans-translation inhibitor, which disrupts bacterial protein synthesis. The inhibitor is naturally fluorescent and looks blue when it is excited by ultraviolet light in the microscope. This is a color version of Image 3481.
Kenneth Keiler, Penn State University
View Media

5895: Bioluminescence in a Tube
5895: Bioluminescence in a Tube
Details about the basic biology and chemistry of the ingredients that produce bioluminescence are allowing scientists to harness it as an imaging tool. Credit: Nathan Shaner, Scintillon Institute.
From Biomedical Beat article July 2017: Chasing Fireflies—and Better Cellular Imaging Techniques
From Biomedical Beat article July 2017: Chasing Fireflies—and Better Cellular Imaging Techniques
Nathan Shaner, Scintillon Institute
View Media
2534: Kinases
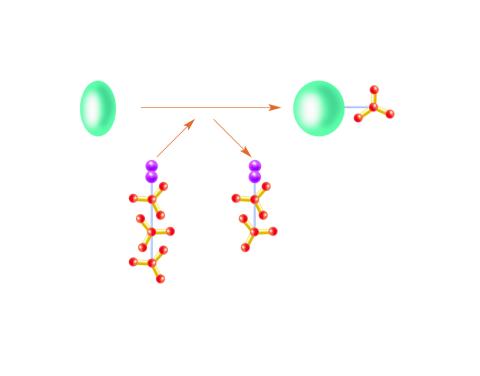
2534: Kinases
Kinases are enzymes that add phosphate groups (red-yellow structures) to proteins (green), assigning the proteins a code. In this reaction, an intermediate molecule called ATP (adenosine triphosphate) donates a phosphate group from itself, becoming ADP (adenosine diphosphate). See image 2535 for a labeled version of this illustration. Featured in Medicines By Design.
Crabtree + Company
View Media
6765: X-ray diffraction pattern from a crystallized cefotaxime-CCD-1 complex
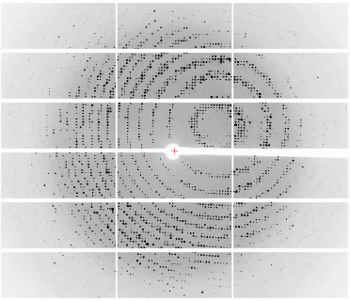
6765: X-ray diffraction pattern from a crystallized cefotaxime-CCD-1 complex
CCD-1 is an enzyme produced by the bacterium Clostridioides difficile that helps it resist antibiotics. Researchers crystallized complexes where a CCD-1 molecule and a molecule of the antibiotic cefotaxime were bound together. Then, they shot X-rays at the complexes to determine their structure—a process known as X-ray crystallography. This image shows the X-ray diffraction pattern of a complex.
Related to images 6764, 6766, and 6767.
Related to images 6764, 6766, and 6767.
Keith Hodgson, Stanford University.
View Media
2523: Plasma membrane
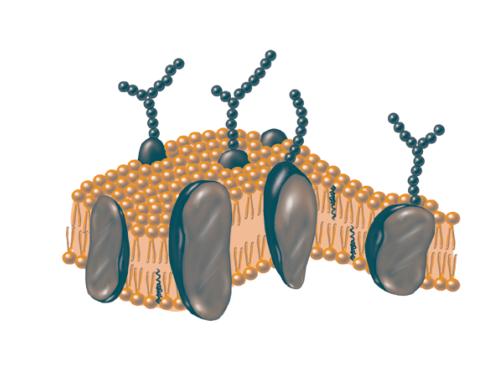
2523: Plasma membrane
The plasma membrane is a cell's protective barrier. See image 2524 for a labeled version of this illustration. Featured in The Chemistry of Health.
Crabtree + Company
View Media

3423: White Poppy (cropped)
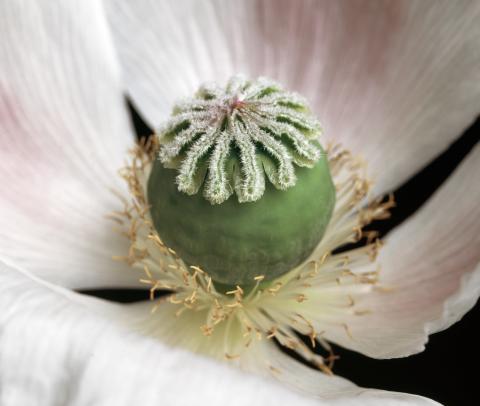
3423: White Poppy (cropped)
A cropped image of a white poppy. View poppy uncropped here 3424.
