Switch to List View
Image and Video Gallery
This is a searchable collection of scientific photos, illustrations, and videos. The images and videos in this gallery are licensed under Creative Commons Attribution Non-Commercial ShareAlike 3.0. This license lets you remix, tweak, and build upon this work non-commercially, as long as you credit and license your new creations under identical terms.
2792: Anti-tumor drug ecteinascidin 743 (ET-743) with hydrogens 03
2792: Anti-tumor drug ecteinascidin 743 (ET-743) with hydrogens 03
Ecteinascidin 743 (ET-743, brand name Yondelis), was discovered and isolated from a sea squirt, Ecteinascidia turbinata, by NIGMS grantee Kenneth Rinehart at the University of Illinois. It was synthesized by NIGMS grantees E.J. Corey and later by Samuel Danishefsky. Multiple versions of this structure are available as entries 2790-2797.
Timothy Jamison, Massachusetts Institute of Technology
View Media
3334: Four timepoints in gastrulation
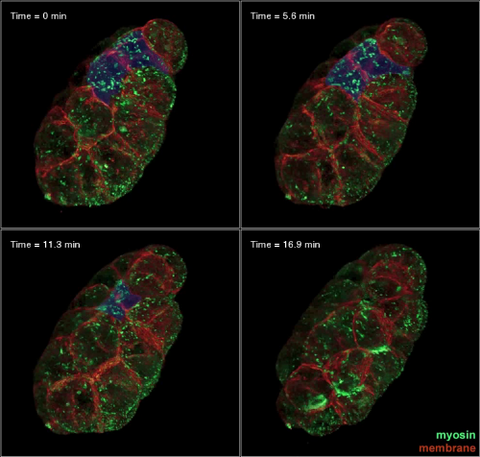
3334: Four timepoints in gastrulation
It has been said that gastrulation is the most important event in a person's life. This part of early embryonic development transforms a simple ball of cells and begins to define cell fate and the body axis. In a study published in Science magazine, NIGMS grantee Bob Goldstein and his research group studied how contractions of actomyosin filaments in C. elegans and Drosophila embryos lead to dramatic rearrangements of cell and embryonic structure. In these images, myosin (green) and plasma membrane (red) are highlighted at four timepoints in gastrulation in the roundworm C. elegans. The blue highlights in the top three frames show how cells are internalized, and the site of closure around the involuting cells is marked with an arrow in the last frame. See related image 3297.
Bob Goldstein, University of North Carolina, Chapel Hill
View Media
2626: Telomeres
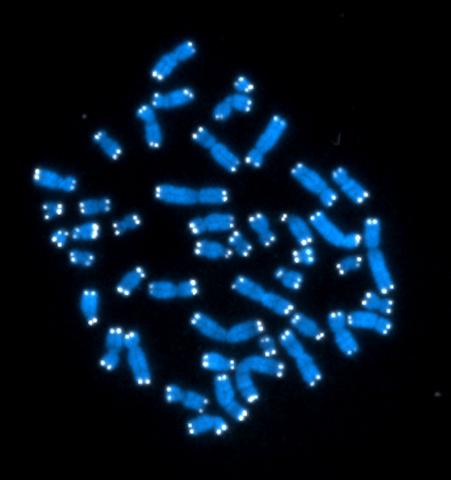
2626: Telomeres
The 46 human chromosomes are shown in blue, with the telomeres appearing as white pinpoints. The DNA has already been copied, so each chromosome is actually made up of two identical lengths of DNA, each with its own two telomeres.
Hesed Padilla-Nash and Thomas Ried, the National Cancer Institute, a part of NIH
View Media
1312: Cell toxins
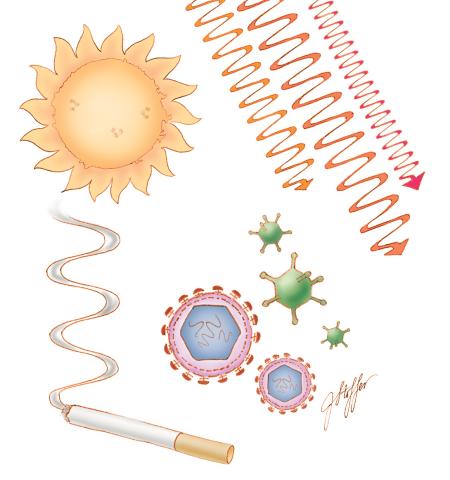
1312: Cell toxins
A number of environmental factors cause DNA mutations that can lead to cancer: toxins in cigarette smoke, sunlight and other radiation, and some viruses.
Judith Stoffer
View Media
6748: Human retinal organoid
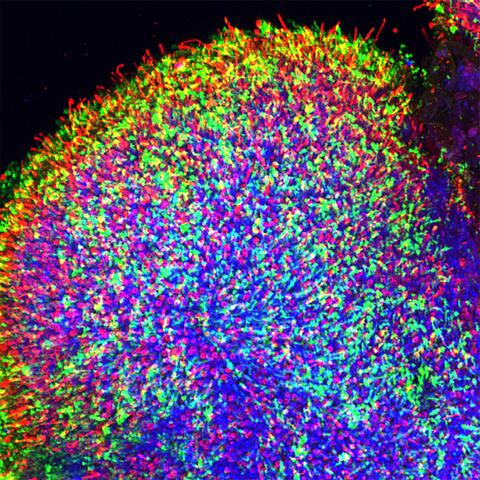
6748: Human retinal organoid
A replica of a human retina grown from stem cells. It shows rod photoreceptors (nerve cells responsible for dark vision) in green and red/green cones (nerve cells responsible for red and green color vision) in red. The cell nuclei are stained blue. This image was captured using a confocal microscope.
Kevin Eliceiri, University of Wisconsin-Madison.
View Media
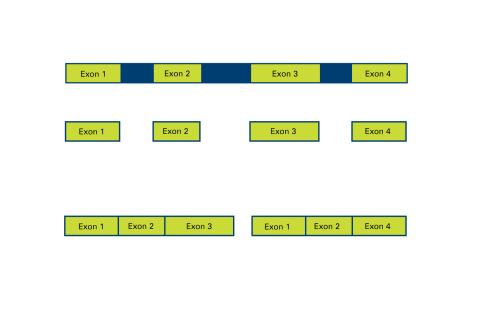
2552: Alternative splicing
2552: Alternative splicing
Arranging exons in different patterns, called alternative splicing, enables cells to make different proteins from a single gene. See image 2553 for a labeled version of this illustration. Featured in The New Genetics.
