Switch to List View
Image and Video Gallery
This is a searchable collection of scientific photos, illustrations, and videos. The images and videos in this gallery are licensed under Creative Commons Attribution Non-Commercial ShareAlike 3.0. This license lets you remix, tweak, and build upon this work non-commercially, as long as you credit and license your new creations under identical terms.
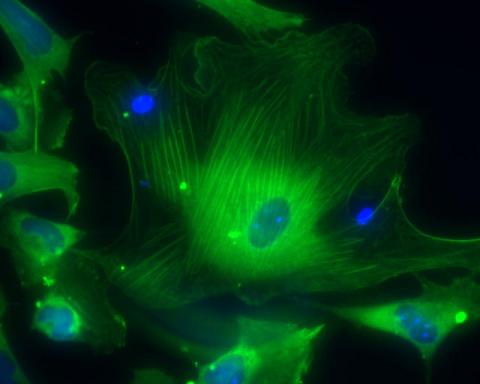
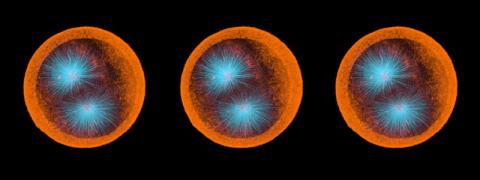
3288: Smooth muscle from human ES cells
3288: Smooth muscle from human ES cells
These smooth muscle cells were derived from human embryonic stem cells. The nuclei are stained blue, and the proteins of the cytoskeleton are stained green. Image and caption information courtesy of the California Institute for Regenerative Medicine.
Alexey Terskikh lab, Burnham Institute for Medical Research, via CIRM
View Media
3268: Fluorescent E. coli bacteria
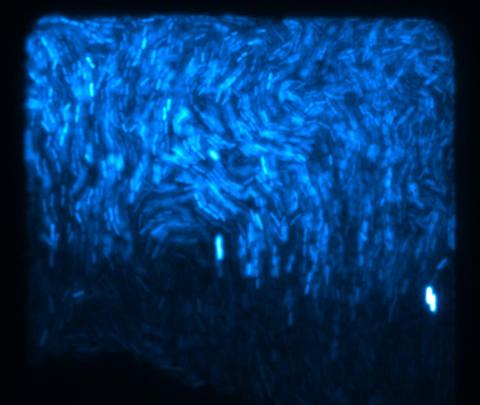
3268: Fluorescent E. coli bacteria
Bioengineers were able to coax bacteria to blink in unison on microfluidic chips. They called each blinking bacterial colony a biopixel. Thousands of fluorescent E. coli bacteria, shown here, make up a biopixel. Related to images 3265 and 3266. From a UC San Diego news release, "Researchers create living 'neon signs' composed of millions of glowing bacteria."
Jeff Hasty Lab, UC San Diego
View Media

2749: Cytoscape network wiring diagram 2
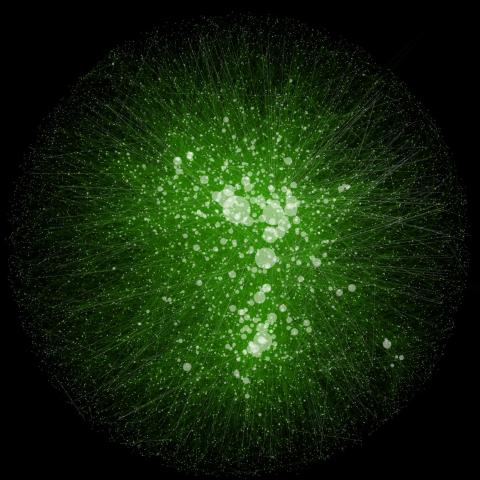
2749: Cytoscape network wiring diagram 2
This image integrates the thousands of known molecular and genetic interactions happening inside our bodies using a computer program called Cytoscape. Images like this are known as network wiring diagrams, but Cytoscape creator Trey Ideker somewhat jokingly calls them "hairballs" because they can be so complicated, intricate and hard to tease apart. Cytoscape comes with tools to help scientists study specific interactions, such as differences between species or between sick and diseased cells. Related to 2737.
Trey Ideker, University of California, San Diego
View Media
3727: Zinc levels in a plant leaf
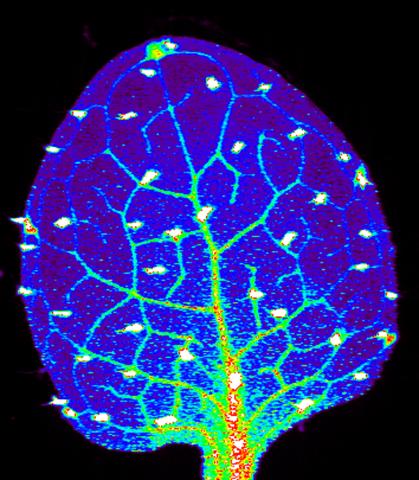
3727: Zinc levels in a plant leaf
Zinc is required for the function of more than 300 enzymes, including those that help regulate gene expression, in various organisms including humans. Researchers study how plants acquire, sequester and distribute zinc to find ways to increase the zinc content of crops to improve human health. Using synchrotron X-ray fluorescence technology, they created this heat map of zinc levels in an Arabidopsis thaliana plant leaf. This image is a winner of the 2015 FASEB Bioart contest and was featured in the NIH Director's blog.
Suzana Car, Dartmouth College
View Media
2753: Xenopus laevis egg
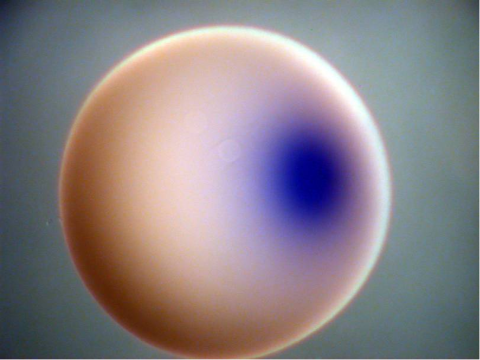
2753: Xenopus laevis egg
Xenopus laevis, the African clawed frog, has long been used as a model organism for studying embryonic development. In this image, RNA encoding the transcription factor Sox 7 (dark blue) is shown to predominate at the vegetal pole, the yolk-rich portion, of a Xenopus laevis frog egg. Sox 7 protein is important to the regulation of embryonic development.