Judy Coyle, Donald Danforth Plant Science Center
View Media
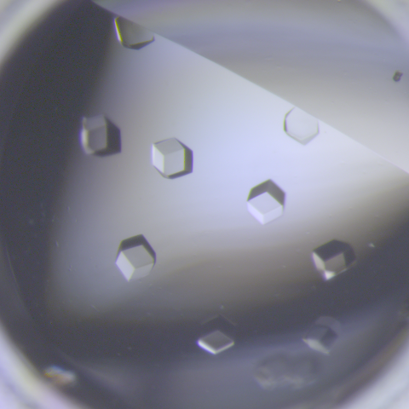

6764: Crystals of CCD-1 in complex with cefotaxime
6764: Crystals of CCD-1 in complex with cefotaxime
CCD-1 is an enzyme produced by the bacterium Clostridioides difficile that helps it resist antibiotics. Here, researchers crystallized bound pairs of CCD-1 molecules and molecules of the antibiotic cefotaxime. This enabled their structure to be studied using X-ray crystallography.
Related to images 6765, 6766, and 6767.
Related to images 6765, 6766, and 6767.
Keith Hodgson, Stanford University.
View Media
3787: In vitro assembly of a cell-signaling pathway
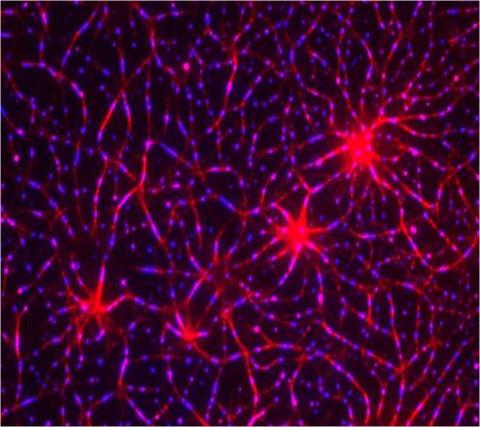
3787: In vitro assembly of a cell-signaling pathway
T cells are white blood cells that are important in defending the body against bacteria, viruses and other pathogens. Each T cell carries proteins, called T-cell receptors, on its surface that are activated when they come in contact with an invader. This activation sets in motion a cascade of biochemical changes inside the T cell to mount a defense against the invasion. Scientists have been interested for some time what happens after a T-cell receptor is activated. One obstacle has been to study how this signaling cascade, or pathway, proceeds inside T cells.
In this image, researchers have created a T-cell receptor pathway consisting of 12 proteins outside the cell on an artificial membrane. The image shows two key steps during the signaling process: clustering of a protein called linker for activation of T cells (LAT) (blue) and polymerization of the cytoskeleton protein actin (red). The findings show that the T-cell receptor signaling proteins self-organize into separate physical and biochemical compartments. This new system of studying molecular pathways outside the cells will enable scientists to better understand how the immune system combats microbes or other agents that cause infection.
To learn more how researchers assembled this T-cell receptor pathway, see this press release from HHMI's Marine Biological Laboratory Whitman Center. Related to video 3786.
In this image, researchers have created a T-cell receptor pathway consisting of 12 proteins outside the cell on an artificial membrane. The image shows two key steps during the signaling process: clustering of a protein called linker for activation of T cells (LAT) (blue) and polymerization of the cytoskeleton protein actin (red). The findings show that the T-cell receptor signaling proteins self-organize into separate physical and biochemical compartments. This new system of studying molecular pathways outside the cells will enable scientists to better understand how the immune system combats microbes or other agents that cause infection.
To learn more how researchers assembled this T-cell receptor pathway, see this press release from HHMI's Marine Biological Laboratory Whitman Center. Related to video 3786.
