Switch to List View
Image and Video Gallery
This is a searchable collection of scientific photos, illustrations, and videos. The images and videos in this gallery are licensed under Creative Commons Attribution Non-Commercial ShareAlike 3.0. This license lets you remix, tweak, and build upon this work non-commercially, as long as you credit and license your new creations under identical terms.
2321: Microtubule breakdown
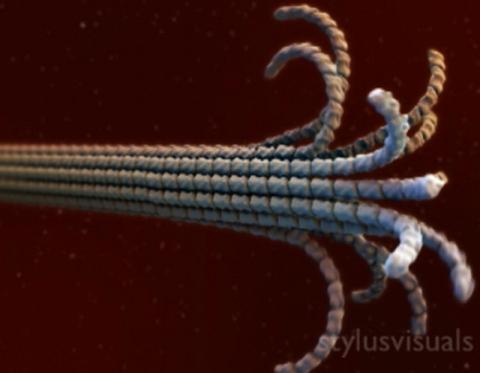
2321: Microtubule breakdown
Like a building supported by a steel frame, a cell contains its own sturdy internal scaffolding made up of proteins, including microtubules. Researchers studying snapshots of microtubules have proposed a model for how these structural elements shorten and lengthen, allowing a cell to move, divide, or change shape. This picture shows an intermediate step in the model: Smaller building blocks called tubulins peel back from the microtubule in thin strips. Knowing the operations of the internal scaffolding will enhance our basic understanding of cellular processes.
Eva Nogales, University of California, Berkeley
View Media
2781: Disease-resistant Arabidopsis leaf
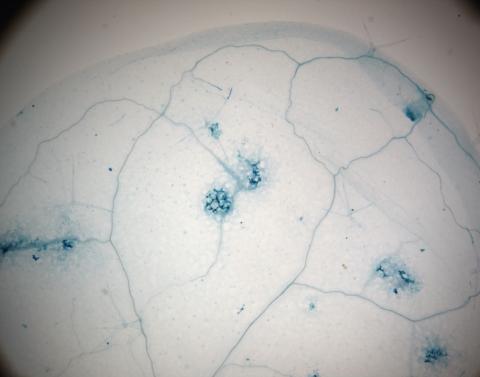
2781: Disease-resistant Arabidopsis leaf
This is a magnified view of an Arabidopsis thaliana leaf a few days after being exposed to the pathogen Hyaloperonospora arabidopsidis. The plant from which this leaf was taken is genetically resistant to the pathogen. The spots in blue show areas of localized cell death where infection occurred, but it did not spread. Compare this response to that shown in Image 2782. Jeff Dangl has been funded by NIGMS to study the interactions between pathogens and hosts that allow or suppress infection.
Jeff Dangl, University of North Carolina, Chapel Hill
View Media

6771: Culex quinquefasciatus mosquito larvae
6771: Culex quinquefasciatus mosquito larvae
Mosquito larvae with genes edited by CRISPR swimming in water. This species of mosquito, Culex quinquefasciatus, can transmit West Nile virus, Japanese encephalitis virus, and avian malaria, among other diseases. The researchers who took this video optimized the gene-editing tool CRISPR for Culex quinquefasciatus that could ultimately help stop the mosquitoes from spreading pathogens. The work is described in the Nature Communications paper "Optimized CRISPR tools and site-directed transgenesis towards gene drive development in Culex quinquefasciatus mosquitoes" by Feng et al. Related to images 6769 and 6770.
Valentino Gantz, University of California, San Diego.
View Media
2578: Cellular aging
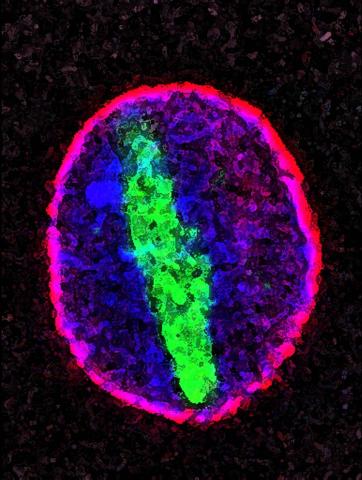
2578: Cellular aging
A protein called tubulin (green) accumulates in the center of a nucleus (outlined in pink) from an aging cell. Normally, this protein is kept out of the nucleus with the help of gatekeepers known as nuclear pore complexes. But NIGMS-funded researchers found that wear and tear to long-lived components of the complexes eventually lowers the gatekeepers' guard. As a result, cytoplasmic proteins like tubulin gain entry to the nucleus while proteins normally confined to the nucleus seep out. The work suggests that finding ways to stop the leakage could slow the cellular aging process and possibly lead to new therapies for age-related diseases.
Maximiliano D'Angelo and Martin Hetzer, Salk Institute
View Media
5883: Beta-galactosidase montage showing cryo-EM improvement--gradient background
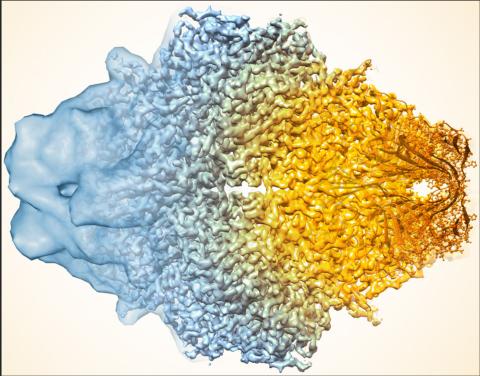
5883: Beta-galactosidase montage showing cryo-EM improvement--gradient background
Composite image of beta-galactosidase showing how cryo-EM’s resolution has improved dramatically in recent years. Older images to the left, more recent to the right. Related to image 5882. NIH Director Francis Collins featured this on his blog on January 14, 2016.
Veronica Falconieri, Sriram Subramaniam Lab, National Cancer Institute
View Media
3360: H1 histamine receptor
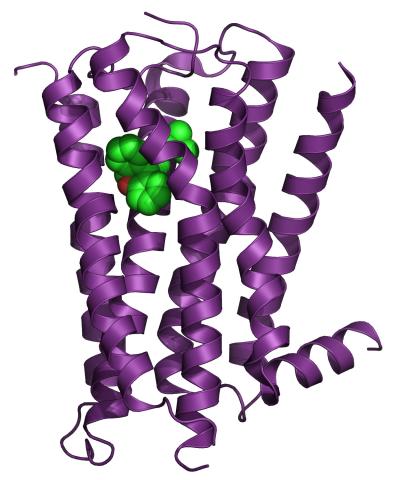
3360: H1 histamine receptor
The receptor is shown bound to an inverse agonist, doxepin.