Crabtree + Company
View Media
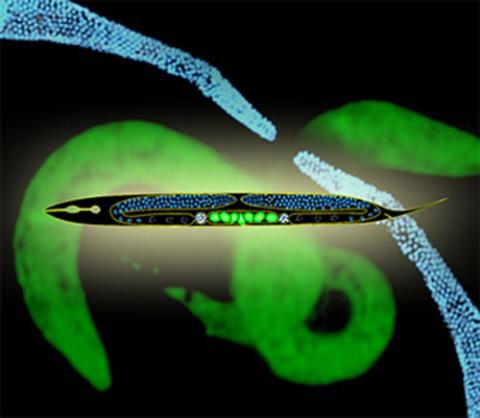
2333: Worms and human infertility
2333: Worms and human infertility
This montage of tiny, transparent C. elegans--or roundworms--may offer insight into understanding human infertility. Researchers used fluorescent dyes to label the worm cells and watch the process of sex cell division, called meiosis, unfold as nuclei (blue) move through the tube-like gonads. Such visualization helps the scientists identify mechanisms that enable these roundworms to reproduce successfully. Because meiosis is similar in all sexually reproducing organisms, what the scientists learn could apply to humans.
Abby Dernburg, Lawrence Berkeley National Laboratory
View Media
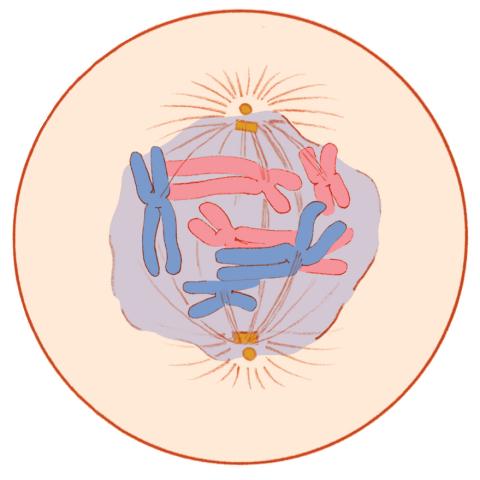
1331: Mitosis - prometaphase
1331: Mitosis - prometaphase
A cell in prometaphase during mitosis: The nuclear membrane breaks apart, and the spindle starts to interact with the chromosomes. Mitosis is responsible for growth and development, as well as for replacing injured or worn out cells throughout the body. For simplicity, mitosis is illustrated here with only six chromosomes.
Judith Stoffer
View Media
5760: Annotated TEM cross-section of C. elegans (roundworm)
5760: Annotated TEM cross-section of C. elegans (roundworm)
The worm Caenorhabditis elegans is a popular laboratory animal because its small size and fairly simple body make it easy to study. Scientists use this small worm to answer many research questions in developmental biology, neurobiology, and genetics. This image, which was taken with transmission electron microscopy (TEM), shows a cross-section through C. elegans, revealing various internal structures labeled in the image. You can find a high-resolution image without the annotations at image 5759.
The image is from a figure in an article published in the journal eLife.
The image is from a figure in an article published in the journal eLife.
Piali Sengupta, Brandeis University
View Media
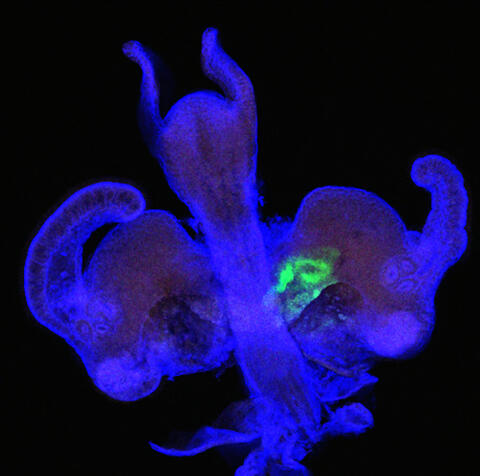
7020: Bacterial symbionts colonizing the crypts of a juvenile Hawaiian bobtail squid light organ
7020: Bacterial symbionts colonizing the crypts of a juvenile Hawaiian bobtail squid light organ
A light organ (~0.5 mm across) of a Hawaiian bobtail squid, Euprymna scolopes, stained blue. At the time of this image, the crypts within the tissues of only one side of the organ had been colonized by green-fluorescent protein-labeled Vibrio fischeri cells, which can be seen here in green. This image was taken using confocal fluorescence microscopy.
Related to images 7016, 7017, 7018, and 7019.
Related to images 7016, 7017, 7018, and 7019.
Margaret J. McFall-Ngai, Carnegie Institution for Science/California Institute of Technology, and Edward G. Ruby, California Institute of Technology.
View Media
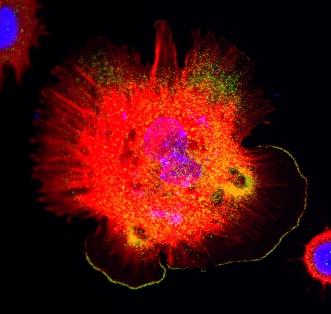
3328: Spreading Cells 01
3328: Spreading Cells 01
Cells move forward with lamellipodia and filopodia supported by networks and bundles of actin filaments. Proper, controlled cell movement is a complex process. Recent research has shown that an actin-polymerizing factor called the Arp2/3 complex is the key component of the actin polymerization engine that drives amoeboid cell motility. ARPC3, a component of the Arp2/3 complex, plays a critical role in actin nucleation. In this photo, the ARPC3+/+ fibroblast cells were fixed and stained with Alexa 546 phalloidin for F-actin (red), Arp2 (green), and DAPI to visualize the nucleus (blue). Arp2, a subunit of the Arp2/3 complex, is localized at the lamellipodia leading edge of ARPC3+/+ fibroblast cells. Related to images 3329, 3330, 3331, 3332, and 3333.
