Switch to List View
Image and Video Gallery
This is a searchable collection of scientific photos, illustrations, and videos. The images and videos in this gallery are licensed under Creative Commons Attribution Non-Commercial ShareAlike 3.0. This license lets you remix, tweak, and build upon this work non-commercially, as long as you credit and license your new creations under identical terms.
6967: Multinucleated cancer cell
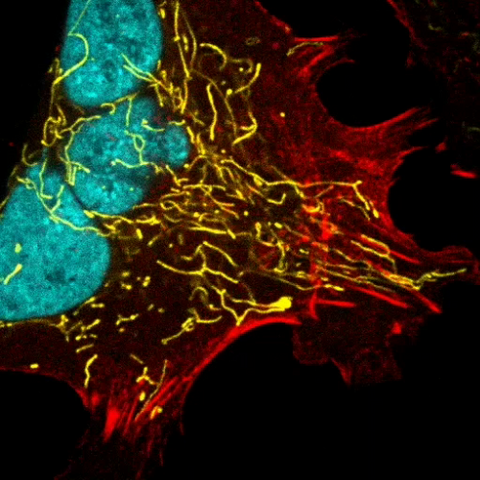
6967: Multinucleated cancer cell
A cancer cell with three nuclei, shown in turquoise. The abnormal number of nuclei indicates that the cell failed to go through cell division, probably more than once. Mitochondria are shown in yellow, and a protein of the cell’s cytoskeleton appears in red. This video was captured using a confocal microscope.
Dylan T. Burnette, Vanderbilt University School of Medicine.
View Media
3639: Cerebellum: the brain's locomotion control center
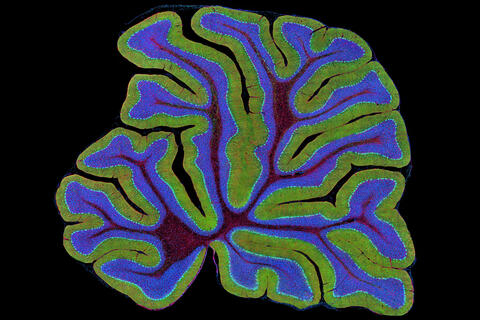
3639: Cerebellum: the brain's locomotion control center
The cerebellum of a mouse is shown here in cross-section. The cerebellum is the brain's locomotion control center. Every time you shoot a basketball, tie your shoe or chop an onion, your cerebellum fires into action. Found at the base of your brain, the cerebellum is a single layer of tissue with deep folds like an accordion. People with damage to this region of the brain often have difficulty with balance, coordination and fine motor skills. For a higher magnification, see image 3371.
This image was part of the Life: Magnified exhibit that ran from June 3, 2014, to January 21, 2015, at Dulles International Airport.
This image was part of the Life: Magnified exhibit that ran from June 3, 2014, to January 21, 2015, at Dulles International Airport.
Thomas Deerinck, National Center for Microscopy and Imaging Research, University of California, San Diego
View Media
2484: RNA Polymerase II
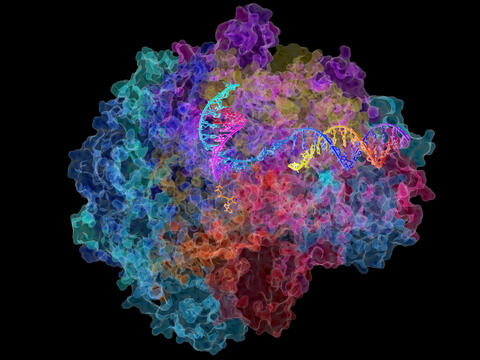
2484: RNA Polymerase II
NIGMS-funded researchers led by Roger Kornberg solved the structure of RNA polymerase II. This is the enzyme in mammalian cells that catalyzes the transcription of DNA into messenger RNA, the molecule that in turn dictates the order of amino acids in proteins. For his work on the mechanisms of mammalian transcription, Kornberg received the Nobel Prize in Chemistry in 2006.
David Bushnell, Ken Westover and Roger Kornberg, Stanford University
View Media
3440: Transcription factor Sox17 controls embryonic development of certain internal organs
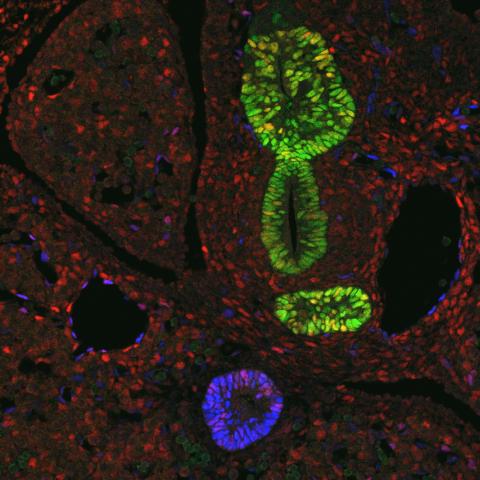
3440: Transcription factor Sox17 controls embryonic development of certain internal organs
During embryonic development, transcription factors (proteins that regulate gene expression) govern the differentiation of cells into separate tissues and organs. Researchers at Cincinnati Children's Hospital Medical Center used mice to study the development of certain internal organs, including the liver, pancreas, duodenum (beginning part of the small intestine), gall bladder and bile ducts. They discovered that transcription factor Sox17 guides some cells to develop into liver cells and others to become part of the pancreas or biliary system (gall bladder, bile ducts and associated structures). The separation of these two distinct cell types (liver versus pancreas/biliary system) is complete by embryonic day 8.5 in mice. The transcription factors PDX1 and Hes1 are also known to be involved in embryonic development of the pancreas and biliary system. This image shows mouse cells at embryonic day 10.5. The green areas show cells that will develop into the pancreas and/or duodenum(PDX1 is labeled green). The blue area near the bottom will become the gall bladder and the connecting tubes (common duct and cystic duct) that attach the gall bladder to the liver and pancreas (Sox17 is labeled blue). The transcription factor Hes1 is labeled red. The image was not published. A similar image (different plane of the section) was published in: Sox17 Regulates Organ Lineage Segregation of Ventral Foregut Progenitor Cells Jason R. Spence, Alex W. Lange, Suh-Chin J. Lin, Klaus H. Kaestner, Andrew M. Lowy, Injune Kim, Jeffrey A. Whitsett and James M. Wells, Developmental Cell, Volume 17, Issue 1, 62-74, 21 July 2009. doi:10.1016/j.devcel.2009.05.012
James M. Wells, Cincinnati Children's Hospital Medical Center
View Media
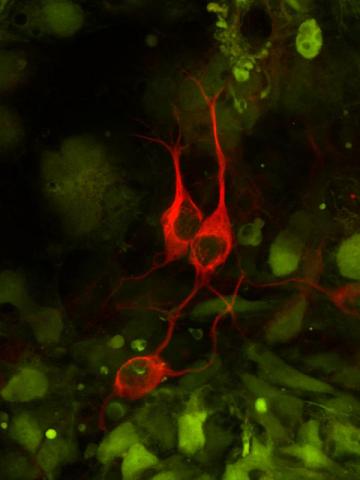
3290: Three neurons and human ES cells
3290: Three neurons and human ES cells
The three neurons (red) visible in this image were derived from human embryonic stem cells. Undifferentiated stem cells are green here. Image and caption information courtesy of the California Institute for Regenerative Medicine.
