Switch to List View
Image and Video Gallery
This is a searchable collection of scientific photos, illustrations, and videos. The images and videos in this gallery are licensed under Creative Commons Attribution Non-Commercial ShareAlike 3.0. This license lets you remix, tweak, and build upon this work non-commercially, as long as you credit and license your new creations under identical terms.
2437: Hydra 01
2437: Hydra 01
Hydra magnipapillata is an invertebrate animal used as a model organism to study developmental questions, for example the formation of the body axis.
Hiroshi Shimizu, National Institute of Genetics in Mishima, Japan
View Media
3746: Serum albumin structure 3
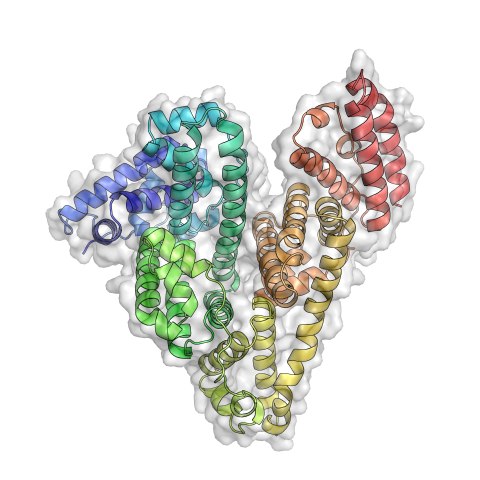
3746: Serum albumin structure 3
Serum albumin (SA) is the most abundant protein in the blood plasma of mammals. SA has a characteristic heart-shape structure and is a highly versatile protein. It helps maintain normal water levels in our tissues and carries almost half of all calcium ions in human blood. SA also transports some hormones, nutrients and metals throughout the bloodstream. Despite being very similar to our own SA, those from other animals can cause some mild allergies in people. Therefore, some scientists study SAs from humans and other mammals to learn more about what subtle structural or other differences cause immune responses in the body.
Related to entries 3744 and 3745.
Related to entries 3744 and 3745.
Wladek Minor, University of Virginia
View Media
3567: RSV-Infected Cell
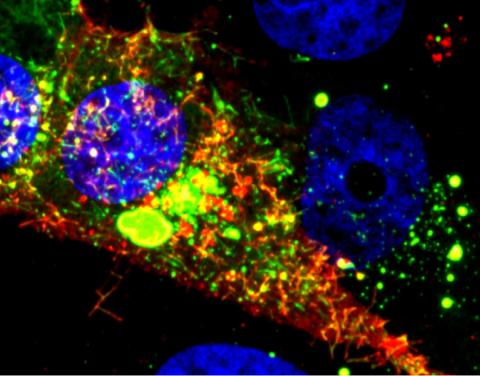
3567: RSV-Infected Cell
Viral RNA (red) in an RSV-infected cell. More information about the research behind this image can be found in a Biomedical Beat Blog posting from January 2014.
Eric Alonas and Philip Santangelo, Georgia Institute of Technology and Emory University
View Media

2531: Drugs enter skin
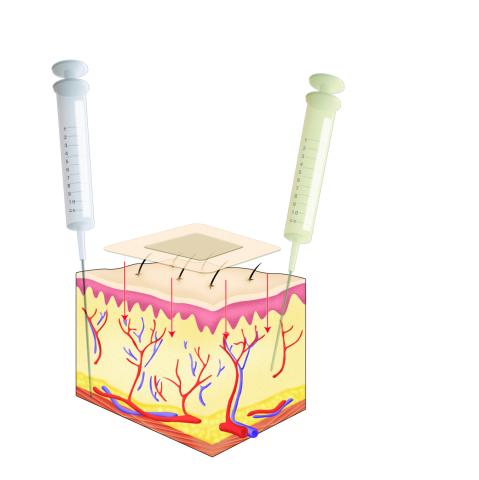
2531: Drugs enter skin
Drugs enter different layers of skin via intramuscular, subcutaneous, or transdermal delivery methods. See image 2532 for a labeled version of this illustration. Featured in Medicines By Design.