Xiaolei Su, HHMI Whitman Center of the Marine Biological Laboratory
View Media
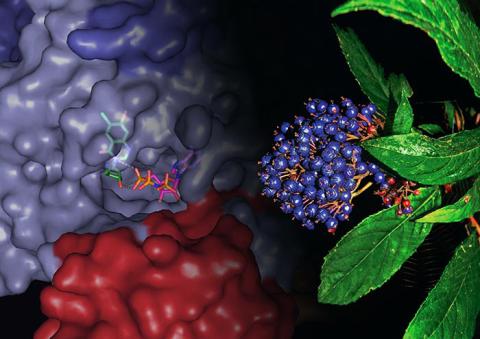
3483: Chang Shan
3483: Chang Shan
For thousands of years, Chinese herbalists have treated malaria using Chang Shan, a root extract from a type of hydrangea that grows in Tibet and Nepal. Recent studies have suggested Chang Shan can also reduce scar formation, treat multiple sclerosis and even slow cancer progression.
Paul Schimmel Lab, Scripps Research Institute
View Media
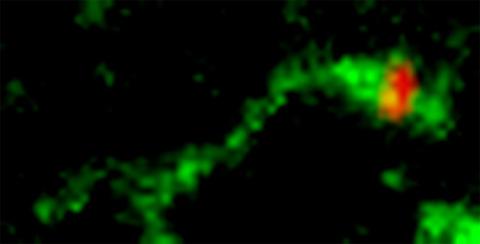
3488: Shiga toxin being sorted inside a cell
3488: Shiga toxin being sorted inside a cell
Shiga toxin (green) is sorted from the endosome into membrane tubules (red), which then pinch off and move to the Golgi apparatus.
Somshuvra Mukhopadhyay, The University of Texas at Austin, and Adam D. Linstedt, Carnegie Mellon University
View Media

3425: Red Poppy
2794: Anti-tumor drug ecteinascidin 743 (ET-743), structure without hydrogens 01
2794: Anti-tumor drug ecteinascidin 743 (ET-743), structure without hydrogens 01
Ecteinascidin 743 (ET-743, brand name Yondelis), was discovered and isolated from a sea squirt, Ecteinascidia turbinata, by NIGMS grantee Kenneth Rinehart at the University of Illinois. It was synthesized by NIGMS grantees E.J. Corey and later by Samuel Danishefsky. Multiple versions of this structure are available as entries 2790-2797.
Timothy Jamison, Massachusetts Institute of Technology
View Media
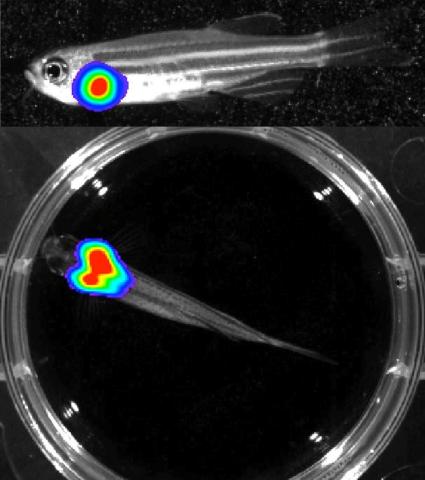
3556: Bioluminescent imaging in adult zebrafish - lateral and overhead view
3556: Bioluminescent imaging in adult zebrafish - lateral and overhead view
Luciferase-based imaging enables visualization and quantification of internal organs and transplanted cells in live adult zebrafish. In this image, a cardiac muscle-restricted promoter drives firefly luciferase expression. This is the lateral and overhead (Bottom) view.
For imagery of the overhead view go to 3557.
For imagery of the lateral view go to 3558.
For more information about the illumated area go to 3559.
For imagery of the overhead view go to 3557.
For imagery of the lateral view go to 3558.
For more information about the illumated area go to 3559.
Kenneth Poss, Duke University
View Media
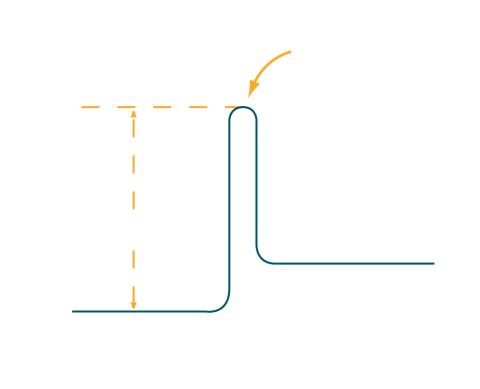
2525: Activation energy
2525: Activation energy
To become products, reactants must overcome an energy hill. See image 2526 for a labeled version of this illustration. Featured in The Chemistry of Health.