Anirvan Ghosh lab, University of California, San Diego, via CIRM
View Media

6751: Petri dish containing C. elegans
6751: Petri dish containing C. elegans
This Petri dish contains microscopic roundworms called Caenorhabditis elegans. Researchers used these particular worms to study how C. elegans senses the color of light in its environment.
H. Robert Horvitz and Dipon Ghosh, Massachusetts Institute of Technology.
View Media
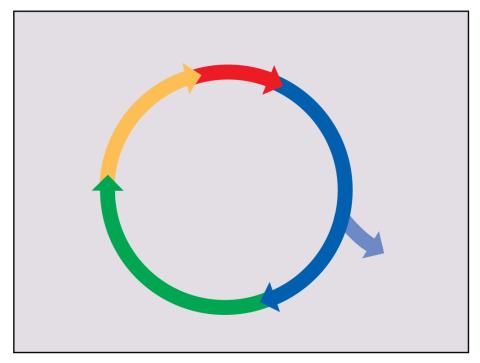
2498: Cell cycle
2498: Cell cycle
Cells progress through a cycle that consists of phases for growth (blue, green, yellow) and division (red). Cells become quiescent when they exit this cycle (purple). See image 2499 for a labeled version of this illustration.
Crabtree + Company
View Media
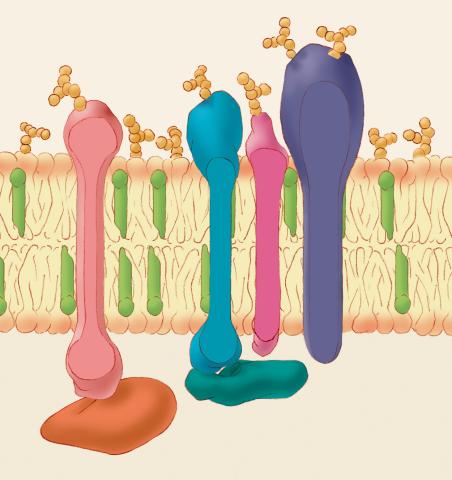
1286: Animal cell membrane
1286: Animal cell membrane
The membrane that surrounds a cell is made up of proteins and lipids. Depending on the membrane's location and role in the body, lipids can make up anywhere from 20 to 80 percent of the membrane, with the remainder being proteins. Cholesterol (green), which is not found in plant cells, is a type of lipid that helps stiffen the membrane.
Judith Stoffer
View Media
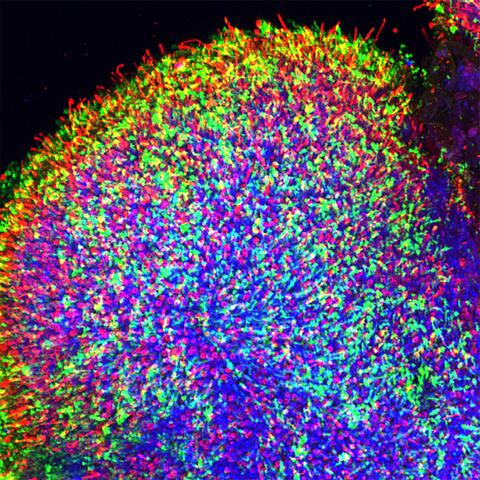
6748: Human retinal organoid
6748: Human retinal organoid
A replica of a human retina grown from stem cells. It shows rod photoreceptors (nerve cells responsible for dark vision) in green and red/green cones (nerve cells responsible for red and green color vision) in red. The cell nuclei are stained blue. This image was captured using a confocal microscope.
Kevin Eliceiri, University of Wisconsin-Madison.
View Media
6612: Ciclo circadiano de un adolescente típico
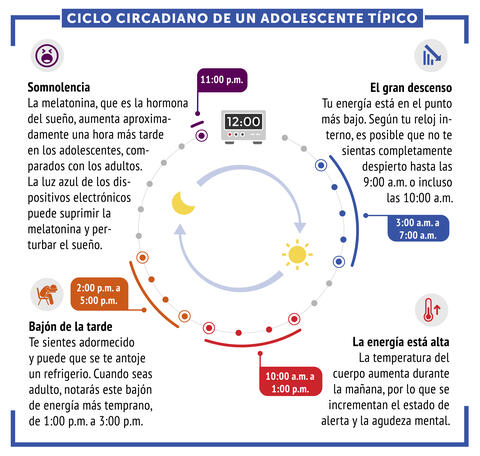
6612: Ciclo circadiano de un adolescente típico
Los ritmos circadianos son cambios físicos, mentales y conductuales que siguen un ciclo de 24 horas. Los ritmos circadianos típicos conducen a un nivel alto de energía durante la mitad del día (de 10 a.m. a 1 p.m.) y un bajón por la tarde. De noche, los ritmos circadianos hacen que la hormona melatonina aumente, lo que hace que la persona se sienta somnolienta.
Vea 6611 para la versión en inglés de esta infografía.
Vea 6611 para la versión en inglés de esta infografía.
NIGMS
View Media
3480: Cancer Cells Glowing from Luciferin
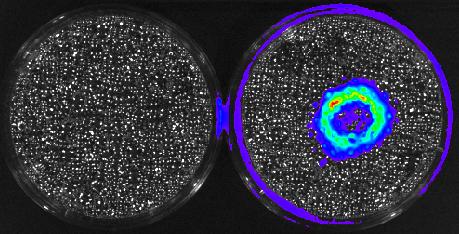
3480: Cancer Cells Glowing from Luciferin
The activator cancer cell culture, right, contains a chemical that causes the cells to emit light when in the presence of immune cells.