Rong Li and Praveen Suraneni, Stowers Institute for Medical Research
View Media
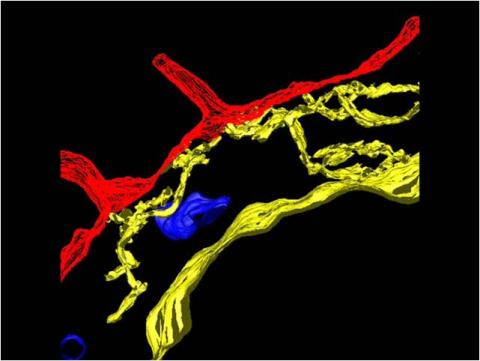
2636: Computer model of cell membrane
2636: Computer model of cell membrane
A computer model of the cell membrane, where the plasma membrane is red, endoplasmic reticulum is yellow, and mitochondria are blue. This image relates to a July 27, 2009 article in Computing Life.
Bridget Wilson, University of New Mexico
View Media
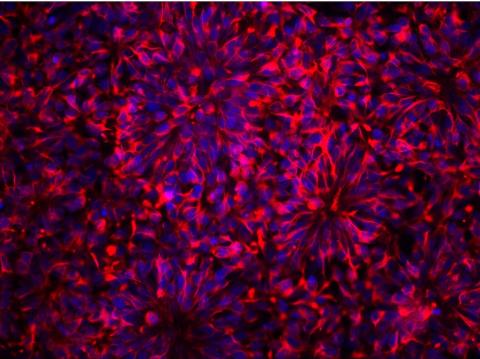
3284: Neurons from human ES cells
3284: Neurons from human ES cells
These neural precursor cells were derived from human embryonic stem cells. The neural cell bodies are stained red, and the nuclei are blue. Image and caption information courtesy of the California Institute for Regenerative Medicine.
Xianmin Zeng lab, Buck Institute for Age Research, via CIRM
View Media
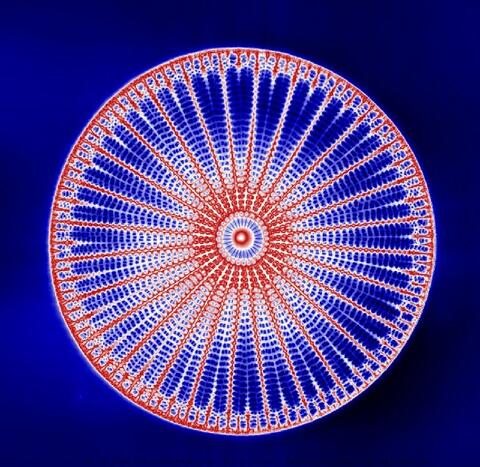

6902: Arachnoidiscus diatom
6902: Arachnoidiscus diatom
An Arachnoidiscus diatom with a diameter of 190µm. Diatoms are microscopic algae that have cell walls made of silica, which is the strongest known biological material relative to its density. In Arachnoidiscus, the cell wall is a radially symmetric pillbox-like shell composed of overlapping halves that contain intricate and delicate patterns. Sometimes, Arachnoidiscus is called “a wheel of glass.”
This image was taken with the orientation-independent differential interference contrast microscope.
This image was taken with the orientation-independent differential interference contrast microscope.
Michael Shribak, Marine Biological Laboratory/University of Chicago.
View Media
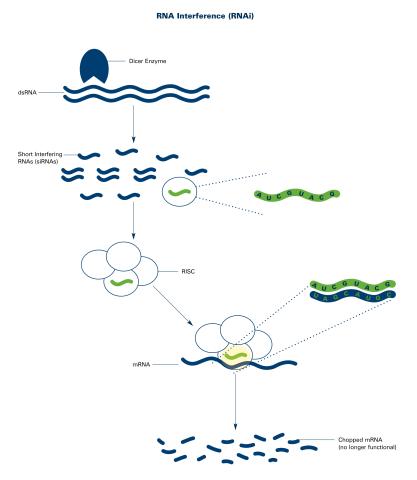
2559: RNA interference (with labels)
2559: RNA interference (with labels)
RNA interference or RNAi is a gene-silencing process in which double-stranded RNAs trigger the destruction of specific RNAs. See 2558 for an unlabeled version of this illustration. Featured in The New Genetics.
Crabtree + Company
View Media
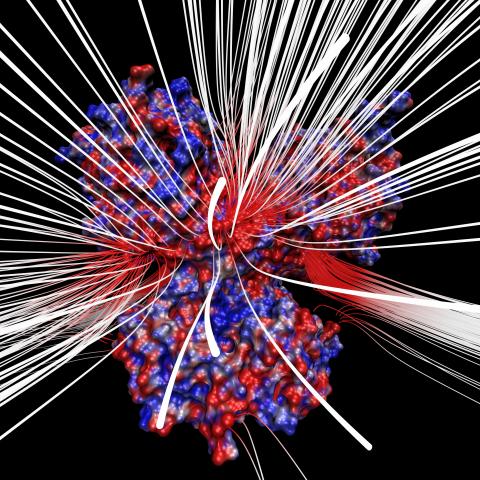

3658: Electrostatic map of human spermine synthase
3658: Electrostatic map of human spermine synthase
From PDB entry 3c6k, Crystal structure of human spermine synthase in complex with spermidine and 5-methylthioadenosine.
Emil Alexov, Clemson University
View Media

3592: Math from the heart
3592: Math from the heart
Watch a cell ripple toward a beam of light that turns on a movement-related protein.
View Media
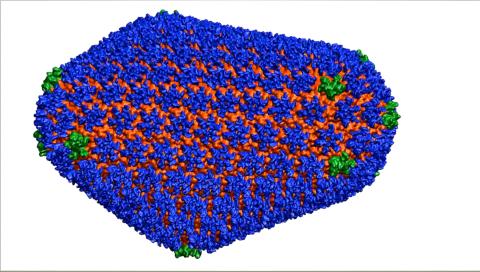

3477: HIV Capsid
3477: HIV Capsid
This image is a computer-generated model of the approximately 4.2 million atoms of the HIV capsid, the shell that contains the virus' genetic material. Scientists determined the exact structure of the capsid and the proteins that it's made of using a variety of imaging techniques and analyses. They then entered these data into a supercomputer that produced the atomic-level image of the capsid. This structural information could be used for developing drugs that target the capsid, possibly leading to more effective therapies. Related to image 6601.