Michael Klymkowsky, University of Colorado, Boulder
View Media
2317: Fruitful dyes
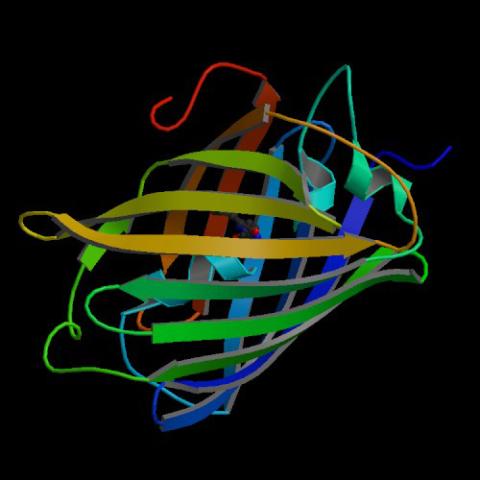
2317: Fruitful dyes
These colorful, computer-generated ribbons show the backbone of a molecule that glows a fluorescent red. The molecule, called mStrawberry, was created by chemists based on a protein found in the ruddy lips of a coral. Scientists use the synthetic molecule and other "fruity" ones like it as a dye to mark and study cell structures.
Roger Y. Tsien, University of California, San Diego
View Media
5882: Beta-galactosidase montage showing cryo-EM improvement--transparent background
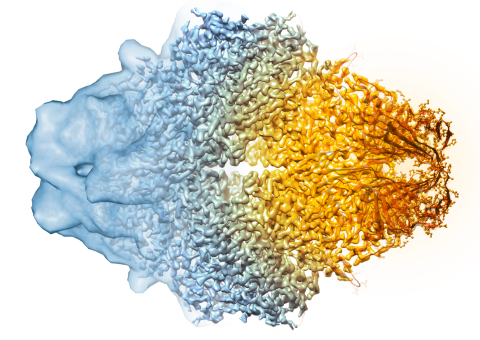
5882: Beta-galactosidase montage showing cryo-EM improvement--transparent background
Composite image of beta-galactosidase showing how cryo-EM’s resolution has improved dramatically in recent years. Older images to the left, more recent to the right. Related to image 5883. NIH Director Francis Collins featured this on his blog on January 14, 2016.
Veronica Falconieri, Sriram Subramaniam Lab, National Cancer Institute
View Media
2426: Zinc finger
2426: Zinc finger
The structure of a gene-regulating zinc finger protein bound to DNA.
Jeremy M. Berg, National Institute of General Medical Sciences
View Media
7004: Protein kinases as cancer chemotherapy targets
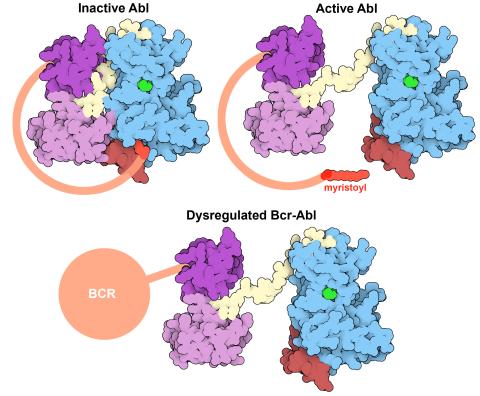
7004: Protein kinases as cancer chemotherapy targets
Protein kinases—enzymes that add phosphate groups to molecules—are cancer chemotherapy targets because they play significant roles in almost all aspects of cell function, are tightly regulated, and contribute to the development of cancer and other diseases if any alterations to their regulation occur. Genetic abnormalities affecting the c-Abl tyrosine kinase are linked to chronic myelogenous leukemia, a cancer of immature cells in the bone marrow. In the noncancerous form of the protein, binding of a myristoyl group to the kinase domain inhibits the activity of the protein until it is needed (top left shows the inactive form, top right shows the open and active form). The cancerous variant of the protein, called Bcr-Abl, lacks this autoinhibitory myristoyl group and is continually active (bottom). ATP is shown in green bound in the active site of the kinase.
Find these in the RCSB Protein Data Bank: c-Abl tyrosine kinase and regulatory domains (PDB entry 1OPL) and F-actin binding domain (PDB entry 1ZZP).
Find these in the RCSB Protein Data Bank: c-Abl tyrosine kinase and regulatory domains (PDB entry 1OPL) and F-actin binding domain (PDB entry 1ZZP).
Amy Wu and Christine Zardecki, RCSB Protein Data Bank.
View Media
6999: HIV enzyme
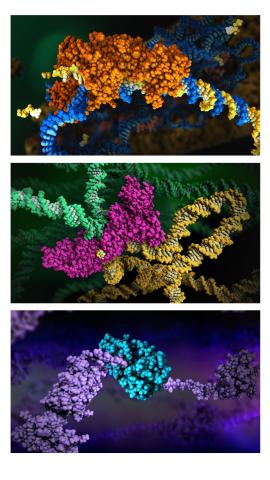
6999: HIV enzyme
These images model the molecular structures of three enzymes with critical roles in the life cycle of the human immunodeficiency virus (HIV). At the top, reverse transcriptase (orange) creates a DNA copy (yellow) of the virus's RNA genome (blue). In the middle image, integrase (magenta) inserts this DNA copy in the DNA genome (green) of the infected cell. At the bottom, much later in the viral life cycle, protease (turquoise) chops up a chain of HIV structural protein (purple) to generate the building blocks for making new viruses. See these enzymes in action on PDB 101’s video A Molecular View of HIV Therapy.
Amy Wu and Christine Zardecki, RCSB Protein Data Bank.