Mark Sellmyer, Stanford University School of Medicine
View Media

3492: Glowing bacteria make a pretty postcard
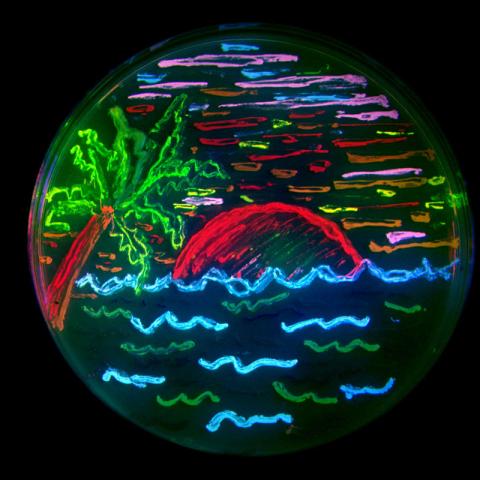
3492: Glowing bacteria make a pretty postcard
This tropical scene, reminiscent of a postcard from Key West, is actually a petri dish containing an artistic arrangement of genetically engineered bacteria. The image showcases eight of the fluorescent proteins created in the laboratory of the late Roger Y. Tsien, a cell biologist at the University of California, San Diego. Tsien, along with Osamu Shimomura of the Marine Biology Laboratory and Martin Chalfie of Columbia University, share the 2008 Nobel Prize in chemistry for their work on green fluorescent protein-a naturally glowing molecule from jellyfish that has become a powerful tool for studying molecules inside living cells.
Nathan C. Shaner, The Scintillon Institute
View Media
3288: Smooth muscle from human ES cells
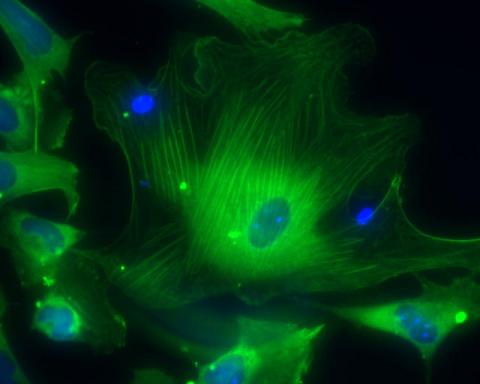
3288: Smooth muscle from human ES cells
These smooth muscle cells were derived from human embryonic stem cells. The nuclei are stained blue, and the proteins of the cytoskeleton are stained green. Image and caption information courtesy of the California Institute for Regenerative Medicine.
Alexey Terskikh lab, Burnham Institute for Medical Research, via CIRM
View Media
2603: Induced stem cells from adult skin 01
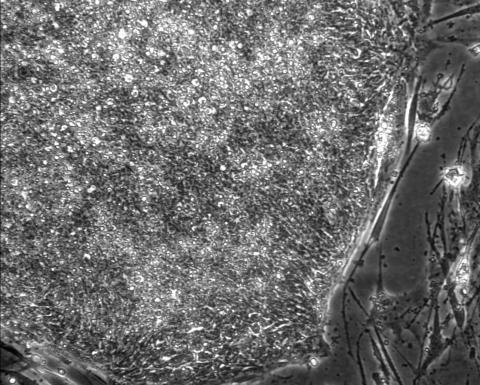
2603: Induced stem cells from adult skin 01
These cells are induced stem cells made from human adult skin cells that were genetically reprogrammed to mimic embryonic stem cells. The induced stem cells were made potentially safer by removing the introduced genes and the viral vector used to ferry genes into the cells, a loop of DNA called a plasmid. The work was accomplished by geneticist Junying Yu in the laboratory of James Thomson, a University of Wisconsin-Madison School of Medicine and Public Health professor and the director of regenerative biology for the Morgridge Institute for Research.
James Thomson, University of Wisconsin-Madison
View Media
1291: Olfactory system
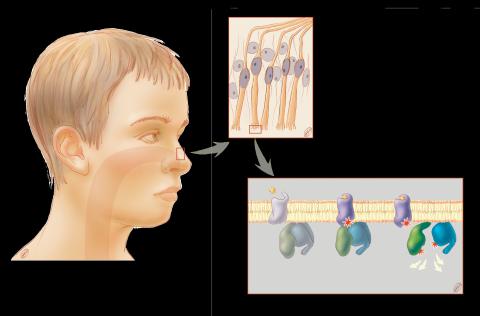
1291: Olfactory system
Sensory organs have cells equipped for detecting signals from the environment, such as odors. Receptors in the membranes of nerve cells in the nose bind to odor molecules, triggering a cascade of chemical reactions tranferred by G proteins into the cytoplasm.
Judith Stoffer
View Media
5779: Microsporidia in roundworm 3
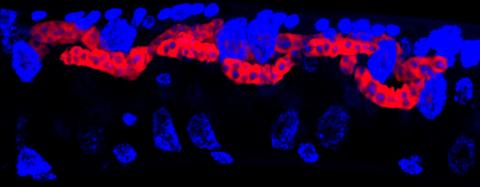
5779: Microsporidia in roundworm 3
Many disease-causing microbes manipulate their host’s metabolism and cells for their own ends. Microsporidia—which are parasites closely related to fungi—infect and multiply inside animal cells, and take the rearranging of cells’ interiors to a new level. They reprogram animal cells such that the cells start to fuse, causing them to form long, continuous tubes. As shown in this image of the roundworm Caenorhabditis elegans, microsporidia (shown in red) have invaded the worm’s gut cells (the large blue dots are the cells' nuclei) and have instructed the cells to merge. The cell fusion enables the microsporidia to thrive and propagate in the expanded space. Scientists study microsporidia in worms to gain more insight into how these parasites manipulate their host cells. This knowledge might help researchers devise strategies to prevent or treat infections with microsporidia.
For more on the research into microsporidia, see this news release from the University of California San Diego. Related to images 5777 and 5778.
For more on the research into microsporidia, see this news release from the University of California San Diego. Related to images 5777 and 5778.
Keir Balla and Emily Troemel, University of California San Diego
View Media
6930: Mouse brain 2
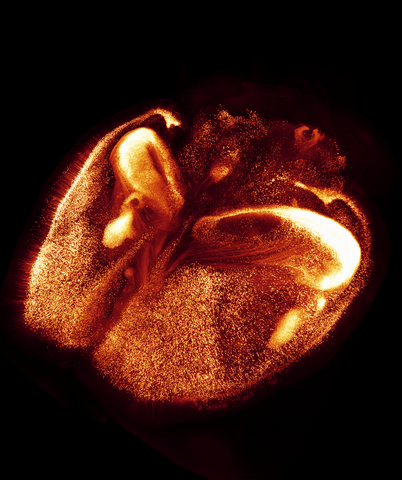
6930: Mouse brain 2
A mouse brain that was genetically modified so that subpopulations of its neurons glow. Researchers often study mice because they share many genes with people and can shed light on biological processes, development, and diseases in humans.
This image was captured using a light sheet microscope.
Related to image 6929 and video 6931.