Juan R. Perilla and the Theoretical and Computational Biophysics Group, University of Illinois at Urbana-Champaign
View Media
2439: Hydra 03
2439: Hydra 03
Hydra magnipapillata is an invertebrate animal used as a model organism to study developmental questions, for example the formation of the body axis.
Hiroshi Shimizu, National Institute of Genetics in Mishima, Japan
View Media
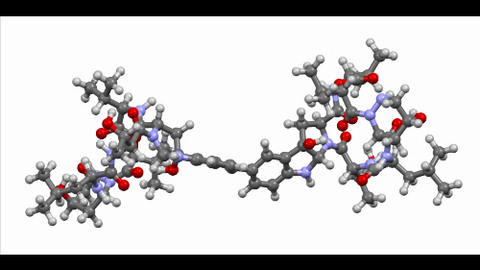
6851: Himastatin, 360-degree view
6851: Himastatin, 360-degree view
A 360-degree view of the molecule himastatin, which was first isolated from the bacterium Streptomyces himastatinicus. Himastatin shows antibiotic activity. The researchers who created this video developed a new, more concise way to synthesize himastatin so it can be studied more easily.
More information about the research that produced this video can be found in the Science paper “Total synthesis of himastatin” by D’Angelo et al.
Related to images 6848 and 6850.
More information about the research that produced this video can be found in the Science paper “Total synthesis of himastatin” by D’Angelo et al.
Related to images 6848 and 6850.
Mohammad Movassaghi, Massachusetts Institute of Technology.
View Media

2371: NMR spectrometer
2371: NMR spectrometer
This photo shows a Varian Unity Inova 900 MHz, 21.1 T standard bore magnet Nuclear Magnetic Resonnance (NMR) spectrometer. NMR spectroscopy provides data used to determine the structures of proteins in solution, rather than in crystal form, as in X-ray crystallography. The technique is limited to smaller proteins or protein fragments in a high throughput approach.
Center for Eukaryotic Structural Genomics
View Media
2491: VDAC-1 (2)
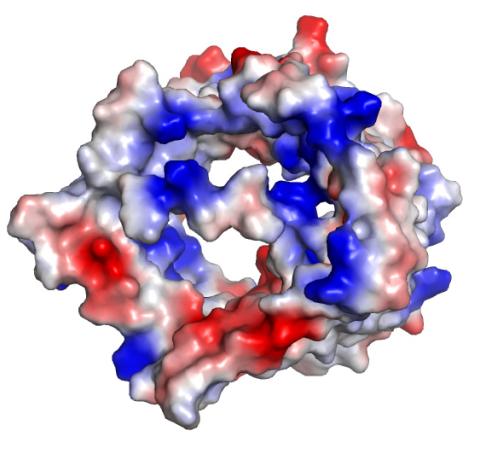
2491: VDAC-1 (2)
The structure of the pore-forming protein VDAC-1 from humans. This molecule mediates the flow of products needed for metabolism--in particular the export of ATP--across the outer membrane of mitochondria, the power plants for eukaryotic cells. VDAC-1 is involved in metabolism and the self-destruction of cells--two biological processes central to health.
Related to images 2494, 2495, and 2488.
Related to images 2494, 2495, and 2488.
Gerhard Wagner, Harvard Medical School
View Media
3754: Circadian rhythm neurons in the fruit fly brain
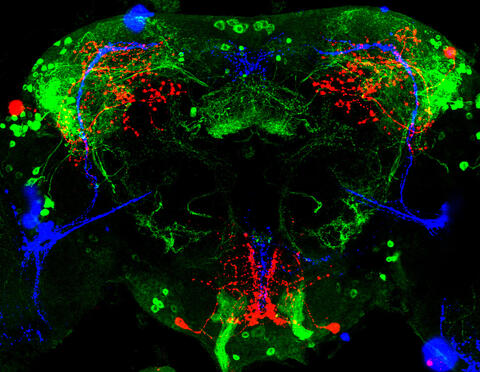
3754: Circadian rhythm neurons in the fruit fly brain
Some nerve cells (neurons) in the brain keep track of the daily cycle. This time-keeping mechanism, called the circadian clock, is found in all animals including us. The circadian clock controls our daily activities such as sleep and wakefulness. Researchers are interested in finding the neuron circuits involved in this time keeping and how the information about daily time in the brain is relayed to the rest of the body. In this image of a brain of the fruit fly Drosophila the time-of-day information flowing through the brain has been visualized by staining the neurons involved: clock neurons (shown in blue) function as "pacemakers" by communicating with neurons that produce a short protein called leucokinin (LK) (red), which, in turn, relays the time signal to other neurons, called LK-R neurons (green). This signaling cascade set in motion by the pacemaker neurons helps synchronize the fly's daily activity with the 24-hour cycle. To learn more about what scientists have found out about circadian pacemaker neurons in the fruit fly see this news release by New York University. This work was featured in the Biomedical Beat blog post Cool Image: A Circadian Circuit.
Justin Blau, New York University
View Media
3326: Cytochrome structure with anticancer drug
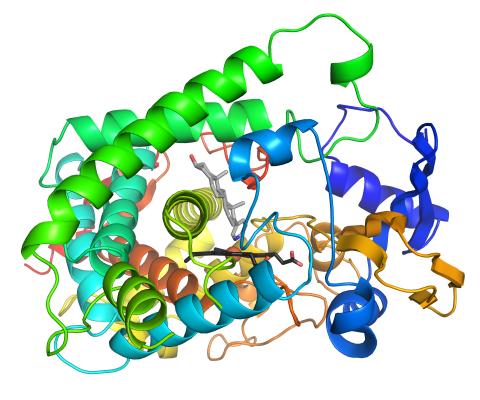
3326: Cytochrome structure with anticancer drug
This image shows the structure of the CYP17A1 enzyme (ribbons colored from blue N-terminus to red C-terminus), with the associated heme colored black. The prostate cancer drug abiraterone is colored gray. Cytochrome P450 enzymes bind to and metabolize a variety of chemicals, including drugs. Cytochrome P450 17A1 also helps create steroid hormones. Emily Scott's lab is studying how CYP17A1 could be selectively inhibited to treat prostate cancer. She and graduate student Natasha DeVore elucidated the structure shown using X-ray crystallography. Dr. Scott created the image (both white bg and transparent bg) for the NIGMS image gallery. See the "Medium-Resolution Image" for a PNG version of the image that is transparent.