View Media
5777: Microsporidia in roundworm 1
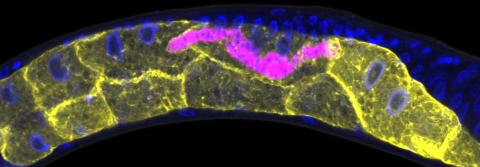
5777: Microsporidia in roundworm 1
Many disease-causing microbes manipulate their host’s metabolism and cells for their own ends. Microsporidia—which are parasites closely related to fungi—infect and multiply inside animal cells, and take the rearranging of cells’ interiors to a new level. They reprogram animal cells such that the cells start to fuse, causing them to form long, continuous tubes. As shown in this image of the roundworm Caenorhabditis elegans, microsporidia (shown in magenta) have invaded the worm’s gut cells (shown in yellow; the cells’ nuclei are shown in blue) and have instructed the cells to merge. The cell fusion enables the microsporidia to thrive and propagate in the expanded space. Scientists study microsporidia in worms to gain more insight into how these parasites manipulate their host cells. This knowledge might help researchers devise strategies to prevent or treat infections with microsporidia. For more on the research into microsporidia, see this news release from the University of California San Diego. Related to images 5778 and 5779.
Keir Balla and Emily Troemel, University of California San Diego
View Media
3765: Trypanosoma brucei, the cause of sleeping sickness
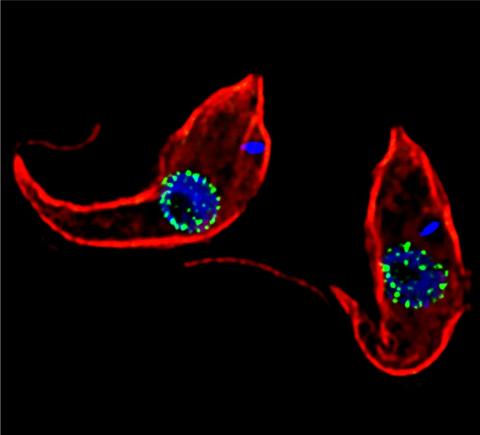
3765: Trypanosoma brucei, the cause of sleeping sickness
Trypanosoma brucei is a single-cell parasite that causes sleeping sickness in humans. Scientists have been studying trypanosomes for some time because of their negative effects on human and also animal health, especially in sub-Saharan Africa. Moreover, because these organisms evolved on a separate path from those of animals and plants more than a billion years ago, researchers study trypanosomes to find out what traits they may harbor that are common to or different from those of other eukaryotes (i.e., those organisms having a nucleus and mitochondria). This image shows the T. brucei cell membrane in red, the DNA in the nucleus and kinetoplast (a structure unique to protozoans, including trypanosomes, which contains mitochondrial DNA) in blue and nuclear pore complexes (which allow molecules to pass into or out of the nucleus) in green. Scientists have found that the trypanosome nuclear pore complex has a unique mechanism by which it attaches to the nuclear envelope. In addition, the trypanosome nuclear pore complex differs from those of other eukaryotes because its components have a near-complete symmetry, and it lacks almost all of the proteins that in other eukaryotes studied so far are required to assemble the pore.
Michael Rout, Rockefeller University
View Media
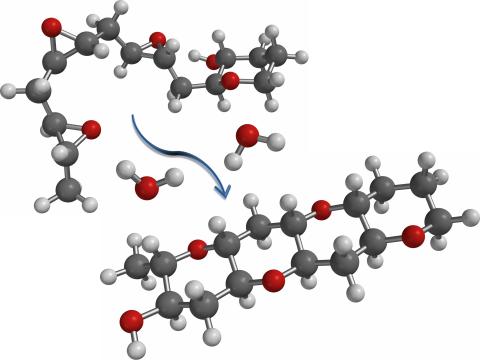
2490: Cascade reaction promoted by water
2490: Cascade reaction promoted by water
This illustration of an epoxide-opening cascade promoted by water emulates the proposed biosynthesis of some of the Red Tide toxins.
Tim Jamison, Massachusetts Institute of Technology
View Media
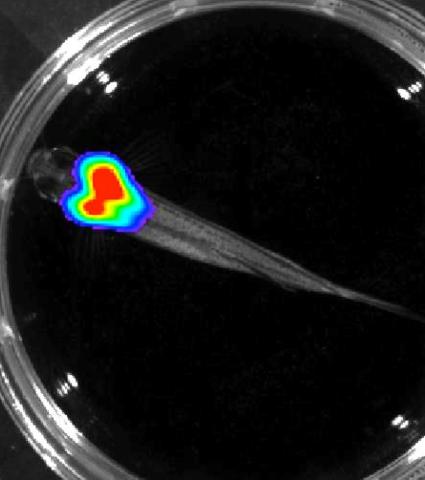
3557: Bioluminescent imaging in adult zebrafish - overhead view
3557: Bioluminescent imaging in adult zebrafish - overhead view
Luciferase-based imaging enables visualization and quantification of internal organs and transplanted cells in live adult zebrafish. In this image, a cardiac muscle-restricted promoter drives firefly luciferase expression.
For imagery of both the lateral and overhead view go to 3556.
For imagery of the lateral view go to 3558.
For more information about the illumated area go to 3559.
For imagery of both the lateral and overhead view go to 3556.
For imagery of the lateral view go to 3558.
For more information about the illumated area go to 3559.
Kenneth Poss, Duke University
View Media
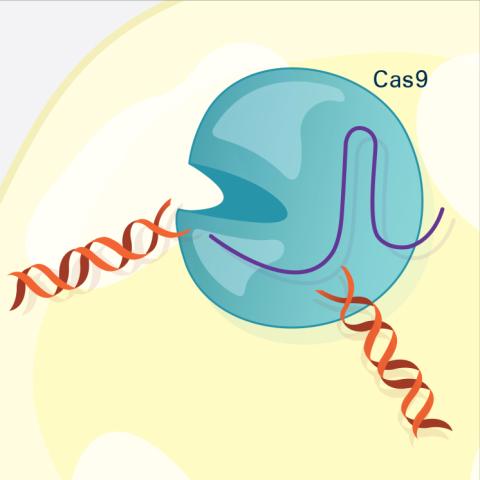
6487: CRISPR Illustration Frame 3
6487: CRISPR Illustration Frame 3
This illustration shows, in simplified terms, how the CRISPR-Cas9 system can be used as a gene-editing tool. The CRISPR system has two components joined together: a finely tuned targeting device (a small strand of RNA programmed to look for a specific DNA sequence) and a strong cutting device (an enzyme called Cas9 that can cut through a double strand of DNA). In this frame (3 of 4), the Cas9 enzyme cuts both strands of the DNA.