This image was captured using a light sheet microscope.
Related to image 6929 and video 6931.
Prayag Murawala, MDI Biological Laboratory and Hannover Medical School.
View Media
6661: Zebrafish embryo showing vasculature
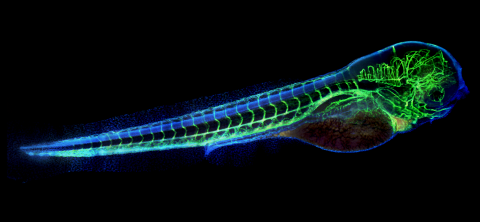
6661: Zebrafish embryo showing vasculature
A zebrafish embryo. The blue areas are cell bodies, the green lines are blood vessels, and the red glow is blood. This image was created by stitching together five individual images captured with a hyperspectral multipoint confocal fluorescence microscope that was developed at the Eliceiri Lab.
Kevin Eliceiri, University of Wisconsin-Madison.
View Media
3334: Four timepoints in gastrulation
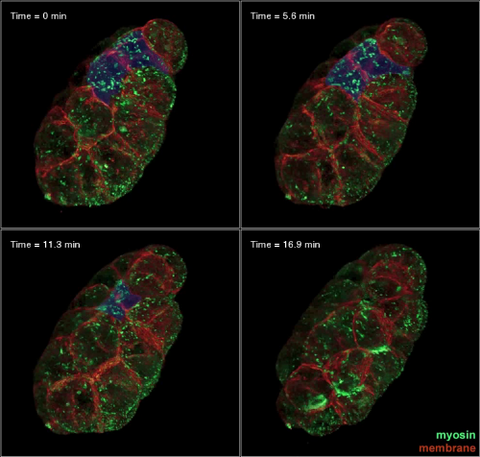
3334: Four timepoints in gastrulation
It has been said that gastrulation is the most important event in a person's life. This part of early embryonic development transforms a simple ball of cells and begins to define cell fate and the body axis. In a study published in Science magazine, NIGMS grantee Bob Goldstein and his research group studied how contractions of actomyosin filaments in C. elegans and Drosophila embryos lead to dramatic rearrangements of cell and embryonic structure. In these images, myosin (green) and plasma membrane (red) are highlighted at four timepoints in gastrulation in the roundworm C. elegans. The blue highlights in the top three frames show how cells are internalized, and the site of closure around the involuting cells is marked with an arrow in the last frame. See related image 3297.
Bob Goldstein, University of North Carolina, Chapel Hill
View Media
3771: Molecular model of freshly made Rous sarcoma virus (RSV)
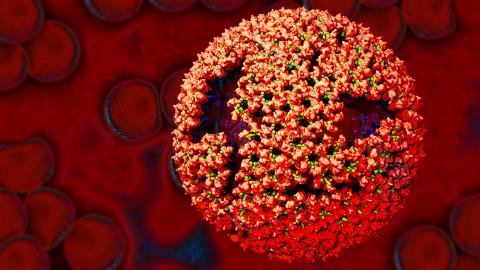
3771: Molecular model of freshly made Rous sarcoma virus (RSV)
Viruses have been the foes of animals and other organisms for time immemorial. For almost as long, they've stayed well hidden from view because they are so tiny (they aren't even cells, so scientists call the individual virus a "particle"). This image shows a molecular model of a particle of the Rous sarcoma virus (RSV), a virus that infects and sometimes causes cancer in chickens. In the background is a photo of red blood cells. The particle shown is "immature" (not yet capable of infecting new cells) because it has just budded from an infected chicken cell and entered the bird's bloodstream. The outer shell of the immature virus is made up of a regular assembly of large proteins (shown in red) that are linked together with short protein molecules called peptides (green). This outer shell covers and protects the proteins (blue) that form the inner shell of the particle. But as you can see, the protective armor of the immature virus contains gaping holes. As the particle matures, the short peptides are removed and the large proteins rearrange, fusing together into a solid sphere capable of infecting new cells. While still immature, the particle is vulnerable to drugs that block its development. Knowing the structure of the immature particle may help scientists develop better medications against RSV and similar viruses in humans. Scientists used sophisticated computational tools to reconstruct the RSV atomic structure by crunching various data on the RSV proteins to simulate the entire structure of immature RSV.
Boon Chong Goh, University of Illinois at Urbana-Champaign
View Media
3609: Pollen grains: male germ cells in plants and a cause of seasonal allergies
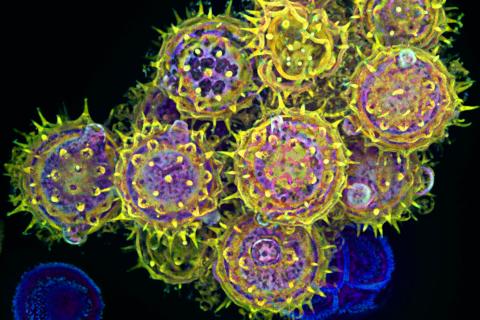
3609: Pollen grains: male germ cells in plants and a cause of seasonal allergies
Those of us who get sneezy and itchy-eyed every spring or fall may have pollen grains, like those shown here, to blame. Pollen grains are the male germ cells of plants, released to fertilize the corresponding female plant parts. When they are instead inhaled into human nasal passages, they can trigger allergies.
This image was part of the Life: Magnified exhibit that ran from June 3, 2014, to January 21, 2015, at Dulles International Airport.
This image was part of the Life: Magnified exhibit that ran from June 3, 2014, to January 21, 2015, at Dulles International Airport.
Edna, Gil, and Amit Cukierman, Fox Chase Cancer Center, Philadelphia, Pa.
View Media

5810: Tongue 1
5810: Tongue 1
Microscopy image of tongue. One in a series of two, see image 5811
National Center for Microscopy and Imaging Research (NCMIR)
View Media
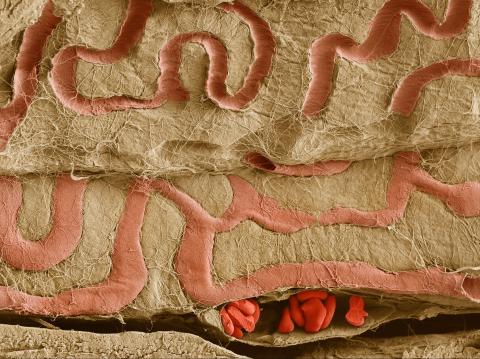
3739: Scanning electron microscopy of the ECM on the surface of a calf muscle
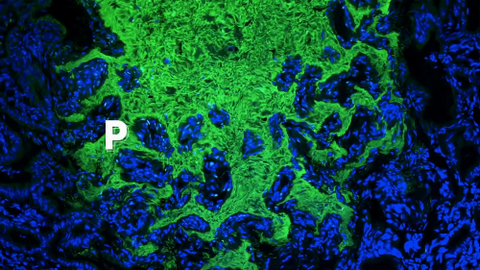
3739: Scanning electron microscopy of the ECM on the surface of a calf muscle
This image shows the extracellular matrix (ECM) on the surface of a soleus (lower calf) muscle in light brown and blood vessels in pink. Near the bottom of the photo, a vessel is opened up to reveal red blood cells. Scientists know less about the ECM in muscle than in other tissues, but it's increasingly clear that the ECM is critical to muscle function, and disruption of the ECM has been associated with many muscle disorders. The ECM in muscles stores and releases growth factors, suggesting that it might play a role in cellular communication.