Crabtree + Company
View Media
6352: CRISPR surveillance complex
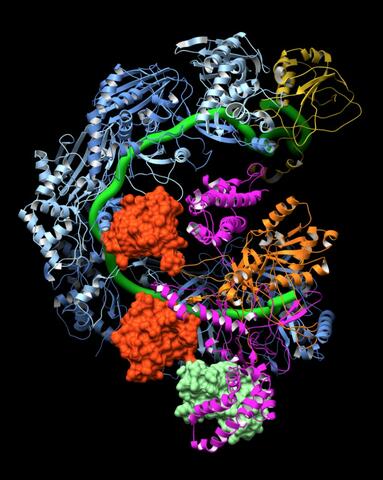
6352: CRISPR surveillance complex
This image shows how the CRISPR surveillance complex is disabled by two copies of anti-CRISPR protein AcrF1 (red) and one AcrF2 (light green). These anti-CRISPRs block access to the CRISPR RNA (green tube) preventing the surveillance complex from scanning and targeting invading viral DNA for destruction.
NRAMM National Resource for Automated Molecular Microscopy http://nramm.nysbc.org/nramm-images/ Source: Bridget Carragher
View Media
3333: Polarized cells- 02
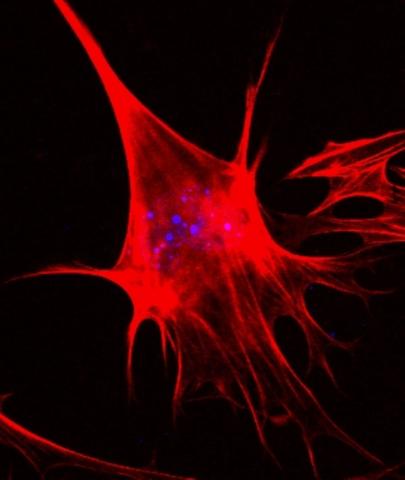
3333: Polarized cells- 02
Cells move forward with lamellipodia and filopodia supported by networks and bundles of actin filaments. Proper, controlled cell movement is a complex process. Recent research has shown that an actin-polymerizing factor called the Arp2/3 complex is the key component of the actin polymerization engine that drives amoeboid cell motility. ARPC3, a component of the Arp2/3 complex, plays a critical role in actin nucleation. In this photo, the ARPC3-/- fibroblast cells were fixed and stained with Alexa 546 phalloidin for F-actin (red) and DAPI to visualize the nucleus (blue). In the absence of functional Arp2/3 complex, ARPC3-/- fibroblast cells' leading edge morphology is significantly altered with filopodia-like structures. Related to images 3328, 3329, 3330, 3331, and 3332.
Rong Li and Praveen Suraneni, Stowers Institute for Medical Research
View Media
2561: Histones in chromatin (with labels)
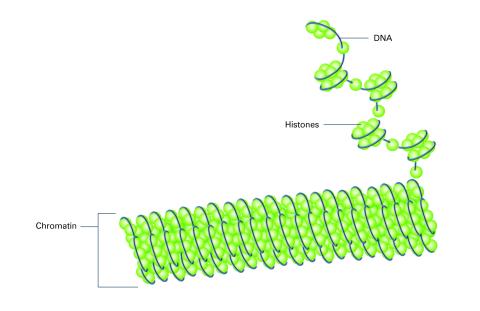
2561: Histones in chromatin (with labels)
Histone proteins loop together with double-stranded DNA to form a structure that resembles beads on a string. See image 2560 for an unlabeled version of this illustration. Featured in The New Genetics.