For an explanation and overview of the CRISPR-Cas9 system, see the iBiology video, and find the full CRIPSR illustration here.
For an explanation and overview of the CRISPR-Cas9 system, see the iBiology video, and find the full CRIPSR illustration here.
National Institute of General Medical Sciences.
View Media

2406: Hen egg lysozyme (2)
2406: Hen egg lysozyme (2)
A crystal of hen egg lysozyme protein created for X-ray crystallography, which can reveal detailed, three-dimensional protein structures.
Alex McPherson, University of California, Irvine
View Media
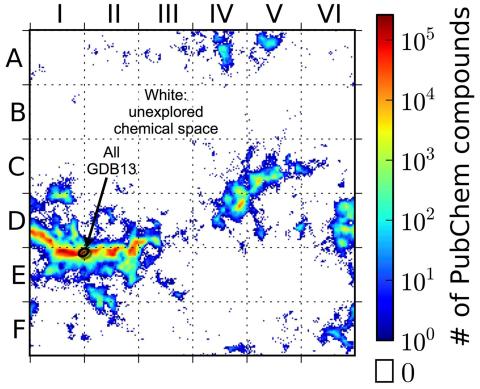
3458: Computer algorithm
3458: Computer algorithm
This computer algorithm plots all feasible small carbon-based molecules as though they were cities on a map and identifies huge, unexplored spaces that may help fuel research into new drug therapies. Featured in the May 16, 2013 issue of Biomedical Beat.
Aaron Virshup, Julia Contreras-Garcia, Peter Wipf, Weitao Yang and David Beratan, University of Pittsburgh Center for Chemical Methodologies and Library Development
View Media
3262: Caulobacter
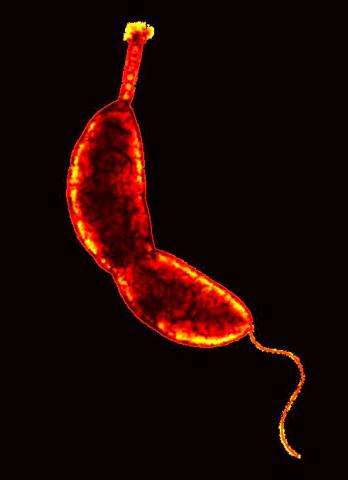
3262: Caulobacter
A study using Caulobacter crescentus showed that some bacteria use just-in-time processing, much like that used in industrial delivery, to make the glue that allows them to attach to surfaces, an important step in the infection process for many disease-causing bacteria. In the image shown, this freshwater bacterium has a holdfast at the top and a propelling flagellum at the end. From an Indiana University news release.
Yves Brun, Indiana University
View Media

6536: Sepsis Infographic
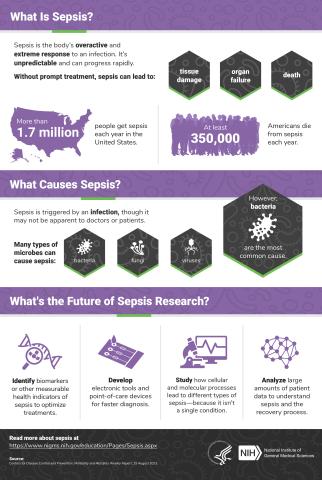
6536: Sepsis Infographic
Sepsis is the body’s overactive and extreme response to an infection. More than 1.7 million people get sepsis each year in the United States. Without prompt treatment, sepsis can lead to tissue damage, organ failure, and death. Many NIGMS-supported researchers are working to improve sepsis diagnosis and treatment. Learn more with our sepsis featured topic page.
See 6551 for the Spanish version of this infographic.
See 6551 for the Spanish version of this infographic.
National Institute of General Medical Sciences
View Media
2566: Haplotypes
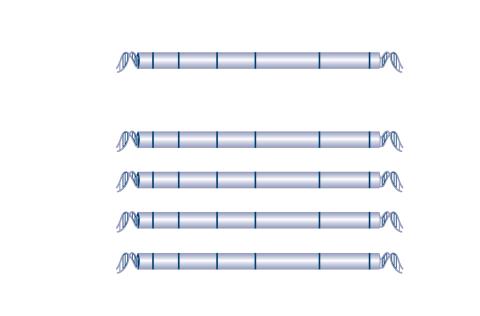
2566: Haplotypes
Haplotypes are combinations of gene variants that are likely to be inherited together within the same chromosomal region. In this example, an original haplotype (top) evolved over time to create three newer haplotypes that each differ by a few nucleotides (red). See image 2567 for a labeled version of this illustration. Featured in The New Genetics.
Crabtree + Company
View Media

3690: Microscopy image of bird-and-flower DNA origami
3690: Microscopy image of bird-and-flower DNA origami
An atomic force microscopy image shows DNA folded into an intricate, computer-designed structure. Image is featured on Biomedical Beat blog post Cool Image: DNA Origami. See also related image 3689 .
Hao Yan, Arizona State University
View Media
6489: CRISPR Illustration Frame 5
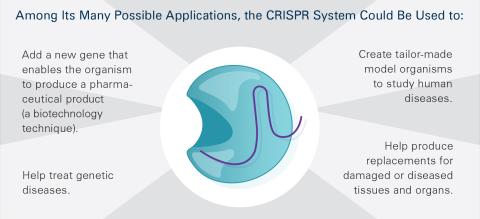
6489: CRISPR Illustration Frame 5
This illustration shows, in simplified terms, how the CRISPR-Cas9 system can be used as a gene-editing tool. This is the fifthframe in a series of five. The CRISPR system has two components joined together: a finely tuned targeting device (a small strand of RNA programmed to look for a specific DNA sequence) and a strong cutting device (an enzyme called Cas9 that can cut through a double strand of DNA). For an explanation and overview of the CRISPR-Cas9 system, see the NIGMS Biomedical Beat blog entry, Field Focus: Precision Gene Editing with CRISPR and the iBiology video, Genome Engineering with CRISPR-Cas9: Birth of a Breakthrough Technology.