Tom Deerinck, National Center for Microscopy and Imaging Research (NCMIR)
View Media
2578: Cellular aging
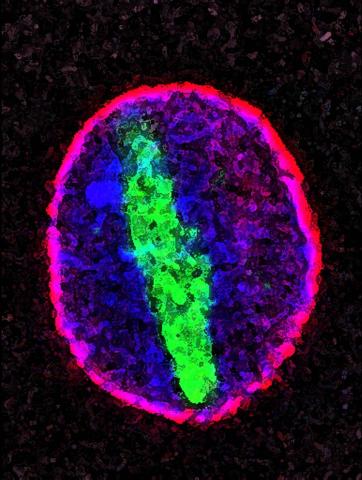
2578: Cellular aging
A protein called tubulin (green) accumulates in the center of a nucleus (outlined in pink) from an aging cell. Normally, this protein is kept out of the nucleus with the help of gatekeepers known as nuclear pore complexes. But NIGMS-funded researchers found that wear and tear to long-lived components of the complexes eventually lowers the gatekeepers' guard. As a result, cytoplasmic proteins like tubulin gain entry to the nucleus while proteins normally confined to the nucleus seep out. The work suggests that finding ways to stop the leakage could slow the cellular aging process and possibly lead to new therapies for age-related diseases.
Maximiliano D'Angelo and Martin Hetzer, Salk Institute
View Media
3253: Pulsating response to stress in bacteria
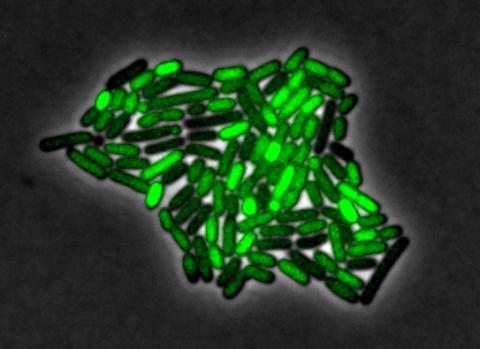
3253: Pulsating response to stress in bacteria
By attaching fluorescent proteins to the genetic circuit responsible for B. subtilis's stress response, researchers can observe the cells' pulses as green flashes. In response to a stressful environment like one lacking food, B. subtilis activates a large set of genes that help it respond to the hardship. Instead of leaving those genes on as previously thought, researchers discovered that the bacteria flip the genes on and off, increasing the frequency of these pulses with increasing stress. See entry 3254 for the related video.
Michael Elowitz, Caltech University
View Media
6614: Los ritmos circadianos y el núcleo supraquiasmático
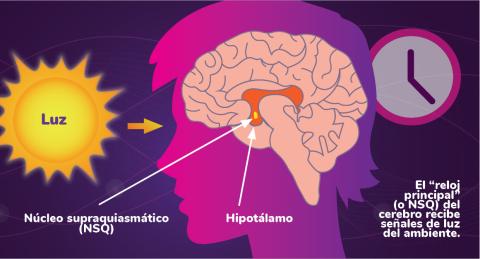
6614: Los ritmos circadianos y el núcleo supraquiasmático
Los ritmos circadianos son cambios físicos, mentales y de comportamiento que siguen un ciclo de 24 horas. Los ritmos circadianos se ven influenciados por la luz y están regulados por el núcleo supraquiasmático del cerebro, a veces denominado el reloj principal.
Vea 6613 para la versión en inglés de esta infografía.
Vea 6613 para la versión en inglés de esta infografía.
NIGMS
View Media
2746: Active site of sulfite oxidase
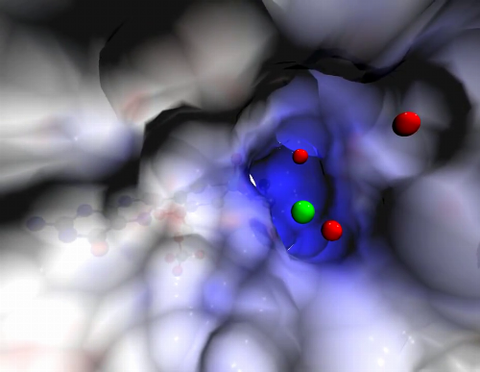
2746: Active site of sulfite oxidase
Sulfite oxidase is an enzyme that is essential for normal neurological development in children. This video shows the active site of the enzyme and its molybdenum cofactor visible as a faint ball-and-stick representation buried within the protein. The positively charged channel (blue) at the active site contains a chloride ion (green) and three water molecules (red). As the protein oscillates, one can see directly down the positively charged channel. At the bottom is the molybdenum atom of the active site (light blue) and its oxo group (red) that is transferred to sulfite to form sulfate in the catalytic reaction.
John Enemark, University of Arizona
View Media
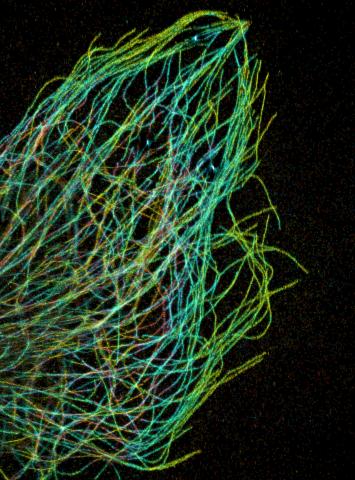
3611: Tiny strands of tubulin, a protein in a cell's skeleton
3611: Tiny strands of tubulin, a protein in a cell's skeleton
Just as our bodies rely on bones for structural support, our cells rely on a cellular skeleton. In addition to helping cells keep their shape, this cytoskeleton transports material within cells and coordinates cell division. One component of the cytoskeleton is a protein called tubulin, shown here as thin strands.
This image was part of the Life: Magnified exhibit that ran from June 3, 2014, to January 21, 2015, at Dulles International Airport.
This image was part of the Life: Magnified exhibit that ran from June 3, 2014, to January 21, 2015, at Dulles International Airport.