Raymond Stevens, The Scripps Research Institute
View Media
2556: Dicer generates microRNAs
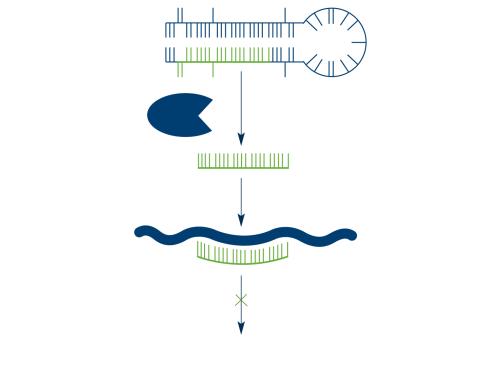
2556: Dicer generates microRNAs
The enzyme Dicer generates microRNAs by chopping larger RNA molecules into tiny Velcro®-like pieces. MicroRNAs stick to mRNA molecules and prevent the mRNAs from being made into proteins. See image 2557 for a labeled version of this illustration. Featured in The New Genetics.
Crabtree + Company
View Media
3675: NCMIR kidney-1
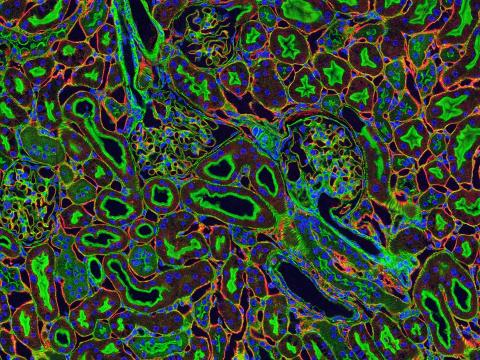
3675: NCMIR kidney-1
Stained kidney tissue. The kidney is an essential organ responsible for disposing wastes from the body and for maintaining healthy ion levels in the blood. It also secretes two hormones, erythropoietin (EPO) and calcitriol (a derivative of vitamin D), into the blood. It works like a purifier by pulling break-down products of metabolism, such as urea and ammonium, from the blood stream for excretion in urine. Related to image 3725.
Tom Deerinck, National Center for Microscopy and Imaging Research (NCMIR)
View Media
3387: NCMIR human spinal nerve
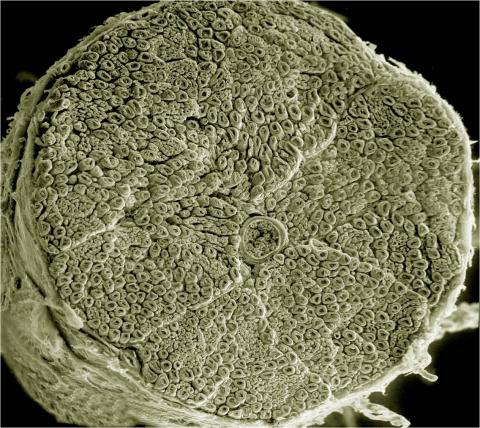
3387: NCMIR human spinal nerve
Spinal nerves are part of the peripheral nervous system. They run within the spinal column to carry nerve signals to and from all parts of the body. The spinal nerves enable all the movements we do, from turning our heads to wiggling our toes, control the movements of our internal organs, such as the colon and the bladder, as well as allow us to feel touch and the location of our limbs.
Tom Deerinck, National Center for Microscopy and Imaging Research (NCMIR)
View Media
3723: Fluorescent microscopy of kidney tissue
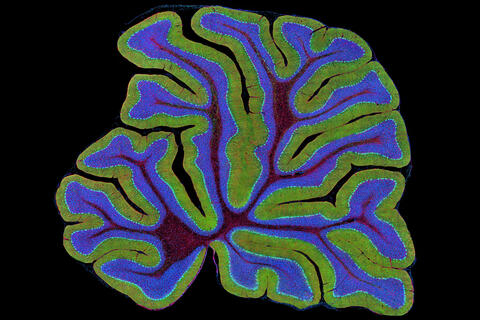
3723: Fluorescent microscopy of kidney tissue
Serum albumin (SA) is the most abundant protein in the blood plasma of mammals. SA has a characteristic heart-shape structure and is a highly versatile protein. It helps maintain normal water levels in our tissues and carries almost half of all calcium ions in human blood. SA also transports some hormones, nutrients and metals throughout the bloodstream. Despite being very similar to our own SA, those from other animals can cause some mild allergies in people. Therefore, some scientists study SAs from humans and other mammals to learn more about what subtle structural or other differences cause immune responses in the body.
Related to entries 3725 and 3675.
Related to entries 3725 and 3675.
Tom Deerinck , National Center for Microscopy and Imaging Research
View Media
3361: A2A adenosine receptor
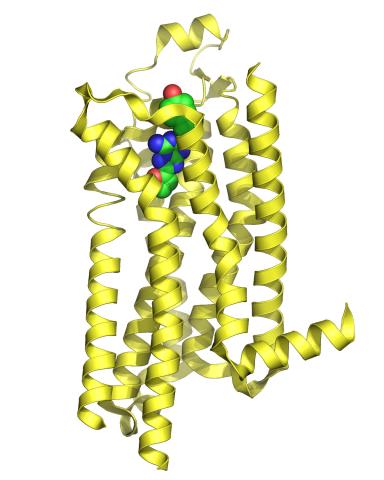
3361: A2A adenosine receptor
The receptor is shown bound to an inverse agonist, ZM241385.