Emily Scott, University of Kansas
View Media
1251: Crab larva eye
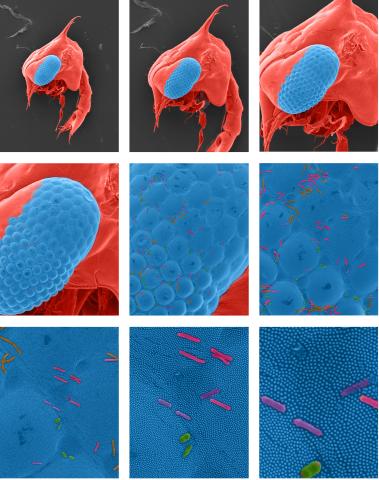
1251: Crab larva eye
Colorized scanning electron micrographs progressively zoom in on the eye of a crab larva. In the higher-resolution frames, bacteria are visible on the eye.
Tina Weatherby Carvalho, University of Hawaii at Manoa
View Media

1287: Mitochondria
1287: Mitochondria
Bean-shaped mitochondria are cells' power plants. These organelles have their own DNA and replicate independently. The highly folded inner membranes are the site of energy generation.
Judith Stoffer
View Media
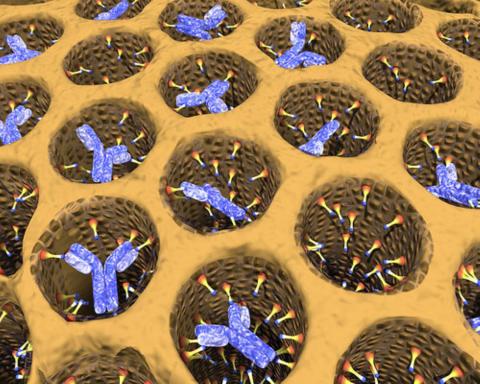

2750: Antibodies in silica honeycomb
2750: Antibodies in silica honeycomb
Antibodies are among the most promising therapies for certain forms of cancer, but patients must take them intravenously, exposing healthy tissues to the drug and increasing the risk of side effects. A team of biochemists packed the anticancer antibodies into porous silica particles to deliver a heavy dose directly to tumors in mice.
Chenghong Lei, Pacific Northwest National Laboratory & Karl Erik Hellstrom, University of Washington
View Media
2780: Arabidopsis leaf injected with a pathogen
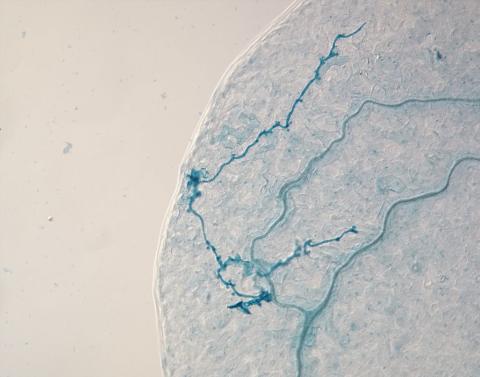
2780: Arabidopsis leaf injected with a pathogen
This is a magnified view of an Arabidopsis thaliana leaf eight days after being infected with the pathogen Hyaloperonospora arabidopsidis, which is closely related to crop pathogens that cause 'downy mildew' diseases. It is also more distantly related to the agent that caused the Irish potato famine. The veins of the leaf are light blue; in darker blue are the pathogen's hyphae growing through the leaf. The small round blobs along the length of the hyphae are called haustoria; each is invading a single plant cell to suck nutrients from the cell. Jeff Dangl and other NIGMS-supported researchers investigate how this pathogen and other like it use virulence mechanisms to suppress host defense and help the pathogens grow.
Jeff Dangl, University of North Carolina, Chapel Hill
View Media
1158: Bacteria shapes
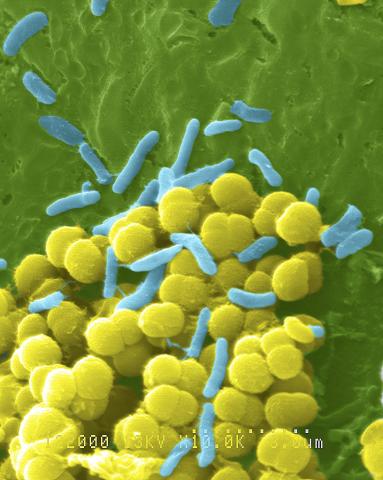
1158: Bacteria shapes
A colorized scanning electron micrograph of bacteria. Scanning electron microscopes allow scientists to see the three-dimensional surface of their samples.
Tina Weatherby Carvalho, University of Hawaii at Manoa
View Media
6806: Wild-type and mutant fruit fly ovaries
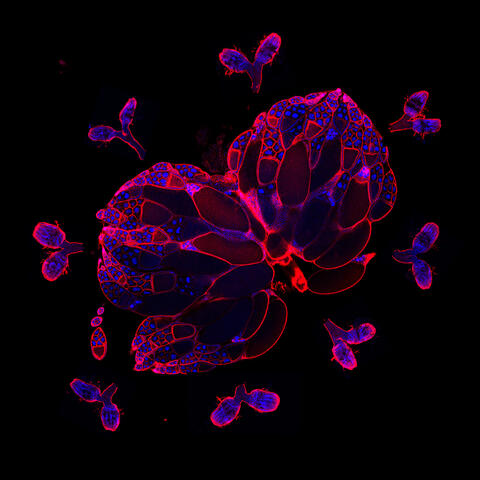
6806: Wild-type and mutant fruit fly ovaries
The two large, central, round shapes are ovaries from a typical fruit fly (Drosophila melanogaster). The small butterfly-like structures surrounding them are fruit fly ovaries where researchers suppressed the expression of a gene that controls microtubule polymerization and is necessary for normal development. This image was captured using a confocal laser scanning microscope.