Pakorn Kanchanawong, National University of Singapore and National Heart, Lung, and Blood Institute, National Institutes of Health; and Clare Waterman, National Heart, Lung, and Blood Institute, National Institutes of Health
View Media
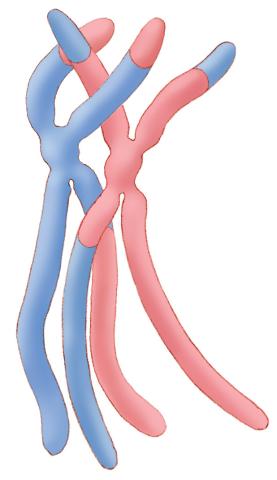
1314: Chromosomes after crossing over
1314: Chromosomes after crossing over
Duplicated pair of chromosomes have exchanged material.
Judith Stoffer
View Media
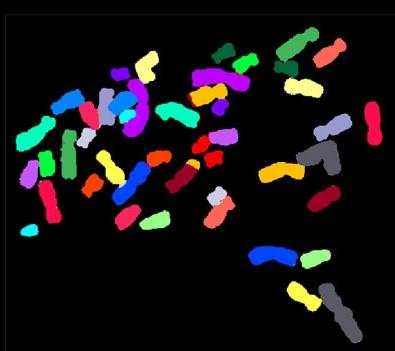
2312: Color-coded chromosomes
2312: Color-coded chromosomes
By mixing fluorescent dyes like an artist mixes paints, scientists are able to color code individual chromosomes. The technique, abbreviated multicolor-FISH, allows researchers to visualize genetic abnormalities often linked to disease. In this image, "painted" chromosomes from a person with a hereditary disease called Werner Syndrome show where a piece of one chromosome has fused to another (see the gold-tipped maroon chromosome in the center). As reported by molecular biologist Jan Karlseder of the Salk Institute for Biological Studies, such damage is typical among people with this rare syndrome.
Anna Jauch, Institute of Human Genetics, Heidelberg, Germany
View Media
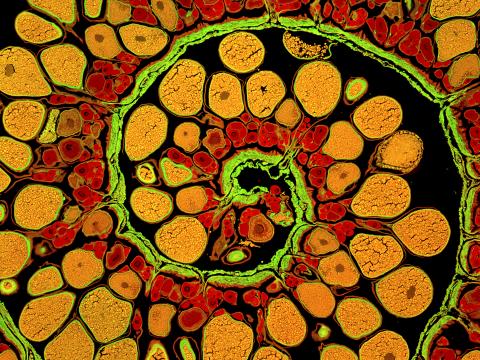
3620: Anglerfish ovary cross-section
3620: Anglerfish ovary cross-section
This image captures the spiral-shaped ovary of an anglerfish in cross-section. Once matured, these eggs will be released in a gelatinous, floating mass. For some species of anglerfish, this egg mass can be up to 3 feet long and include nearly 200,000 eggs.
This image was part of the Life: Magnified exhibit that ran from June 3, 2014, to January 21, 2015, at Dulles International Airport.
This image was part of the Life: Magnified exhibit that ran from June 3, 2014, to January 21, 2015, at Dulles International Airport.
James E. Hayden, The Wistar Institute, Philadelphia, Pa.
View Media
3788: Yeast cells pack a punch
3788: Yeast cells pack a punch
Although they are tiny, microbes that are growing in confined spaces can generate a lot of pressure. In this video, yeast cells grow in a small chamber called a microfluidic bioreactor. As the cells multiply, they begin to bump into and squeeze each other, resulting in periodic bursts of cells moving into different parts of the chamber. The continually growing cells also generate a lot of pressure--the researchers conducting these experiments found that the pressure generated by the cells can be almost five times higher than that in a car tire--about 150 psi, or 10 times the atmospheric pressure. Occasionally, this pressure even caused the small reactor to burst. By tracking the growth of the yeast or other cells and measuring the mechanical forces generated, scientists can simulate microbial growth in various places such as water pumps, sewage lines or catheters to learn how damage to these devices can be prevented. To learn more how researchers used small bioreactors to gauge the pressure generated by growing microbes, see this press release from UC Berkeley.
Oskar Hallatschek, UC Berkeley
View Media
6539: Pathways: What is Basic Science?
6539: Pathways: What is Basic Science?
Learn about basic science, sometimes called “pure” or “fundamental” science, and how it contributes to the development of medical treatments. Discover more resources from NIGMS’ Pathways collaboration with Scholastic. View the video on YouTube for closed captioning.
National Institute of General Medical Sciences
View Media
2513: Life of an AIDS virus
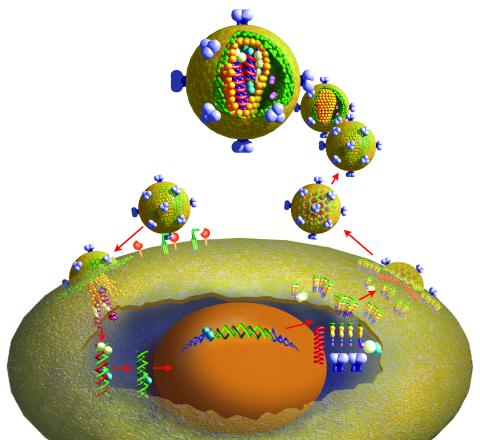
2513: Life of an AIDS virus
HIV is a retrovirus, a type of virus that carries its genetic material not as DNA but as RNA. Long before anyone had heard of HIV, researchers in labs all over the world studied retroviruses, tracing out their life cycle and identifying the key proteins the viruses use to infect cells. When HIV was identified as a retrovirus, these studies gave AIDS researchers an immediate jump-start. The previously identified viral proteins became initial drug targets. See images 2514 and 2515 for labeled versions of this illustration. Featured in The Structures of Life.
Crabtree + Company
View Media
1058: Lily mitosis 01
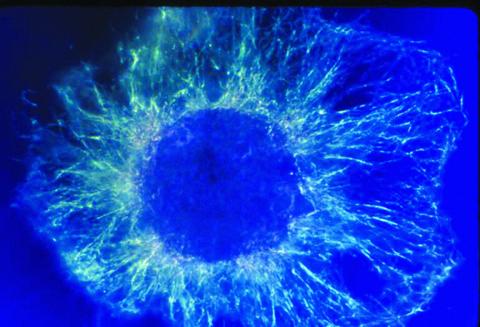
1058: Lily mitosis 01
A light microscope image shows the chromosomes, stained dark blue, in a dividing cell of an African globe lily (Scadoxus katherinae). This is one frame of a time-lapse sequence that shows cell division in action. The lily is considered a good organism for studying cell division because its chromosomes are much thicker and easier to see than human ones.