Raymond Stevens, The Scripps Research Institute
View Media
3254: Pulsating response to stress in bacteria - video
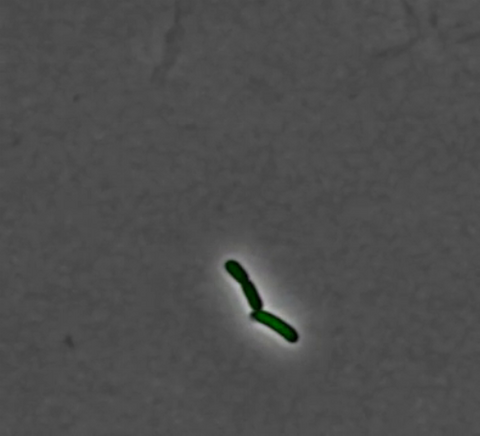
3254: Pulsating response to stress in bacteria - video
By attaching fluorescent proteins to the genetic circuit responsible for B. subtilis's stress response, researchers can observe the cells' pulses as green flashes. This video shows flashing cells as they multiply over the course of more than 12 hours. In response to a stressful environment like one lacking food, B. subtilis activates a large set of genes that help it respond to the hardship. Instead of leaving those genes on as previously thought, researchers discovered that the bacteria flip the genes on and off, increasing the frequency of these pulses with increasing stress. See entry 3253 for a related still image.
Michael Elowitz, Caltech University
View Media
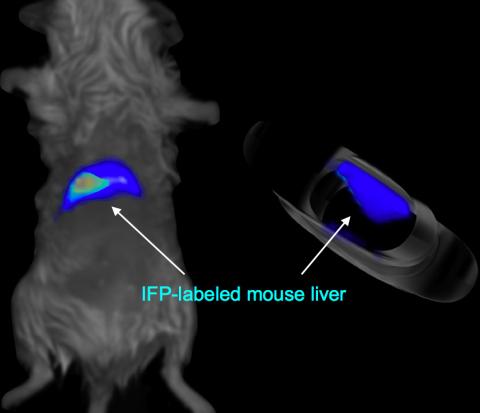
2601: Mouse liver labeled with fluorescent probe
2601: Mouse liver labeled with fluorescent probe
A mouse liver glows after being tagged with specially designed infrared-fluorescent protein (IFP). Since its discovery in 1962, green fluorescent protein (GFP) has become an invaluable resource in biomedical imaging. But because of its short wavelength, the light that makes GFP glow doesn't penetrate far in whole animals. So University of California, San Diego cell biologist Roger Tsien--who shared the 2008 Nobel Prize in chemistry for groundbreaking work with GFP--made infrared-fluorescent proteins (IFPs) that shine under longer-wavelength light, allowing whole-body imaging in small animals.
Xiaokun Shu, University of California, San Diego
View Media
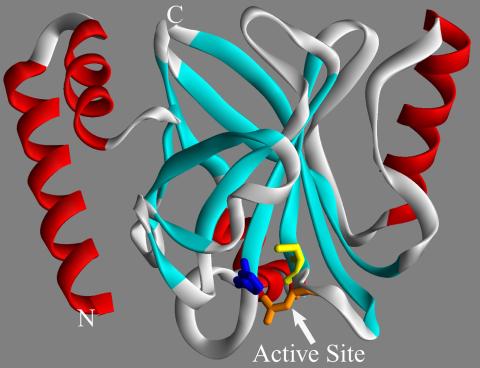
2386: Sortase b from B. anthracis
2386: Sortase b from B. anthracis
Structure of sortase b from the bacterium B. anthracis, which causes anthrax. Sortase b is an enzyme used to rob red blood cells of iron, which the bacteria need to survive.
Midwest Center for Structural Genomics, PSI
View Media
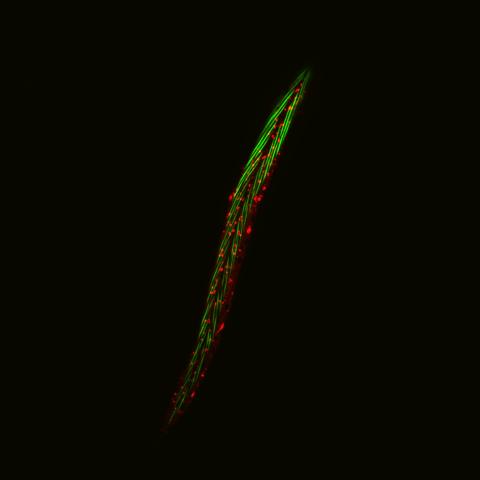
6581: Fluorescent C. elegans showing muscle and ribosomal protein
6581: Fluorescent C. elegans showing muscle and ribosomal protein
C. elegans, a tiny roundworm, with a ribosomal protein glowing red and muscle fibers glowing green. Researchers used these worms to study a molecular pathway that affects aging. The ribosomal protein is involved in protein translation and may play a role in dietary restriction-induced longevity. Image created using confocal microscopy.
View group of roundworms here 6582.
View closeup of roundworms here 6583.
View group of roundworms here 6582.
View closeup of roundworms here 6583.
Jarod Rollins, Mount Desert Island Biological Laboratory.
View Media
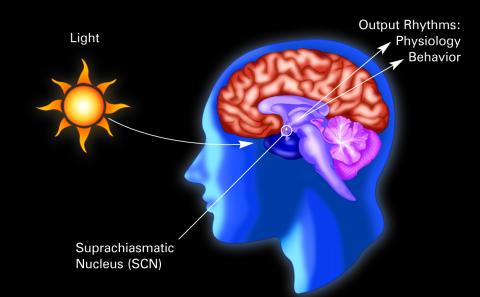
2569: Circadian rhythm (with labels)
2569: Circadian rhythm (with labels)
The human body keeps time with a master clock called the suprachiasmatic nucleus or SCN. Situated inside the brain, it's a tiny sliver of tissue about the size of a grain of rice, located behind the eyes. It sits quite close to the optic nerve, which controls vision, and this means that the SCN "clock" can keep track of day and night. The SCN helps control sleep and maintains our circadian rhythm--the regular, 24-hour (or so) cycle of ups and downs in our bodily processes such as hormone levels, blood pressure, and sleepiness. The SCN regulates our circadian rhythm by coordinating the actions of billions of miniature "clocks" throughout the body. These aren't actually clocks, but rather are ensembles of genes inside clusters of cells that switch on and off in a regular, 24-hour (or so) cycle in our physiological day.