Related to image 6807.
Related to image 6807.
Vladimir I. Gelfand, Feinberg School of Medicine, Northwestern University.
View Media

3475: Automated Worm Sorter - 4
3475: Automated Worm Sorter - 4
Georgia Tech associate professor Hang Lu holds a microfluidic chip that is part of a system that uses artificial intelligence and cutting-edge image processing to automatically examine large number of nematodes used for genetic research.
Georgia Tech/Gary Meek
View Media
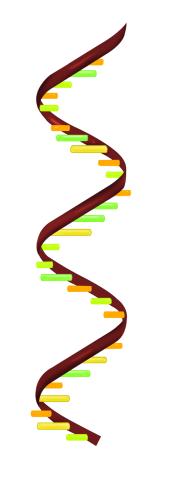
2554: RNA strand
2554: RNA strand
Ribonucleic acid (RNA) has a sugar-phosphate backbone and the bases adenine (A), cytosine (C), guanine (G), and uracil (U). See image 2555 for a labeled version of this illustration. Featured in The New Genetics.
Crabtree + Company
View Media
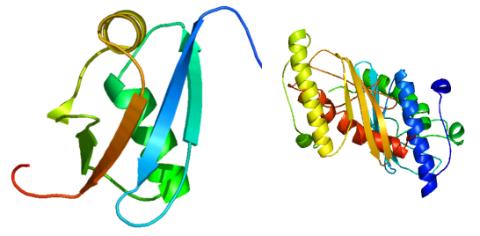
2388: Ubiquitin-fold modifier 1 from C. elegans
2388: Ubiquitin-fold modifier 1 from C. elegans
Solution NMR structure of protein target WR41 (left) from C. elegans. Noting the unanticipated structural similarity to the ubiquitin protein (Ub) found in all eukaryotic cells, researchers discovered that WR41 is a Ub-like modifier, ubiquitin-fold modifier 1 (Ufm1), on a newly uncovered ubiquitin-like pathway. Subsequently, the PSI group also determined the three-dimensional structure of protein target HR41 (right) from humans, the E2 ligase for Ufm1, using both NMR and X-ray crystallography.
Northeast Structural Genomics Consortium
View Media
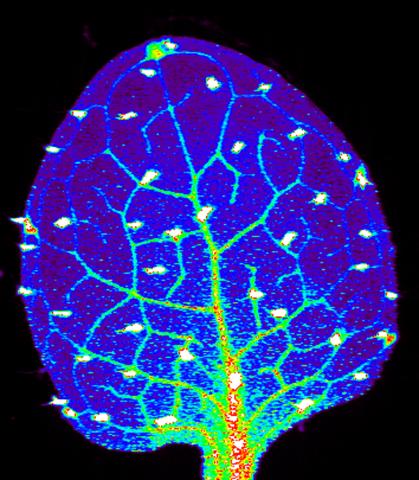
3727: Zinc levels in a plant leaf
3727: Zinc levels in a plant leaf
Zinc is required for the function of more than 300 enzymes, including those that help regulate gene expression, in various organisms including humans. Researchers study how plants acquire, sequester and distribute zinc to find ways to increase the zinc content of crops to improve human health. Using synchrotron X-ray fluorescence technology, they created this heat map of zinc levels in an Arabidopsis thaliana plant leaf. This image is a winner of the 2015 FASEB Bioart contest and was featured in the NIH Director's blog.
Suzana Car, Dartmouth College
View Media
2315: Fly cells live
2315: Fly cells live
If a picture is worth a thousand words, what's a movie worth? For researchers studying cell migration, a "documentary" of fruit fly cells (bright green) traversing an egg chamber could answer longstanding questions about cell movement. Historically, researchers have been unable to watch this cell migration unfold in living ovarian tissue in real time. But by developing a culture medium that allows fly eggs to survive outside their ovarian homes, scientists can observe the nuances of cell migration as it happens. Such details may shed light on how immune cells move to a wound and why cancer cells spread to other sites. See 3594 for still image.
Denise Montell, Johns Hopkins University School of Medicine
View Media

2667: Glowing fish
2667: Glowing fish
Professor Marc Zimmer's family pets, including these fish, glow in the dark in response to blue light. Featured in the September 2009 issue of Findings.
View Media

3272: Ear hair cells derived from embryonic stem cells
3272: Ear hair cells derived from embryonic stem cells
Mouse embryonic stem cells matured into this bundle of hair cells similar to the ones that transmit sound in the ear. These cells could one day be transplanted as a therapy for some forms of deafness, or they could be used to screen drugs to treat deafness. The hairs are shown at 23,000 times magnification via scanning electron microscopy. Image and caption information courtesy of the California Institute for Regenerative Medicine.
Stefen Heller, Stanford University, via CIRM
View Media
2574: Simulation of uncontrolled avian flu outbreak
2574: Simulation of uncontrolled avian flu outbreak
This video simulation shows what an uncontrolled outbreak of transmissible avian flu among people living in Thailand might look like. Red indicates new cases while green indicates areas where the epidemic has finished. The video shows the spread of infection and recovery over 300 days in Thailand and neighboring countries.
Neil M. Ferguson, Imperial College London
View Media
3478: DDR2 Receptors Attach to Collagen in Breast Tumor
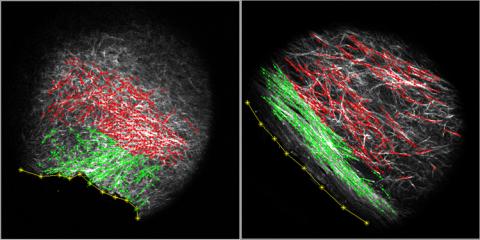
3478: DDR2 Receptors Attach to Collagen in Breast Tumor
On the left, the boundary of a breast tumor (yellow) attaches to collagen fibers that are closest to it (green) using DDR2. On the right, a tumor without DDR2 remains disconnected from the collagen.