Andrew S. Bajer, University of Oregon, Eugene
View Media

2387: Thymidylate synthase complementing protein from Thermotoga maritime
2387: Thymidylate synthase complementing protein from Thermotoga maritime
A model of thymidylate synthase complementing protein from Thermotoga maritime.
Joint Center for Structural Genomics, PSI
View Media
6577: Transient receptor potential channel TRPV5
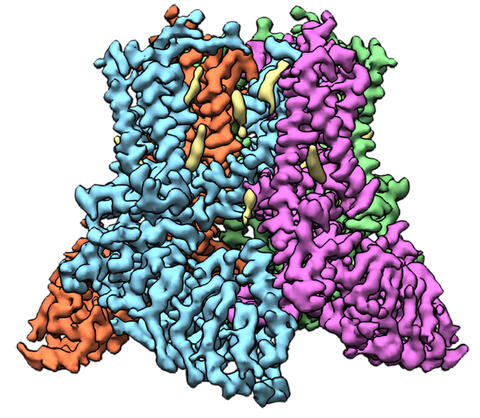
6577: Transient receptor potential channel TRPV5
A 3D reconstruction of a transient receptor potential channel called TRPV5 that was created based on cryo-electron microscopy images. TRPV5 is primarily found in kidney cells and is essential for reabsorbing calcium into the blood.
Vera Moiseenkova-Bell, University of Pennsylvania.
View Media
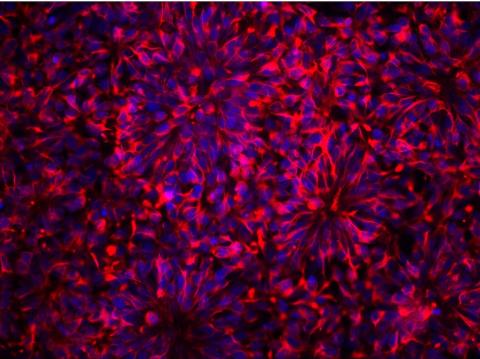
3284: Neurons from human ES cells
3284: Neurons from human ES cells
These neural precursor cells were derived from human embryonic stem cells. The neural cell bodies are stained red, and the nuclei are blue. Image and caption information courtesy of the California Institute for Regenerative Medicine.
Xianmin Zeng lab, Buck Institute for Age Research, via CIRM
View Media
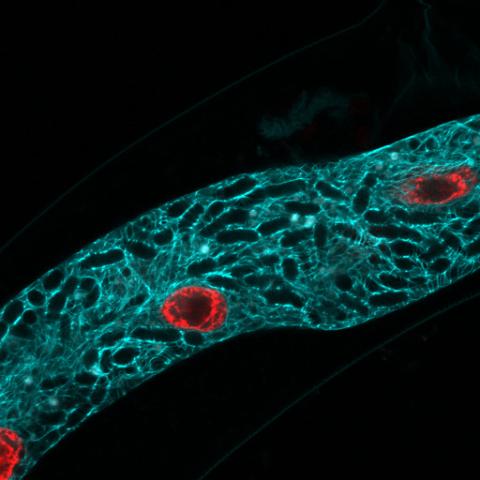
5778: Microsporidia in roundworm 2
5778: Microsporidia in roundworm 2
Many disease-causing microbes manipulate their host’s metabolism and cells for their own ends. Microsporidia—which are parasites closely related to fungi—infect and multiply inside animal cells, and take the rearranging of cells’ interiors to a new level. They reprogram animal cells such that the cells start to fuse, causing them to form long, continuous tubes. As shown in this image of the roundworm Caenorhabditis elegans, microsporidia (dark oval shapes) invaded the worm’s gut cells (long tube; the cell nuclei are shown in red) and have instructed the cells to merge. The cell fusion enables the microsporidia to thrive and propagate in the expanded space. Scientists study microsporidia in worms to gain more insight into how these parasites manipulate their host cells. This knowledge might help researchers devise strategies to prevent or treat infections with microsporidia.
For more on the research into microsporidia, see this news release from the University of California San Diego. Related to images 5777 and 5779.
For more on the research into microsporidia, see this news release from the University of California San Diego. Related to images 5777 and 5779.
Keir Balla and Emily Troemel, University of California San Diego
View Media

6607: Cryo-ET cell cross-section visualizing insulin vesicles
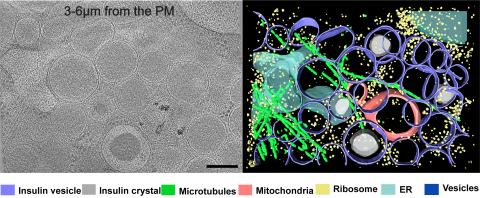
6607: Cryo-ET cell cross-section visualizing insulin vesicles
On the left, a cross-section slice of a rat pancreas cell captured using cryo-electron tomography (cryo-ET). On the right, a color-coded, 3D version of the image highlighting cell structures. Visible features include insulin vesicles (purple rings), insulin crystals (gray circles), microtubules (green rods), ribosomes (small yellow circles). The black line at the bottom right of the left image represents 200 nm. Related to image 6608.
Xianjun Zhang, University of Southern California.
View Media
6556: Floral pattern in a mixture of two bacterial species, Acinetobacter baylyi and Escherichia coli, grown on a semi-solid agar for 72 hour
6556: Floral pattern in a mixture of two bacterial species, Acinetobacter baylyi and Escherichia coli, grown on a semi-solid agar for 72 hour
Floral pattern emerging as two bacterial species, motile Acinetobacter baylyi and non-motile Escherichia coli (green), are grown together for 72 hours on 0.5% agar surface from a small inoculum in the center of a Petri dish.
See 6557 for a photo of this process at 24 hours on 0.75% agar surface.
See 6553 for a photo of this process at 48 hours on 1% agar surface.
See 6555 for another photo of this process at 48 hours on 1% agar surface.
See 6550 for a video of this process.
See 6557 for a photo of this process at 24 hours on 0.75% agar surface.
See 6553 for a photo of this process at 48 hours on 1% agar surface.
See 6555 for another photo of this process at 48 hours on 1% agar surface.
See 6550 for a video of this process.
L. Xiong et al, eLife 2020;9: e48885
View Media
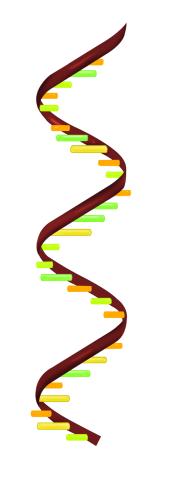
2554: RNA strand
2554: RNA strand
Ribonucleic acid (RNA) has a sugar-phosphate backbone and the bases adenine (A), cytosine (C), guanine (G), and uracil (U). See image 2555 for a labeled version of this illustration. Featured in The New Genetics.