Crabtree + Company
View Media
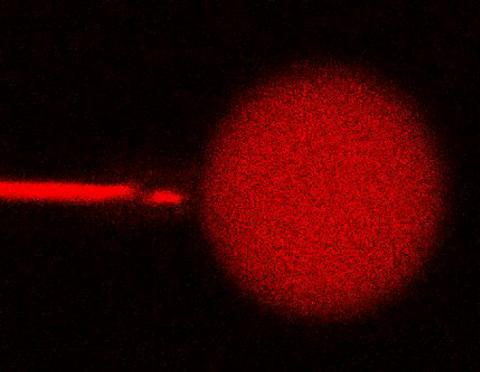
3583: Bee venom toxin destroying a cell
3583: Bee venom toxin destroying a cell
This video condenses 6.5 minutes into less than a minute to show how the toxin in bee venom, called melittin, destroys an animal or bacterial cell. What looks like a red balloon is an artificial cell filled with red dye. Melittin molecules are colored green and float on the cell's surface like twigs on a pond. As melittin accumulates on the cell's membrane, the membrane expands to accommodate it. In the video, the membrane stretches into a column on the left. When melittin levels reach a critical threshold, countless pinhole leaks burst open in the membrane. The cell's vital fluids (red dye in the video) leak out through these pores. Within minutes, the cell collapses.
Huey Huang, Rice University
View Media
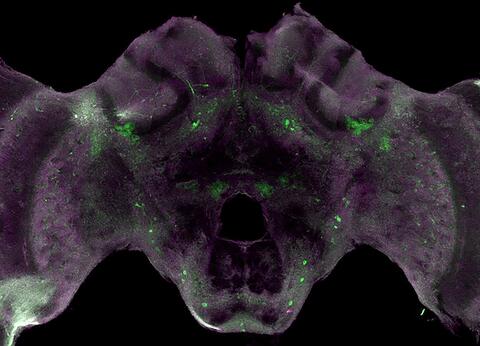
6755: Honeybee brain
6755: Honeybee brain
Insect brains, like the honeybee brain shown here, are very different in shape from human brains. Despite that, bee and human brains have a lot in common, including many of the genes and neurochemicals they rely on in order to function. The bright-green spots in this image indicate the presence of tyrosine hydroxylase, an enzyme that allows the brain to produce dopamine. Dopamine is involved in many important functions, such as the ability to experience pleasure. This image was captured using confocal microscopy.
Gene Robinson, University of Illinois at Urbana-Champaign.
View Media
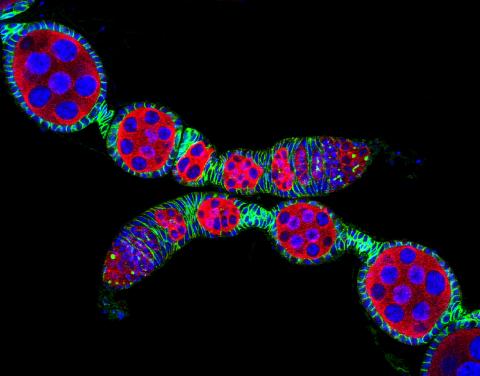
5772: Confocal microscopy image of two Drosophila ovarioles
5772: Confocal microscopy image of two Drosophila ovarioles
Ovarioles in female insects are tubes in which egg cells (called oocytes) form at one end and complete their development as they reach the other end of the tube. This image, taken with a confocal microscope, shows ovarioles in a very popular lab animal, the fruit fly Drosophila. The basic structure of ovarioles supports very rapid egg production, with some insects (like termites) producing several thousand eggs per day. Each insect ovary typically contains four to eight ovarioles, but this number varies widely depending on the insect species.
Scientists use insect ovarioles, for example, to study the basic processes that help various insects, including those that cause disease (like some mosquitos and biting flies), reproduce very quickly.
Scientists use insect ovarioles, for example, to study the basic processes that help various insects, including those that cause disease (like some mosquitos and biting flies), reproduce very quickly.
2004 Olympus BioScapes Competition
View Media
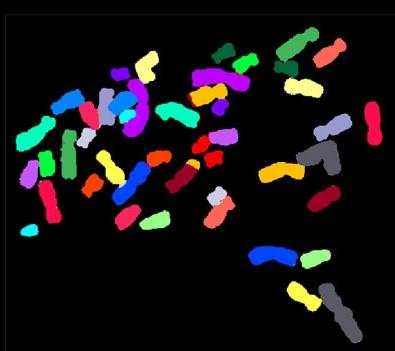
2312: Color-coded chromosomes
2312: Color-coded chromosomes
By mixing fluorescent dyes like an artist mixes paints, scientists are able to color code individual chromosomes. The technique, abbreviated multicolor-FISH, allows researchers to visualize genetic abnormalities often linked to disease. In this image, "painted" chromosomes from a person with a hereditary disease called Werner Syndrome show where a piece of one chromosome has fused to another (see the gold-tipped maroon chromosome in the center). As reported by molecular biologist Jan Karlseder of the Salk Institute for Biological Studies, such damage is typical among people with this rare syndrome.
Anna Jauch, Institute of Human Genetics, Heidelberg, Germany
View Media
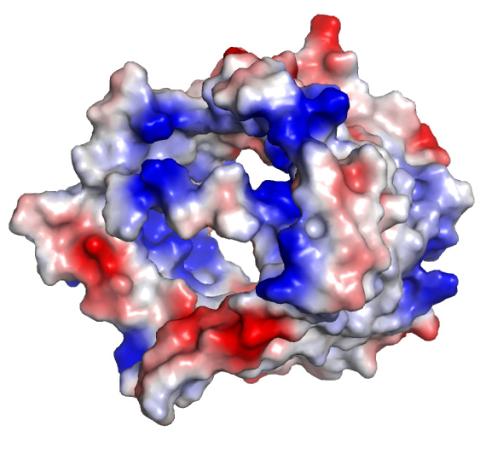
2494: VDAC-1 (3)
2494: VDAC-1 (3)
The structure of the pore-forming protein VDAC-1 from humans. This molecule mediates the flow of products needed for metabolism--in particular the export of ATP--across the outer membrane of mitochondria, the power plants for eukaryotic cells. VDAC-1 is involved in metabolism and the self-destruction of cells--two biological processes central to health.
Related to images 2491, 2495, and 2488.
Related to images 2491, 2495, and 2488.