Callie Corsa and Suzanne Ponik, Washington University School of Medicine in St. Louis
View Media
3765: Trypanosoma brucei, the cause of sleeping sickness
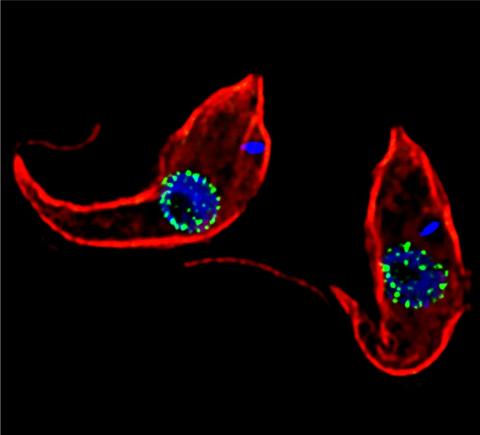
3765: Trypanosoma brucei, the cause of sleeping sickness
Trypanosoma brucei is a single-cell parasite that causes sleeping sickness in humans. Scientists have been studying trypanosomes for some time because of their negative effects on human and also animal health, especially in sub-Saharan Africa. Moreover, because these organisms evolved on a separate path from those of animals and plants more than a billion years ago, researchers study trypanosomes to find out what traits they may harbor that are common to or different from those of other eukaryotes (i.e., those organisms having a nucleus and mitochondria). This image shows the T. brucei cell membrane in red, the DNA in the nucleus and kinetoplast (a structure unique to protozoans, including trypanosomes, which contains mitochondrial DNA) in blue and nuclear pore complexes (which allow molecules to pass into or out of the nucleus) in green. Scientists have found that the trypanosome nuclear pore complex has a unique mechanism by which it attaches to the nuclear envelope. In addition, the trypanosome nuclear pore complex differs from those of other eukaryotes because its components have a near-complete symmetry, and it lacks almost all of the proteins that in other eukaryotes studied so far are required to assemble the pore.
Michael Rout, Rockefeller University
View Media
2367: Map of protein structures 02
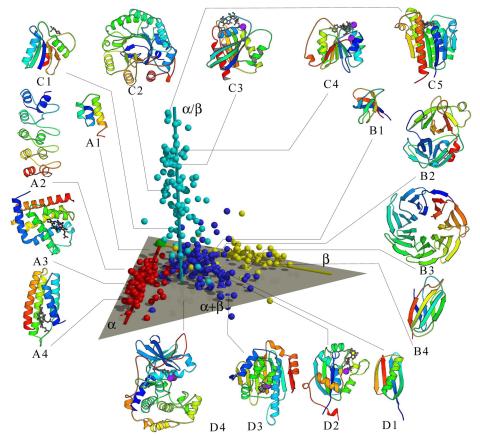
2367: Map of protein structures 02
A global "map of the protein structure universe" indicating the positions of specific proteins. The preponderance of small, less-structured proteins near the origin, with the more highly structured, large proteins towards the ends of the axes, may suggest the evolution of protein structures.
Berkeley Structural Genomics Center, PSI
View Media
6569: Cryo-electron tomography of a Caulobacter bacterium
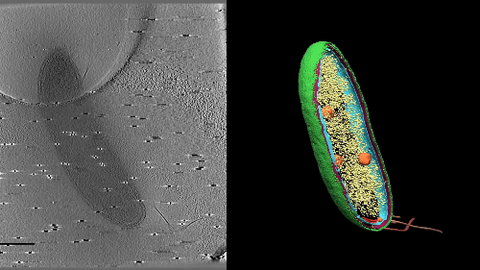
6569: Cryo-electron tomography of a Caulobacter bacterium
3D image of Caulobacter bacterium with various components highlighted: cell membranes (red and blue), protein shell (green), protein factories known as ribosomes (yellow), and storage granules (orange).
Peter Dahlberg, Stanford University.
View Media
2547: Central dogma, illustrated
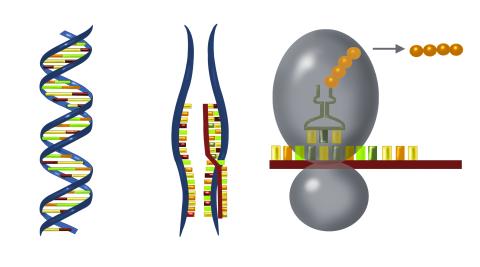
2547: Central dogma, illustrated
DNA encodes RNA, which encodes protein. DNA is transcribed to make messenger RNA (mRNA). The mRNA sequence (dark red strand) is complementary to the DNA sequence (blue strand). On ribosomes, transfer RNA (tRNA) reads three nucleotides at a time in mRNA to bring together the amino acids that link up to make a protein. See image 2548 for a labeled version of this illustration and 2549 for a labeled and numbered version. Featured in The New Genetics.
Crabtree + Company
View Media
3556: Bioluminescent imaging in adult zebrafish - lateral and overhead view
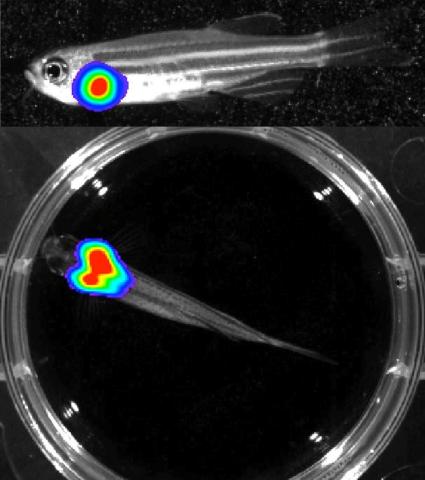
3556: Bioluminescent imaging in adult zebrafish - lateral and overhead view
Luciferase-based imaging enables visualization and quantification of internal organs and transplanted cells in live adult zebrafish. In this image, a cardiac muscle-restricted promoter drives firefly luciferase expression. This is the lateral and overhead (Bottom) view.
For imagery of the overhead view go to 3557.
For imagery of the lateral view go to 3558.
For more information about the illumated area go to 3559.
For imagery of the overhead view go to 3557.
For imagery of the lateral view go to 3558.
For more information about the illumated area go to 3559.