Crabtree + Company
View Media

7011: Hawaiian bobtail squid
7011: Hawaiian bobtail squid
An adult Hawaiian bobtail squid, Euprymna scolopes, swimming next to a submerged hand.
Related to image 7010 and video 7012.
Related to image 7010 and video 7012.
Margaret J. McFall-Ngai, Carnegie Institution for Science/California Institute of Technology, and Edward G. Ruby, California Institute of Technology.
View Media
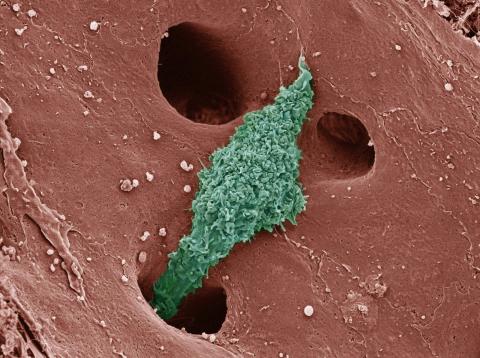
6535: Kupffer cell residing in the liver
6535: Kupffer cell residing in the liver
Kupffer cells appear in the liver during the early stages of mammalian development and stay put throughout life to protect liver cells, clean up old red blood cells, and regulate iron levels. Source article Replenishing the Liver’s Immune Protections. Posted on December 12th, 2019 by Dr. Francis Collins.
Thomas Deerinck, National Center for Microscopy and Imaging Research, University of California, San Diego.
View Media

2397: Bovine milk alpha-lactalbumin (1)
2397: Bovine milk alpha-lactalbumin (1)
A crystal of bovine milk alpha-lactalbumin protein created for X-ray crystallography, which can reveal detailed, three-dimensional protein structures.
Alex McPherson, University of California, Irvine
View Media

1294: Stem cell differentiation
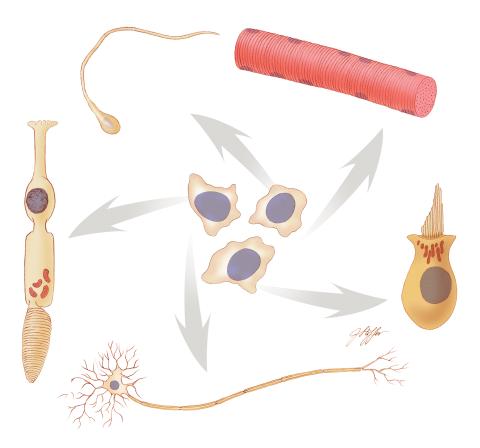
1294: Stem cell differentiation
Undifferentiated embryonic stem cells cease to exist a few days after conception. In this image, ES cells are shown to differentiate into sperm, muscle fiber, hair cells, nerve cells, and cone cells.
Judith Stoffer
View Media
2756: Xenopus laevis embryos
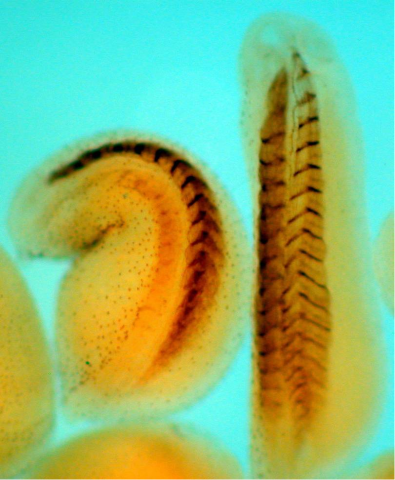
2756: Xenopus laevis embryos
Xenopus laevis, the African clawed frog, has long been used as a model organism for studying embryonic development. The frog embryo on the left lacks the developmental factor Sizzled. A normal embryo is shown on the right.
Michael Klymkowsky, University of Colorado, Boulder
View Media
3558: Bioluminescent imaging in adult zebrafish - lateral view
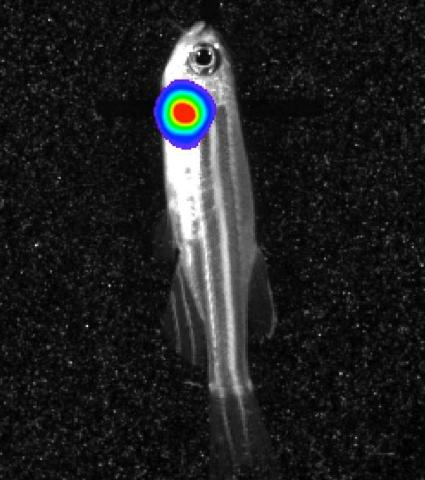
3558: Bioluminescent imaging in adult zebrafish - lateral view
Luciferase-based imaging enables visualization and quantification of internal organs and transplanted cells in live adult zebrafish. In this image, a cardiac muscle-restricted promoter drives firefly luciferase expression (lateral view).
For imagery of both the lateral and overhead view go to 3556.
For imagery of the overhead view go to 3557.
For more information about the illumated area go to 3559.
For imagery of both the lateral and overhead view go to 3556.
For imagery of the overhead view go to 3557.
For more information about the illumated area go to 3559.
Kenneth Poss, Duke University
View Media
6969: Snowflake yeast 1
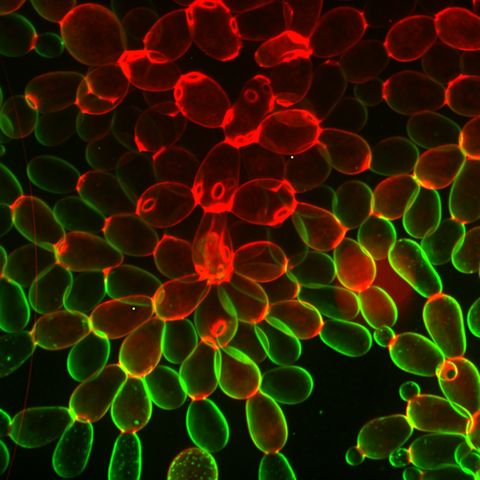
6969: Snowflake yeast 1
Multicellular yeast called snowflake yeast that researchers created through many generations of directed evolution from unicellular yeast. Stained cell membranes (green) and cell walls (red) reveal the connections between cells. Younger cells take up more cell membrane stain, while older cells take up more cell wall stain, leading to the color differences seen here. This image was captured using spinning disk confocal microscopy.
Related to images 6970 and 6971.
Related to images 6970 and 6971.
William Ratcliff, Georgia Institute of Technology.
View Media