Gerhard Wagner, Harvard Medical School
View Media
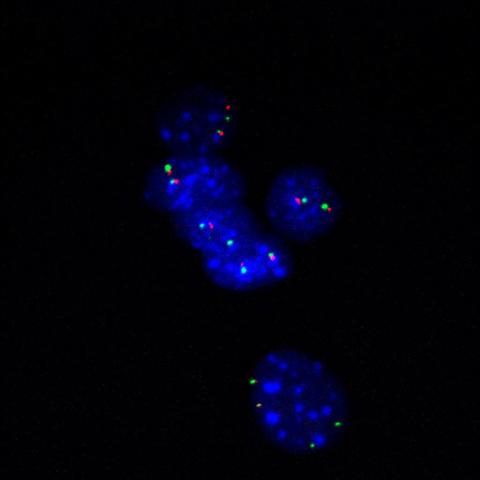
3296: Fluorescence in situ hybridization (FISH) in mouse ES cells shows DNA interactions
3296: Fluorescence in situ hybridization (FISH) in mouse ES cells shows DNA interactions
Researchers used fluorescence in situ hybridization (FISH) to confirm the presence of long range DNA-DNA interactions in mouse embryonic stem cells. Here, two loci labeled in green (Oct4) and red that are 13 Mb apart on linear DNA are frequently found to be in close proximity. DNA-DNA colocalizations like this are thought to both reflect and contribute to cell type specific gene expression programs.
Kathrin Plath, University of California, Los Angeles
View Media
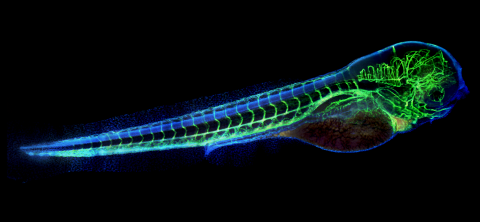
6661: Zebrafish embryo showing vasculature
6661: Zebrafish embryo showing vasculature
A zebrafish embryo. The blue areas are cell bodies, the green lines are blood vessels, and the red glow is blood. This image was created by stitching together five individual images captured with a hyperspectral multipoint confocal fluorescence microscope that was developed at the Eliceiri Lab.
Kevin Eliceiri, University of Wisconsin-Madison.
View Media

3656: Fruit fly ovary_2
3656: Fruit fly ovary_2
A fruit fly ovary, shown here, contains as many as 20 eggs. Fruit flies are not merely tiny insects that buzz around overripe fruit--they are a venerable scientific tool. Research on the flies has shed light on many aspects of human biology, including biological rhythms, learning, memory and neurodegenerative diseases. Another reason fruit flies are so useful in a lab (and so successful in fruit bowls) is that they reproduce rapidly. About three generations can be studied in a single month. Related to image 3607.
Denise Montell, University of California, Santa Barbara
View Media
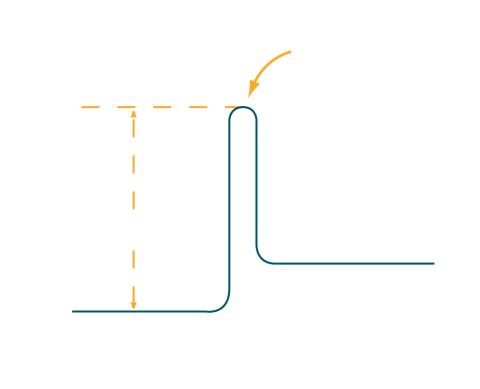
2525: Activation energy
2525: Activation energy
To become products, reactants must overcome an energy hill. See image 2526 for a labeled version of this illustration. Featured in The Chemistry of Health.
Crabtree + Company
View Media
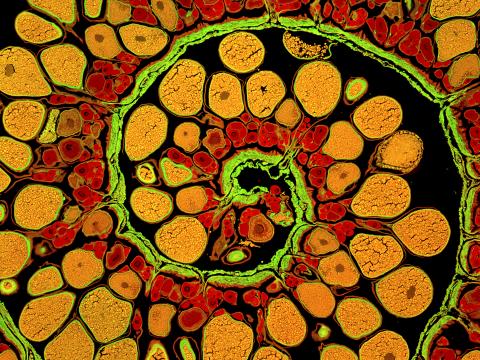
3620: Anglerfish ovary cross-section
3620: Anglerfish ovary cross-section
This image captures the spiral-shaped ovary of an anglerfish in cross-section. Once matured, these eggs will be released in a gelatinous, floating mass. For some species of anglerfish, this egg mass can be up to 3 feet long and include nearly 200,000 eggs.
This image was part of the Life: Magnified exhibit that ran from June 3, 2014, to January 21, 2015, at Dulles International Airport.
This image was part of the Life: Magnified exhibit that ran from June 3, 2014, to January 21, 2015, at Dulles International Airport.
James E. Hayden, The Wistar Institute, Philadelphia, Pa.
View Media

6344: Drosophila
6344: Drosophila
Two adult fruit flies (Drosophila)
Dr. Vicki Losick, MDI Biological Laboratory, www.mdibl.org
View Media
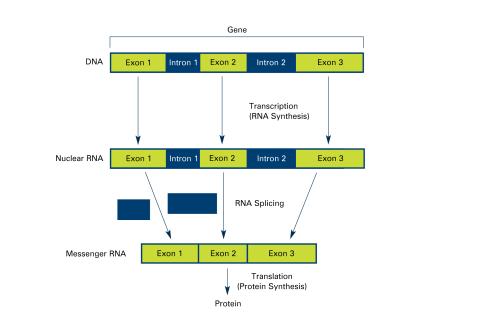
2551: Introns (with labels)
2551: Introns (with labels)
Genes are often interrupted by stretches of DNA (introns, blue) that do not contain instructions for making a protein. The DNA segments that do contain protein-making instructions are known as exons (green). See image 2550 for an unlabeled version of this illustration. Featured in The New Genetics.
Crabtree + Company
View Media
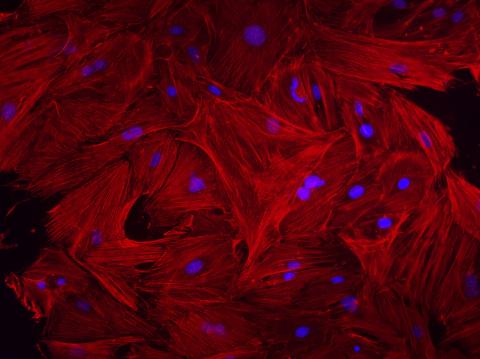
3281: Mouse heart fibroblasts
3281: Mouse heart fibroblasts
This image shows mouse fetal heart fibroblast cells. The muscle protein actin is stained red, and the cell nuclei are stained blue. The image was part of a study investigating stem cell-based approaches to repairing tissue damage after a heart attack. Image and caption information courtesy of the California Institute for Regenerative Medicine.