Crabtree + Company
View Media
3416: X-ray co-crystal structure of Src kinase bound to a DNA-templated macrocycle inhibitor 4
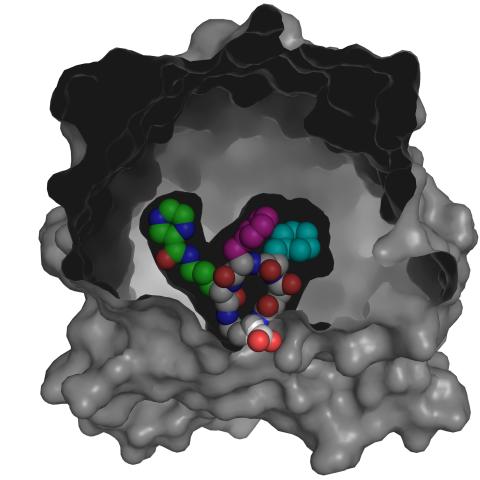
3416: X-ray co-crystal structure of Src kinase bound to a DNA-templated macrocycle inhibitor 4
X-ray co-crystal structure of Src kinase bound to a DNA-templated macrocycle inhibitor. Related to 3413, 3414, 3415, 3417, 3418, and 3419.
Markus A. Seeliger, Stony Brook University Medical School and David R. Liu, Harvard University
View Media
7000: Plastic-eating enzymes
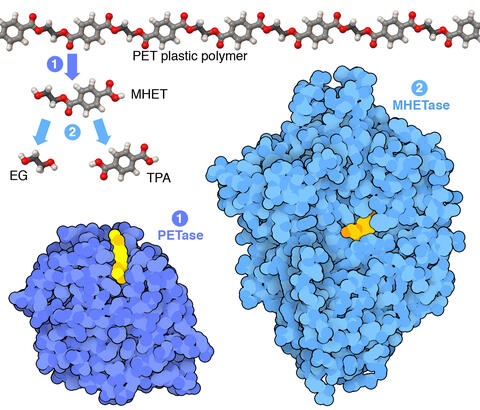
7000: Plastic-eating enzymes
PETase enzyme degrades polyester plastic (polyethylene terephthalate, or PET) into monohydroxyethyl terephthalate (MHET). Then, MHETase enzyme degrades MHET into its constituents ethylene glycol (EG) and terephthalic acid (TPA).
Find these in the RCSB Protein Data Bank: PET hydrolase (PDB entry 5XH3) and MHETase (PDB entry 6QGA).
Find these in the RCSB Protein Data Bank: PET hydrolase (PDB entry 5XH3) and MHETase (PDB entry 6QGA).
Amy Wu and Christine Zardecki, RCSB Protein Data Bank.
View Media

3786: Movie of in vitro assembly of a cell-signaling pathway
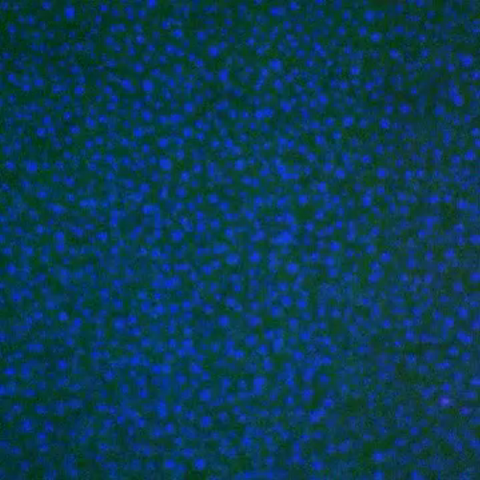
3786: Movie of in vitro assembly of a cell-signaling pathway
T cells are white blood cells that are important in defending the body against bacteria, viruses and other pathogens. Each T cell carries proteins, called T-cell receptors, on its surface that are activated when they come in contact with an invader. This activation sets in motion a cascade of biochemical changes inside the T cell to mount a defense against the invasion. Scientists have been interested for some time what happens after a T-cell receptor is activated. One obstacle has been to study how this signaling cascade, or pathway, proceeds inside T cells.