View Media
2511: X-ray crystallography
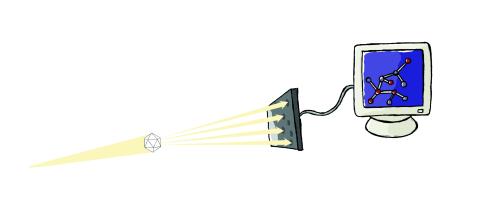
2511: X-ray crystallography
X-ray crystallography allows researchers to see structures too small to be seen by even the most powerful microscopes. To visualize the arrangement of atoms within molecules, researchers can use the diffraction patterns obtained by passing X-ray beams through crystals of the molecule. This is a common way for solving the structures of proteins. See image 2512 for a labeled version of this illustration. Featured in The Structures of Life.
Crabtree + Company
View Media
3566: Mouse colon with gut bacteria
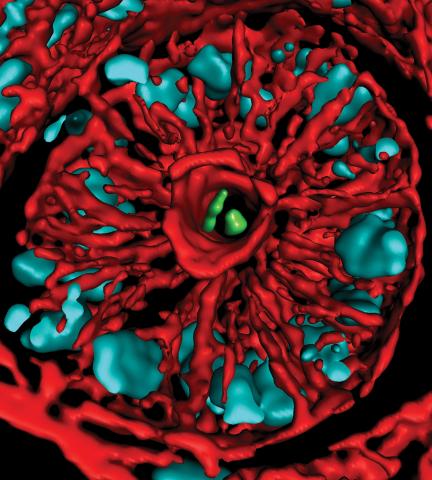
3566: Mouse colon with gut bacteria
A section of mouse colon with gut bacteria (center, in green) residing within a protective pocket. Understanding how microorganisms colonize the gut could help devise ways to correct for abnormal changes in bacterial communities that are associated with disorders like inflammatory bowel disease.
Sarkis K. Mazmanian, California Institute of Technology
View Media
6933: Zebrafish head vasculature video
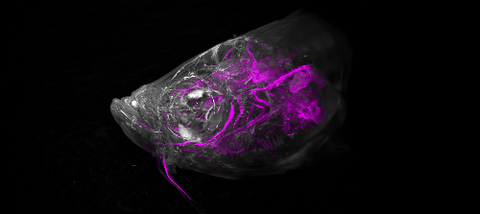
6933: Zebrafish head vasculature video
Various views of a zebrafish head with blood vessels shown in purple. Researchers often study zebrafish because they share many genes with humans, grow and reproduce quickly, and have see-through eggs and embryos, which make it easy to study early stages of development.
This video was captured using a light sheet microscope.
Related to image 6934.
This video was captured using a light sheet microscope.
Related to image 6934.
Prayag Murawala, MDI Biological Laboratory and Hannover Medical School.
View Media
1339: Egg comparison
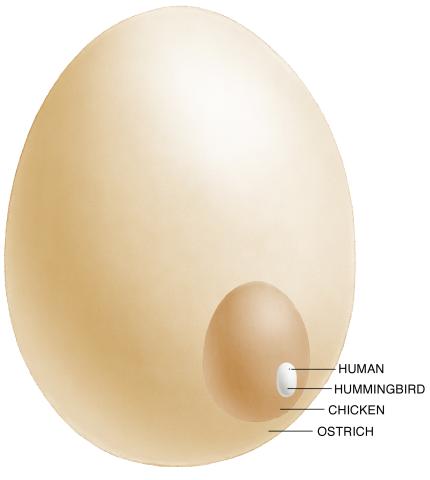
1339: Egg comparison
The largest human cell (by volume) is the egg. Human eggs are 150 micrometers in diameter and you can just barely see one with a naked eye. In comparison, consider the eggs of chickens...or ostriches!
Judith Stoffer
View Media
2299: 2-D NMR
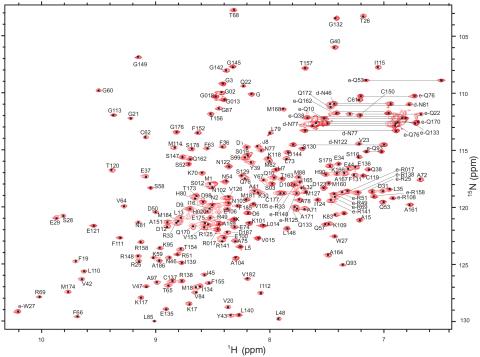
2299: 2-D NMR
A two-dimensional NMR spectrum of a protein, in this case a 2D 1H-15N HSQC NMR spectrum of a 228 amino acid DNA/RNA-binding protein.
Dr. Xiaolian Gao's laboratory at the University of Houston
View Media
2351: tRNA splicing enzyme endonuclease in humans
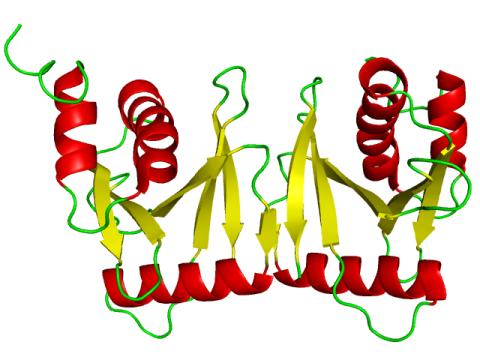
2351: tRNA splicing enzyme endonuclease in humans
An NMR solution structure model of the transfer RNA splicing enzyme endonuclease in humans (subunit Sen15). This represents the first structure of a eukaryotic tRNA splicing endonuclease subunit.