Kara McCloskey lab, University of California, Merced, via CIRM
View Media
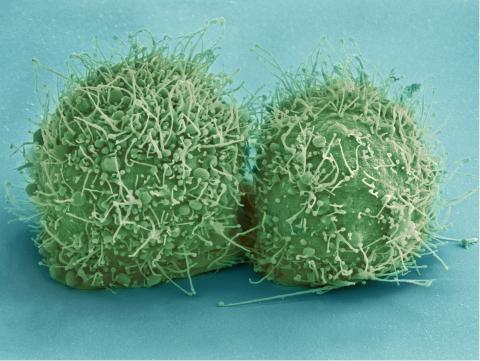
3518: HeLa cells
3518: HeLa cells
Scanning electron micrograph of just-divided HeLa cells. Zeiss Merlin HR-SEM. See related images 3519, 3520, 3521, 3522.
National Center for Microscopy and Imaging Research
View Media
3571: HIV-1 virus in the colon
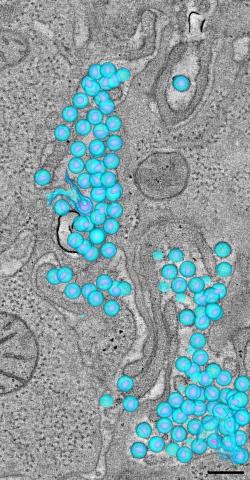
3571: HIV-1 virus in the colon
A tomographic reconstruction of the colon shows the location of large pools of HIV-1 virus particles (in blue) located in the spaces between adjacent cells. The purple objects within each sphere represent the conical cores that are one of the structural hallmarks of the HIV virus.
Mark Ladinsky, California Institute of Technology
View Media
2558: RNA interference
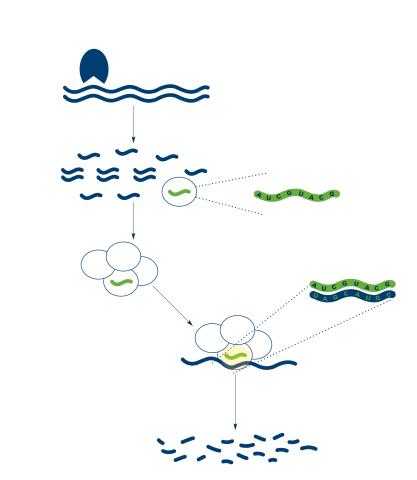
2558: RNA interference
RNA interference or RNAi is a gene-silencing process in which double-stranded RNAs trigger the destruction of specific RNAs. See 2559 for a labeled version of this illustration. Featured in The New Genetics.
Crabtree + Company
View Media
3426: Regeneration of Mouse Ears
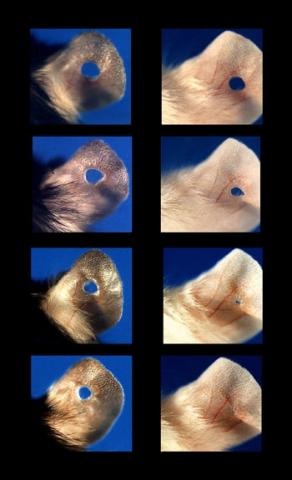
3426: Regeneration of Mouse Ears
Normal mice, like the B6 breed pictured on the left, develop scars when their ears are pierced. The Murphy Roths Large (MRL) mice pictured on the right can grow back lost ear tissue thanks to an inactive version of the p21 gene. When researchers knocked out that same gene in other mouse breeds, their ears also healed completely without scarring. Journal Article: Clark, L.D., Clark, R.K. and Heber-Katz, E. 1998. A new murine model for mammalian wound repair and regeneration. Clin Immunol Immunopathol 88: 35-45.
Ellen Heber-Katz, The Wistar Institute
View Media
2515: Life of an AIDS virus (with labels and stages)
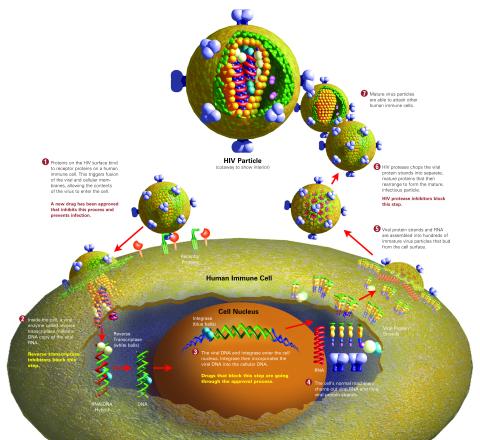
2515: Life of an AIDS virus (with labels and stages)
HIV is a retrovirus, a type of virus that carries its genetic material not as DNA but as RNA. Long before anyone had heard of HIV, researchers in labs all over the world studied retroviruses, tracing out their life cycle and identifying the key proteins the viruses use to infect cells. When HIV was identified as a retrovirus, these studies gave AIDS researchers an immediate jump-start. The previously identified viral proteins became initial drug targets. See images 2513 and 2514 for other versions of this illustration. Featured in The Structures of Life.
Crabtree + Company
View Media
2541: Nucleotides make up DNA
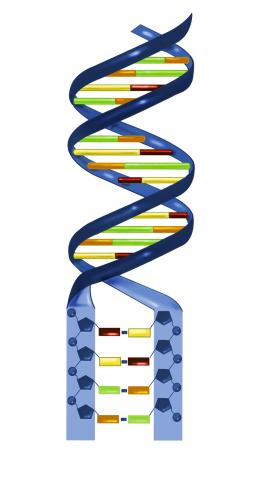
2541: Nucleotides make up DNA
DNA consists of two long, twisted chains made up of nucleotides. Each nucleotide contains one base, one phosphate molecule, and the sugar molecule deoxyribose. The bases in DNA nucleotides are adenine, thymine, cytosine, and guanine. See image 2542 for a labeled version of this illustration. Featured in The New Genetics.