Kenneth Poss, Duke University
View Media

2443: Mapping human genetic variation
2443: Mapping human genetic variation
This map paints a colorful portrait of human genetic variation around the world. Researchers analyzed the DNA of 485 people and tinted the genetic types in different colors to produce one of the most detailed maps of its kind ever made. The map shows that genetic variation decreases with increasing distance from Africa, which supports the idea that humans originated in Africa, spread to the Middle East, then to Asia and Europe, and finally to the Americas. The data also offers a rich resource that scientists could use to pinpoint the genetic basis of diseases prevalent in diverse populations. Featured in the March 19, 2008, issue of Biomedical Beat.
Noah Rosenberg and Martin Soave, University of Michigan
View Media
2457: RAC1 activation in motile fibroblast
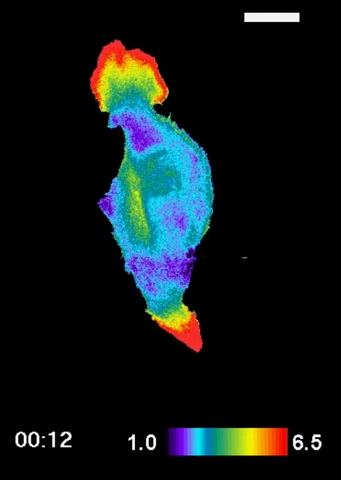
2457: RAC1 activation in motile fibroblast
Novel biosensor system maps the timing and location of Rac protein activation in a living mouse embryo fibroblast.
Klaus Hahn, University of North Carolina, Chapel Hill Medical School
View Media
3633: Cells lining the blood vessel walls
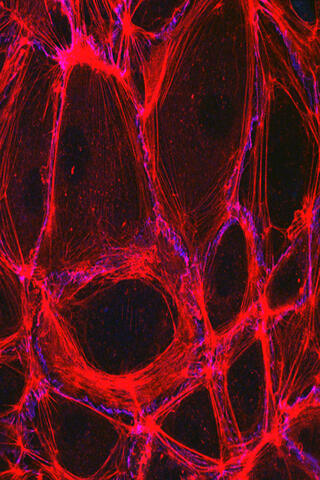
3633: Cells lining the blood vessel walls
The structure of the endothelium, the thin layer of cells that line our arteries and veins, is visible here. The endothelium is like a gatekeeper, controlling the movement of materials into and out of the bloodstream. Endothelial cells are held tightly together by specialized proteins that function like strong ropes (red) and others that act like cement (blue).
This image was part of the Life: Magnified exhibit that ran from June 3, 2014, to January 21, 2015, at Dulles International Airport.
This image was part of the Life: Magnified exhibit that ran from June 3, 2014, to January 21, 2015, at Dulles International Airport.
Christopher V. Carman and Roberta Martinelli, Harvard Medical School.
View Media
1021: Lily mitosis 08
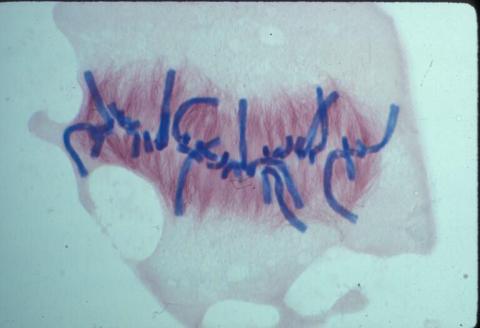
1021: Lily mitosis 08
A light microscope image of a cell from the endosperm of an African globe lily (Scadoxus katherinae). This is one frame of a time-lapse sequence that shows cell division in action. The lily is considered a good organism for studying cell division because its chromosomes are much thicker and easier to see than human ones. Staining shows microtubules in red and chromosomes in blue. Here, condensed chromosomes are clearly visible and lined up.
Related to images 1010, 1011, 1012, 1013, 1014, 1015, 1016, 1017, 1018, and 1019.
Related to images 1010, 1011, 1012, 1013, 1014, 1015, 1016, 1017, 1018, and 1019.
Andrew S. Bajer, University of Oregon, Eugene
View Media
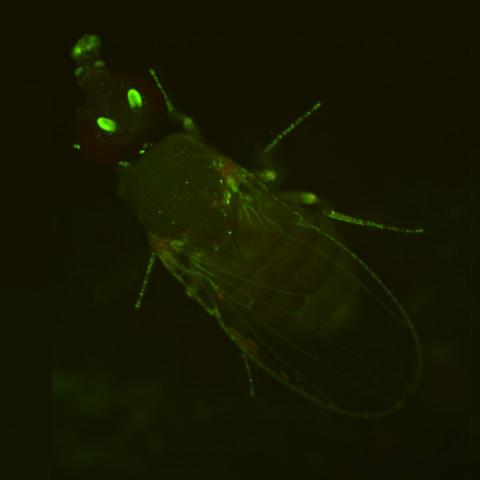

2417: Fly by night
2417: Fly by night
This fruit fly expresses green fluorescent protein (GFP) in the same pattern as the period gene, a gene that regulates circadian rhythm and is expressed in all sensory neurons on the surface of the fly.
Jay Hirsh, University of Virginia
View Media
5777: Microsporidia in roundworm 1
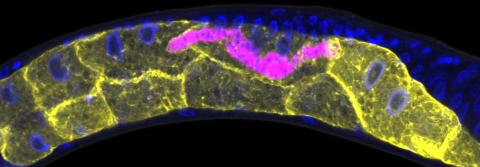
5777: Microsporidia in roundworm 1
Many disease-causing microbes manipulate their host’s metabolism and cells for their own ends. Microsporidia—which are parasites closely related to fungi—infect and multiply inside animal cells, and take the rearranging of cells’ interiors to a new level. They reprogram animal cells such that the cells start to fuse, causing them to form long, continuous tubes. As shown in this image of the roundworm Caenorhabditis elegans, microsporidia (shown in magenta) have invaded the worm’s gut cells (shown in yellow; the cells’ nuclei are shown in blue) and have instructed the cells to merge. The cell fusion enables the microsporidia to thrive and propagate in the expanded space. Scientists study microsporidia in worms to gain more insight into how these parasites manipulate their host cells. This knowledge might help researchers devise strategies to prevent or treat infections with microsporidia. For more on the research into microsporidia, see this news release from the University of California San Diego. Related to images 5778 and 5779.
Keir Balla and Emily Troemel, University of California San Diego
View Media