Center for Eukaryotic Structural Genomics, PSI
View Media
3662: Mitochondrion from insect flight muscle
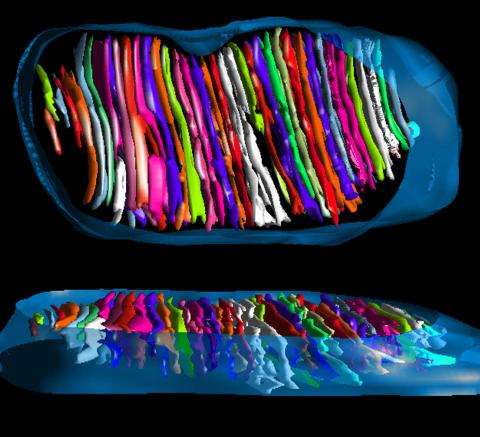
3662: Mitochondrion from insect flight muscle
This is a tomographic reconstruction of a mitochondrion from an insect flight muscle. Mitochondria are cellular compartments that are best known as the powerhouses that convert energy from the food into energy that runs a range of biological processes. Nearly all our cells have mitochondria.
National Center for Microscopy and Imaging Research
View Media
6613: Circadian rhythms and the SCN
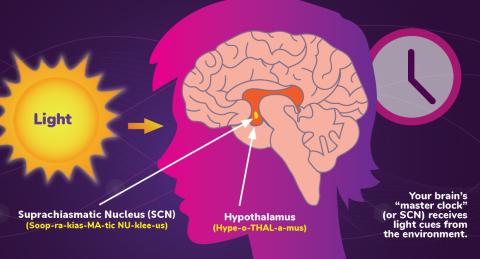
6613: Circadian rhythms and the SCN
Circadian rhythms are physical, mental, and behavioral changes that follow a 24-hour cycle. Circadian rhythms are influenced by light and regulated by the brain’s suprachiasmatic nucleus (SCN), sometimes referred to as a master clock. Learn more in NIGMS’ circadian rhythms fact sheet. See 6614 for the Spanish version of this infographic.
NIGMS
View Media

2305: Beaded bacteriophage
2305: Beaded bacteriophage
This sculpture made of purple and clear glass beads depicts bacteriophage Phi174, a virus that infects bacteria. It rests on a surface that portrays an adaptive landscape, a conceptual visualization. The ridges represent the gene combinations associated with the greatest fitness levels of the virus, as measured by how quickly the virus can reproduce itself. Phi174 is an important model system for studies of viral evolution because its genome can readily be sequenced as it evolves under defined laboratory conditions.
Holly Wichman, University of Idaho. (Surface by A. Johnston; photo by J. Palmersheim)
View Media
6893: Chromatin in human tenocyte
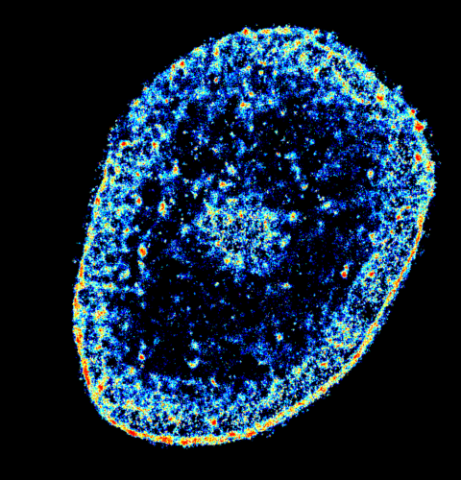
6893: Chromatin in human tenocyte
The nucleus of a degenerating human tendon cell, also known as a tenocyte. It has been color-coded based on the density of chromatin—a substance made up of DNA and proteins. Areas of low chromatin density are shown in blue, and areas of high chromatin density are shown in red. This image was captured using Stochastic Optical Reconstruction Microscopy (STORM).
Related to images 6887 and 6888.
Related to images 6887 and 6888.
Melike Lakadamyali, Perelman School of Medicine at the University of Pennsylvania.
View Media
6486: CRISPR Illustration Frame 2
6486: CRISPR Illustration Frame 2
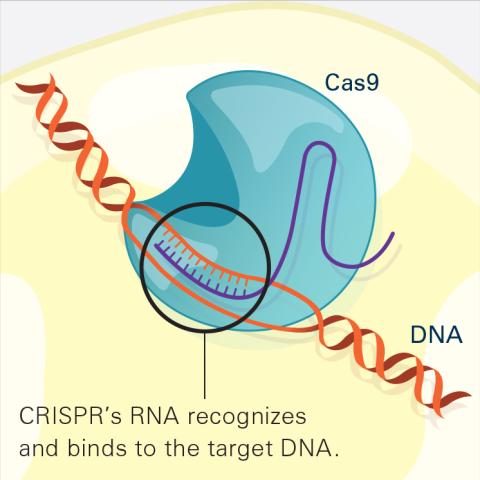
This illustration shows, in simplified terms, how the CRISPR-Cas9 system can be used as a gene-editing tool. The CRISPR system has two components joined together: a finely tuned targeting device (a small strand of RNA programmed to look for a specific DNA sequence) and a strong cutting device (an enzyme called Cas9 that can cut through a double strand of DNA). In this frame (2 of 4), the CRISPR machine locates the target DNA sequence once inserted into a cell.
For an explanation and overview of the CRISPR-Cas9 system, see the iBiology video, and find the full CRIPSR illustration here.
For an explanation and overview of the CRISPR-Cas9 system, see the iBiology video, and find the full CRIPSR illustration here.
National Institute of General Medical Sciences.
View Media

2683: GFP sperm
2683: GFP sperm
Fruit fly sperm cells glow bright green when they express the gene for green fluorescent protein (GFP).