Crabtree + Company
View Media
6465: CRISPR Illustration Frame 1
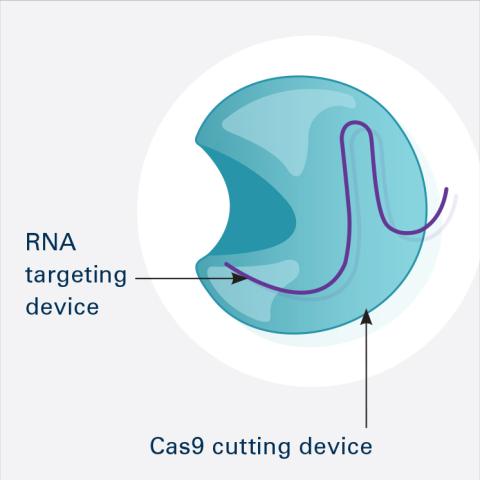
6465: CRISPR Illustration Frame 1
This illustration shows, in simplified terms, how the CRISPR-Cas9 system can be used as a gene-editing tool. This is the first frame in a series of four. The CRISPR system has two components joined together: a finely tuned targeting device (a small strand of RNA programmed to look for a specific DNA sequence) and a strong cutting device (an enzyme called Cas9 that can cut through a double strand of DNA).
For an explanation and overview of the CRISPR-Cas9 system, see the iBiology video, and find the full CRIPSR illustration here.
For an explanation and overview of the CRISPR-Cas9 system, see the iBiology video, and find the full CRIPSR illustration here.
National Institute of General Medical Sciences.
View Media
2517: ATP synthase
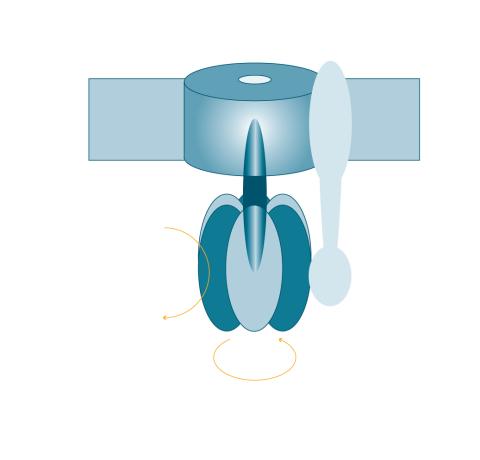
2517: ATP synthase
The world's smallest motor, ATP synthase, generates energy for the cell. See image 2518 for a labeled version of this illustration. Featured in The Chemistry of Health.
Crabtree + Company
View Media
2784: Microtubule dynamics in real time
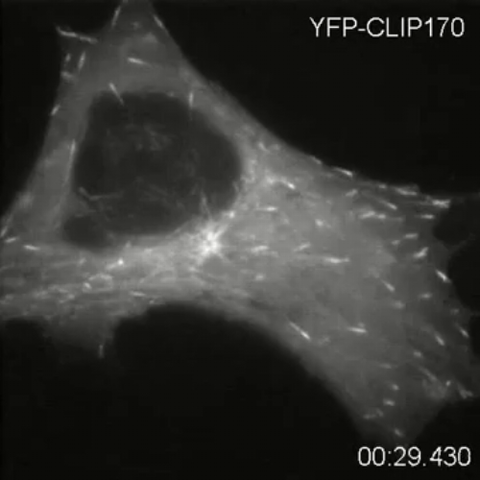
2784: Microtubule dynamics in real time
Cytoplasmic linker protein (CLIP)-170 is a microtubule plus-end-tracking protein that regulates microtubule dynamics and links microtubule ends to different intracellular structures. In this movie, the gene for CLIP-170 has been fused with green fluorescent protein (GFP). When the protein is expressed in cells, the activities can be monitored in real time. Here, you can see CLIP-170 streaming towards the edges of the cell.
Gary Borisy, Marine Biology Laboratory
View Media
1017: Lily mitosis 07
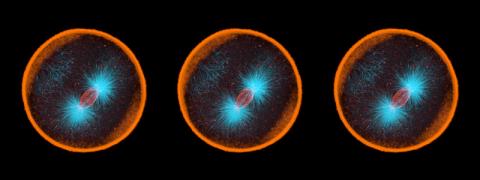
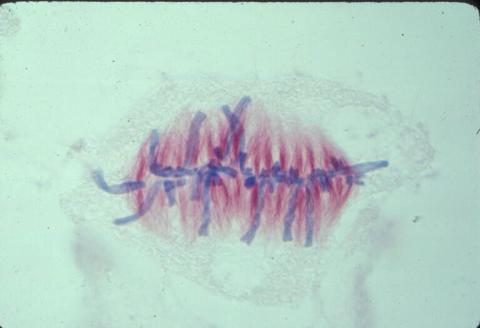
1017: Lily mitosis 07
A light microscope image of a cell from the endosperm of an African globe lily (Scadoxus katherinae). This is one frame of a time-lapse sequence that shows cell division in action. The lily is considered a good organism for studying cell division because its chromosomes are much thicker and easier to see than human ones. Staining shows microtubules in red and chromosomes in blue. Here, condensed chromosomes are clearly visible and have lined up in the middle of the dividing cell.
Related to images 1010, 1011, 1012, 1013, 1014, 1015, 1016, 1018, 1019, and 1021.
Related to images 1010, 1011, 1012, 1013, 1014, 1015, 1016, 1018, 1019, and 1021.
Andrew S. Bajer, University of Oregon, Eugene
View Media
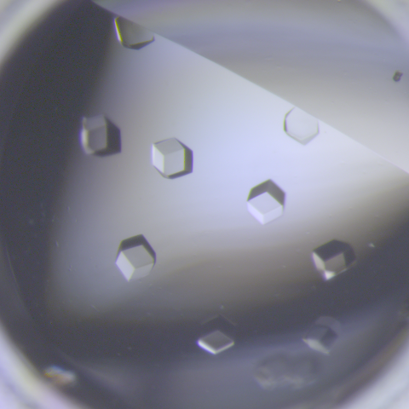

6764: Crystals of CCD-1 in complex with cefotaxime
6764: Crystals of CCD-1 in complex with cefotaxime
CCD-1 is an enzyme produced by the bacterium Clostridioides difficile that helps it resist antibiotics. Here, researchers crystallized bound pairs of CCD-1 molecules and molecules of the antibiotic cefotaxime. This enabled their structure to be studied using X-ray crystallography.
Related to images 6765, 6766, and 6767.
Related to images 6765, 6766, and 6767.
Keith Hodgson, Stanford University.
View Media

3772: The Proteasome: The Cell's Trash Processor in Action
3772: The Proteasome: The Cell's Trash Processor in Action
Our cells are constantly removing and recycling molecular waste. This video shows one way cells process their trash.