In this video, researchers have created a T-cell receptor pathway consisting of 12 proteins outside the cell on an artificial membrane. The video shows three key steps during the signaling process: phosphorylation of the T-cell receptor (green), clustering of a protein called linker for activation of T cells (LAT) (blue) and polymerization of the cytoskeleton protein actin (red). The findings show that the T-cell receptor signaling proteins self-organize into separate physical and biochemical compartments. This new system of studying molecular pathways outside the cells will enable scientists to better understand how the immune system combats microbes or other agents that cause infection.
To learn more how researchers assembled this T-cell receptor pathway, see this press release from HHMI's Marine Biological Laboratory Whitman Center. Related to image 3787.
In this video, researchers have created a T-cell receptor pathway consisting of 12 proteins outside the cell on an artificial membrane. The video shows three key steps during the signaling process: phosphorylation of the T-cell receptor (green), clustering of a protein called linker for activation of T cells (LAT) (blue) and polymerization of the cytoskeleton protein actin (red). The findings show that the T-cell receptor signaling proteins self-organize into separate physical and biochemical compartments. This new system of studying molecular pathways outside the cells will enable scientists to better understand how the immune system combats microbes or other agents that cause infection.
To learn more how researchers assembled this T-cell receptor pathway, see this press release from HHMI's Marine Biological Laboratory Whitman Center. Related to image 3787.
Xiaolei Su, HHMI Whitman Center of the Marine Biological Laboratory
View Media
2519: Bond types
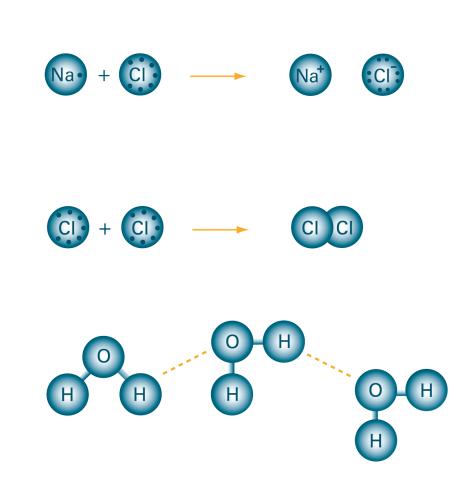
2519: Bond types
Ionic and covalent bonds hold molecules, like sodium chloride and chlorine gas, together. Hydrogen bonds among molecules, notably involving water, also play an important role in biology. See image 2520 for a labeled version of this illustration. Featured in The Chemistry of Health.
Crabtree + Company
View Media
6603: Protein formation
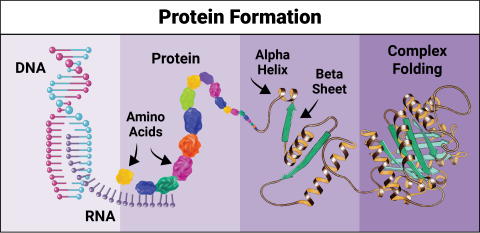
6603: Protein formation
Proteins are 3D structures made up of smaller units. DNA is transcribed to RNA, which in turn is translated into amino acids. Amino acids form a protein strand, which has sections of corkscrew-like coils, called alpha helices, and other sections that fold flat, called beta sheets. The protein then goes through complex folding to produce the 3D structure.
NIGMS, with the folded protein illustration adapted from Jane Richardson, Duke University Medical Center
View Media
6982: Insulin production and fat sensing in fruit flies
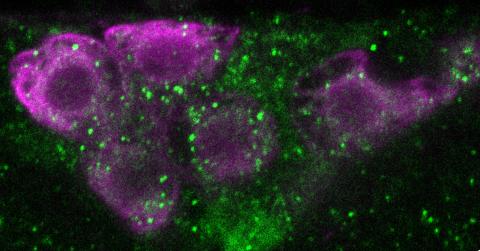
6982: Insulin production and fat sensing in fruit flies
Fourteen neurons (magenta) in the adult Drosophila brain produce insulin, and fat tissue sends packets of lipids to the brain via the lipoprotein carriers (green). This image was captured using a confocal microscope and shows a maximum intensity projection of many slices.