View Media

6344: Drosophila
6344: Drosophila
Two adult fruit flies (Drosophila)
Dr. Vicki Losick, MDI Biological Laboratory, www.mdibl.org
View Media
2727: Proteins related to myotonic dystrophy
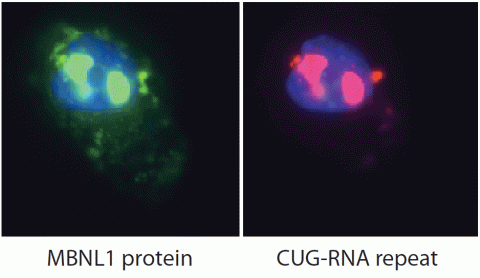
2727: Proteins related to myotonic dystrophy
Myotonic dystrophy is thought to be caused by the binding of a protein called Mbnl1 to abnormal RNA repeats. In these two images of the same muscle precursor cell, the top image shows the location of the Mbnl1 splicing factor (green) and the bottom image shows the location of RNA repeats (red) inside the cell nucleus (blue). The white arrows point to two large foci in the cell nucleus where Mbnl1 is sequestered with RNA.
Manuel Ares, University of California, Santa Cruz
View Media
1306: Vesicular shuttle model
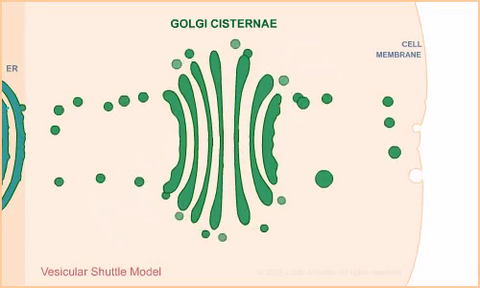
1306: Vesicular shuttle model
Animation for the vesicular shuttle model of Golgi transport.
Judith Stoffer
View Media
6983: Genetic mosaicism in fruit flies
6983: Genetic mosaicism in fruit flies
Fat tissue from the abdomen of a genetically mosaic adult fruit fly. Genetic mosaicism means that the fly has cells with different genotypes even though it formed from a single zygote. This specific mosaicism results in accumulation of a critical fly adipokine (blue-green) within the fat tissue cells that have reduced expression a key nutrient sensing gene (in left panel). The dotted line shows the cells lacking the gene that is present and functioning in the rest of the cells. Nuclei are labelled in magenta. This image was captured using a confocal microscope and shows a maximum intensity projection of many slices.
Related to images 6982, 6984, and 6985.
Related to images 6982, 6984, and 6985.
Akhila Rajan, Fred Hutchinson Cancer Center
View Media
5760: Annotated TEM cross-section of C. elegans (roundworm)
5760: Annotated TEM cross-section of C. elegans (roundworm)
The worm Caenorhabditis elegans is a popular laboratory animal because its small size and fairly simple body make it easy to study. Scientists use this small worm to answer many research questions in developmental biology, neurobiology, and genetics. This image, which was taken with transmission electron microscopy (TEM), shows a cross-section through C. elegans, revealing various internal structures labeled in the image. You can find a high-resolution image without the annotations at image 5759.
The image is from a figure in an article published in the journal eLife.
The image is from a figure in an article published in the journal eLife.
Piali Sengupta, Brandeis University
View Media
5811: NCMIR Tongue 2
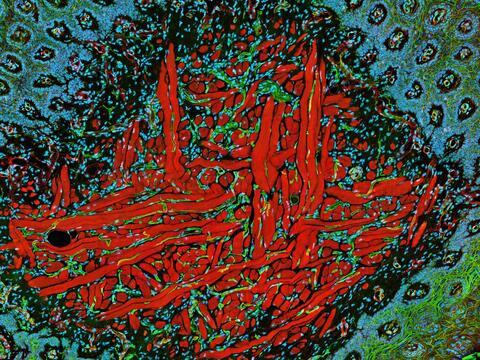
5811: NCMIR Tongue 2
Microscopy image of a tongue. One in a series of two, see image 5810
National Center for Microscopy and Imaging Research (NCMIR)
View Media
1017: Lily mitosis 07
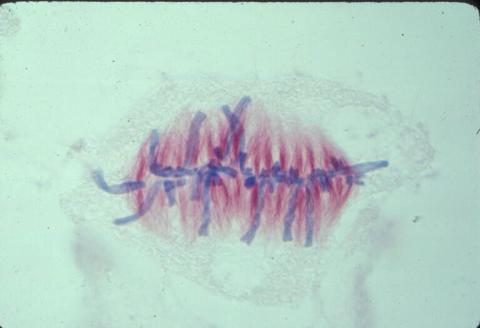
1017: Lily mitosis 07
A light microscope image of a cell from the endosperm of an African globe lily (Scadoxus katherinae). This is one frame of a time-lapse sequence that shows cell division in action. The lily is considered a good organism for studying cell division because its chromosomes are much thicker and easier to see than human ones. Staining shows microtubules in red and chromosomes in blue. Here, condensed chromosomes are clearly visible and have lined up in the middle of the dividing cell.
Related to images 1010, 1011, 1012, 1013, 1014, 1015, 1016, 1018, 1019, and 1021.
Related to images 1010, 1011, 1012, 1013, 1014, 1015, 1016, 1018, 1019, and 1021.
Andrew S. Bajer, University of Oregon, Eugene
View Media
2433: Fruit fly sperm cells
2433: Fruit fly sperm cells
Developing fruit fly spermatids require caspase activity (green) for the elimination of unwanted organelles and cytoplasm via apoptosis.
Hermann Steller, Rockefeller University
View Media
2330: Repairing DNA
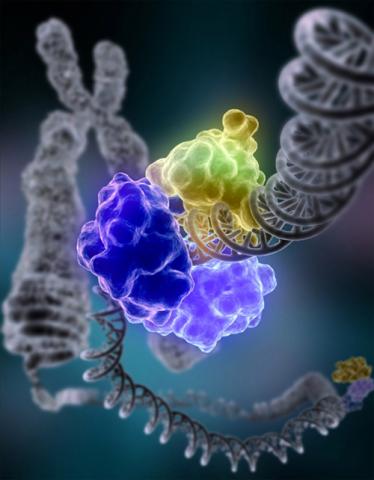
2330: Repairing DNA
Like a watch wrapped around a wrist, a special enzyme encircles the double helix to repair a broken strand of DNA. Without molecules that can mend such breaks, cells can malfunction, die, or become cancerous. Related to image 3493.