View Media

7011: Hawaiian bobtail squid
7011: Hawaiian bobtail squid
An adult Hawaiian bobtail squid, Euprymna scolopes, swimming next to a submerged hand.
Related to image 7010 and video 7012.
Related to image 7010 and video 7012.
Margaret J. McFall-Ngai, Carnegie Institution for Science/California Institute of Technology, and Edward G. Ruby, California Institute of Technology.
View Media
3408: Kluyveromyces polysporus Argonaute bound to guide RNA
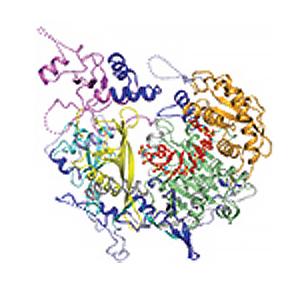
3408: Kluyveromyces polysporus Argonaute bound to guide RNA
A segment of siRNA, shown in red, guides a "slicer" protein called Argonaute (multi-colored twists and corkscrews) to the target RNA molecules.
Kotaro Nakanishi and David Weinberg, Massachusetts Institute of Technology
View Media
3364: Nociceptin/orphanin FQ peptide opioid receptor
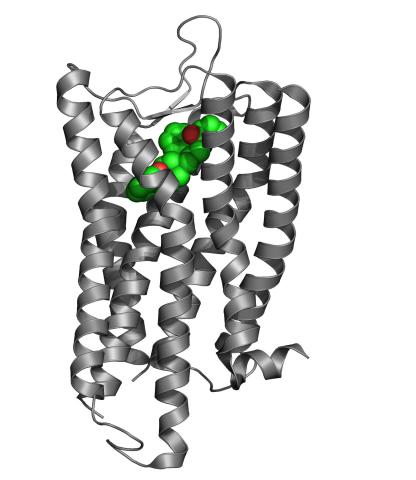
3364: Nociceptin/orphanin FQ peptide opioid receptor
The receptor is shown bound to an antagonist, compound-24
Raymond Stevens, The Scripps Research Institute
View Media
2536: G switch
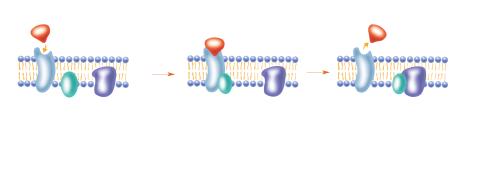
2536: G switch
The G switch allows our bodies to respond rapidly to hormones. See images 2537 and 2538 for labeled versions of this image. Featured in Medicines By Design.
Crabtree + Company
View Media
2442: Hydra 06
2442: Hydra 06
Hydra magnipapillata is an invertebrate animal used as a model organism to study developmental questions, for example the formation of the body axis.
Hiroshi Shimizu, National Institute of Genetics in Mishima, Japan
View Media
2528: A drug's life in the body (with labels)
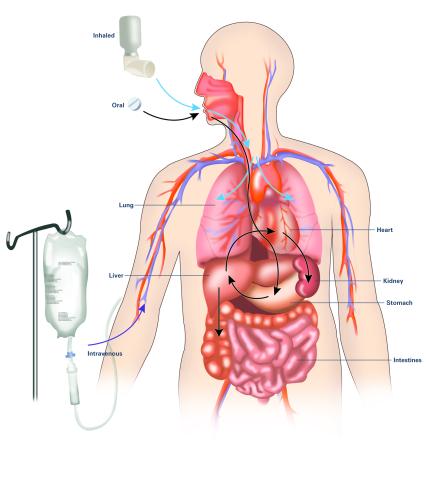
2528: A drug's life in the body (with labels)
A drug's life in the body. Medicines taken by mouth (oral) pass through the liver before they are absorbed into the bloodstream. Other forms of drug administration bypass the liver, entering the blood directly. See 2527 for an unlabeled version of this illustration. Featured in Medicines By Design.
Crabtree + Company
View Media
6612: Ciclo circadiano de un adolescente típico
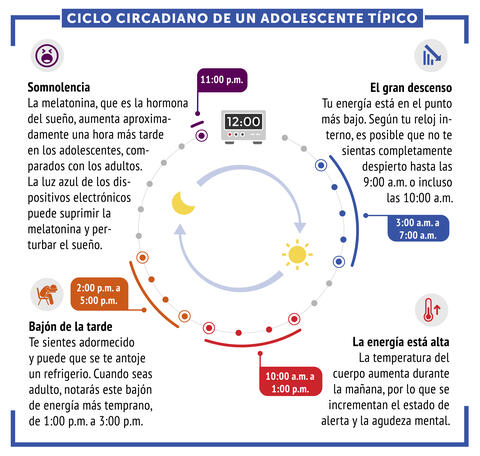
6612: Ciclo circadiano de un adolescente típico
Los ritmos circadianos son cambios físicos, mentales y conductuales que siguen un ciclo de 24 horas. Los ritmos circadianos típicos conducen a un nivel alto de energía durante la mitad del día (de 10 a.m. a 1 p.m.) y un bajón por la tarde. De noche, los ritmos circadianos hacen que la hormona melatonina aumente, lo que hace que la persona se sienta somnolienta.
Vea 6611 para la versión en inglés de esta infografía.
Vea 6611 para la versión en inglés de esta infografía.
NIGMS
View Media
6892: Microtubules and tau aggregates
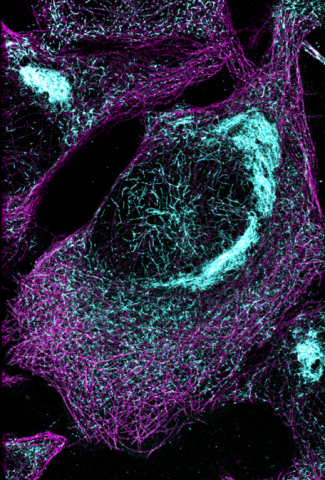
6892: Microtubules and tau aggregates
Microtubules (magenta) and tau protein (light blue) in a cell model of tauopathy. Researchers believe that tauopathy—the aggregation of tau protein—plays a role in Alzheimer’s disease and other neurodegenerative diseases. This image was captured using Stochastic Optical Reconstruction Microscopy (STORM).
Related to images 6889, 6890, and 6891.
Related to images 6889, 6890, and 6891.
Melike Lakadamyali, Perelman School of Medicine at the University of Pennsylvania.
View Media