Related to images 6983, 6984, and 6985.
Related to images 6983, 6984, and 6985.
Akhila Rajan, Fred Hutchinson Cancer Center
View Media

3728: Quorum-sensing inhibitor limits bacterial growth
3728: Quorum-sensing inhibitor limits bacterial growth
To simulate the consequences of disrupting bacterial cell-to-cell communication, called quorum sensing, in the crypts (small chambers within the colon), the researchers experimented with an inhibitor molecule (i.e., antagonist) to turn off quorum sensing in methicillin-resistant Staphylococcus aureus (MRSA), an antibiotic-resistant strain of bacteria that often causes human infections. In this experiment, a medium promoting bacterial growth flows through experimental chambers mimicking the colon environment. The chambers on the right contained no antagonist. In the left chambers, after being added to the flowing medium, the quorum-sensing-inhibiting molecules quickly spread throughout the crevices, inactivating quorum sensing and reducing colonization. These results suggest a potential strategy for addressing MRSA virulence via inhibitors of bacterial communication. You can read more about this research here.
Minyoung Kevin Kim and Bonnie Bassler, Princeton University
View Media
7023: Dynein moving along microtubules
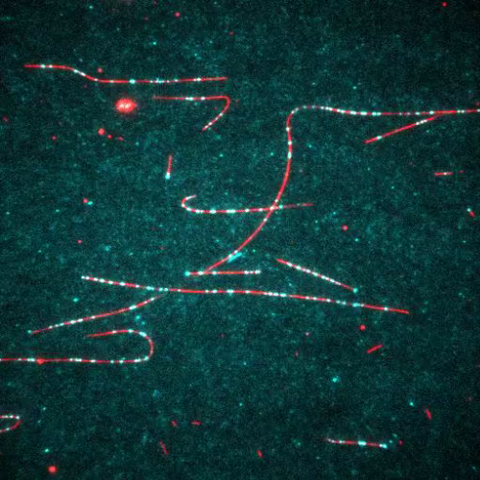
7023: Dynein moving along microtubules
Dynein (green) is a motor protein that “walks” along microtubules (red, part of the cytoskeleton) and carries its cargo along with it. This video was captured through fluorescence microscopy.
Morgan DeSantis, University of Michigan.
View Media
3747: Cryo-electron microscopy revealing the "wasabi receptor"
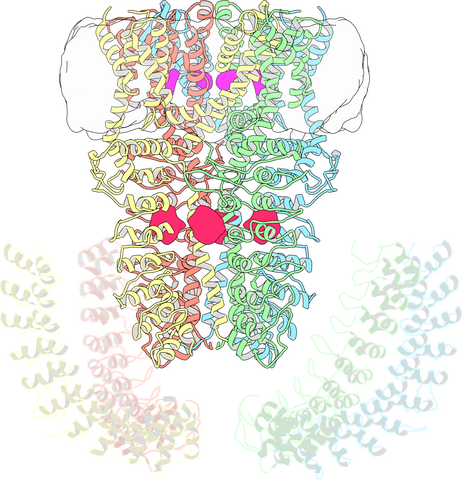
3747: Cryo-electron microscopy revealing the "wasabi receptor"
The TRPA1 protein is responsible for the burn you feel when you taste a bite of sushi topped with wasabi. Known therefore informally as the "wasabi receptor," this protein forms pores in the membranes of nerve cells that sense tastes or odors. Pungent chemicals like wasabi or mustard oil cause the pores to open, which then triggers a tingling or burn on our tongue. This receptor also produces feelings of pain in response to chemicals produced within our own bodies when our tissues are damaged or inflamed. Researchers used cryo-EM to reveal the structure of the wasabi receptor at a resolution of about 4 angstroms (a credit card is about 8 million angstroms thick). This detailed structure can help scientists understand both how we feel pain and how we can limit it by developing therapies to block the receptor. For more on cryo-EM see the blog post Cryo-Electron Microscopy Reveals Molecules in Ever Greater Detail.
Jean-Paul Armache, UCSF
View Media