Tom Ellenberger, Washington University School of Medicine
View Media
2716: Mycobacterium tuberculosis
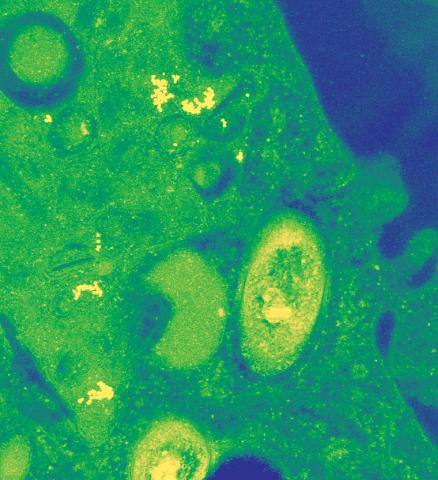
2716: Mycobacterium tuberculosis
Mycobacterium tuberculosis, the bacterium that causes tuberculosis, has infected one-quarter of the world's population and causes more than one million deaths each year, according to the World Health Organization.
Reuben Peters, Iowa State University
View Media
5729: Assembly of the HIV capsid
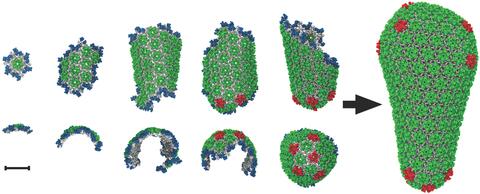
5729: Assembly of the HIV capsid
The HIV capsid is a pear-shaped structure that is made of proteins the virus needs to mature and become infective. The capsid is inside the virus and delivers the virus' genetic information into a human cell. To better understand how the HIV capsid does this feat, scientists have used computer programs to simulate its assembly. This image shows a series of snapshots of the steps that grow the HIV capsid. A model of a complete capsid is shown on the far right of the image for comparison; the green, blue and red colors indicate different configurations of the capsid protein that make up the capsid “shell.” The bar in the left corner represents a length of 20 nanometers, which is less than a tenth the size of the smallest bacterium. Computer models like this also may be used to reconstruct the assembly of the capsids of other important viruses, such as Ebola or the Zika virus. The studies reporting this research were published in Nature Communications and Nature. To learn more about how researchers used computer simulations to track the assembly of the HIV capsid, see this press release from the University of Chicago.
John Grime and Gregory Voth, The University of Chicago
View Media
3330: mDia1 antibody staining-01
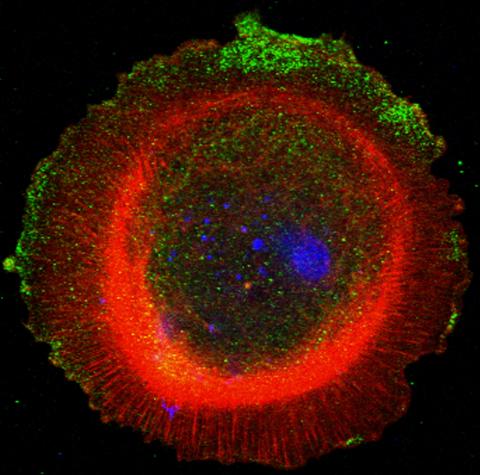
3330: mDia1 antibody staining-01
Cells move forward with lamellipodia and filopodia supported by networks and bundles of actin filaments. Proper, controlled cell movement is a complex process. Recent research has shown that an actin-polymerizing factor called the Arp2/3 complex is the key component of the actin polymerization engine that drives amoeboid cell motility. ARPC3, a component of the Arp2/3 complex, plays a critical role in actin nucleation. In this photo, the ARPC3+/+ fibroblast cells were fixed and stained with Alexa 546 phalloidin for F-actin (red), mDia1 (green), and DAPI to visualize the nucleus (blue). mDia1 is localized at the lamellipodia of ARPC3+/+ fibroblast cells. Related to images 3328, 3329, 3331, 3332, and 3333.
Rong Li and Praveen Suraneni, Stowers Institute for Medical Research
View Media

2392: Sheep hemoglobin crystal
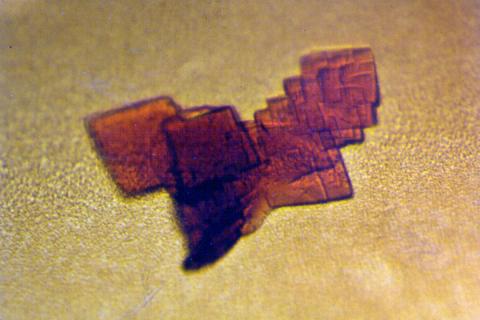
2392: Sheep hemoglobin crystal
A crystal of sheep hemoglobin protein created for X-ray crystallography, which can reveal detailed, three-dimensional protein structures.
Alex McPherson, University of California, Irvine
View Media
3434: Flu virus proteins during self-replication
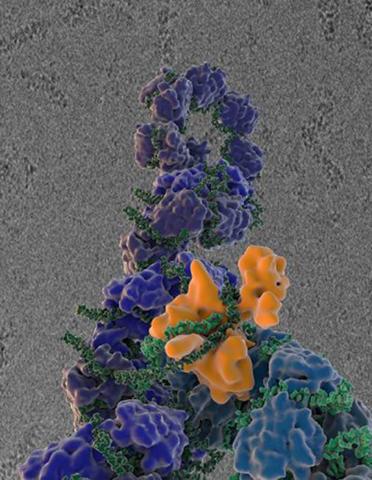
3434: Flu virus proteins during self-replication
Influenza (flu) virus proteins in the act of self-replication. Viral nucleoprotein (blue) encapsidates [encapsulates] the RNA genome (green). The influenza virus polymerase (orange) reads and copies the RNA genome. In the background is an image of influenza virus ribonucleoprotein complexes observed using cryo-electron microscopy. This image is from a November 2012 News Release.
Scripps Research Institute in La Jolla, CA
View Media