Crabtree + Company
View Media
5730: Dynamic cryo-EM model of the human transcription preinitiation complex
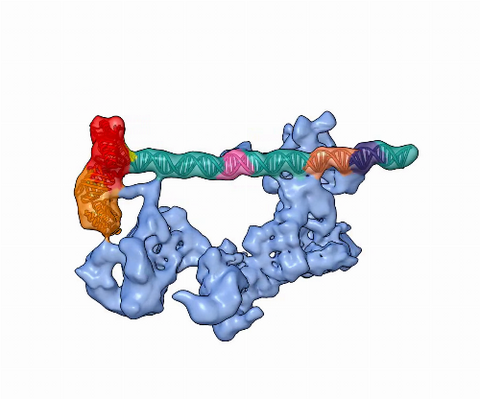
5730: Dynamic cryo-EM model of the human transcription preinitiation complex
Gene transcription is a process by which information encoded in DNA is transcribed into RNA. It's essential for all life and requires the activity of proteins, called transcription factors, that detect where in a DNA strand transcription should start. In eukaryotes (i.e., those that have a nucleus and mitochondria), a protein complex comprising 14 different proteins is responsible for sniffing out transcription start sites and starting the process. This complex represents the core machinery to which an enzyme, named RNA polymerase, can bind to and read the DNA and transcribe it to RNA. Scientists have used cryo-electron microscopy (cryo-EM) to visualize the TFIID-RNA polymerase-DNA complex in unprecedented detail. This animation shows the different TFIID components as they contact DNA and recruit the RNA polymerase for gene transcription.
To learn more about the research that has shed new light on gene transcription, see this news release from Berkeley Lab.
Related to image 3766.
To learn more about the research that has shed new light on gene transcription, see this news release from Berkeley Lab.
Related to image 3766.
Eva Nogales, Berkeley Lab
View Media
6776: Tracking cells in a gastrulating zebrafish embryo
6776: Tracking cells in a gastrulating zebrafish embryo
During development, a zebrafish embryo is transformed from a ball of cells into a recognizable body plan by sweeping convergence and extension cell movements. This process is called gastrulation. Each line in this video represents the movement of a single zebrafish embryo cell over the course of 3 hours. The video was created using time-lapse confocal microscopy. Related to image 6775.
Liliana Solnica-Krezel, Washington University School of Medicine in St. Louis.
View Media
2529: Aspirin
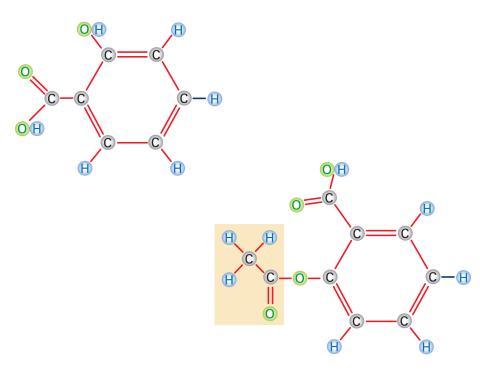
2529: Aspirin
Acetylsalicylate (bottom) is the aspirin of today. Adding a chemical tag called an acetyl group (shaded box, bottom) to a molecule derived from willow bark (salicylate, top) makes the molecule less acidic (and easier on the lining of the digestive tract), but still effective at relieving pain. See image 2530 for a labeled version of this illustration. Featured in Medicines By Design.
Crabtree + Company
View Media
5883: Beta-galactosidase montage showing cryo-EM improvement--gradient background
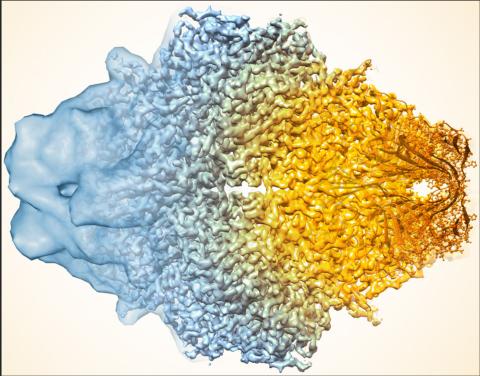
5883: Beta-galactosidase montage showing cryo-EM improvement--gradient background
Composite image of beta-galactosidase showing how cryo-EM’s resolution has improved dramatically in recent years. Older images to the left, more recent to the right. Related to image 5882. NIH Director Francis Collins featured this on his blog on January 14, 2016.
Veronica Falconieri, Sriram Subramaniam Lab, National Cancer Institute
View Media
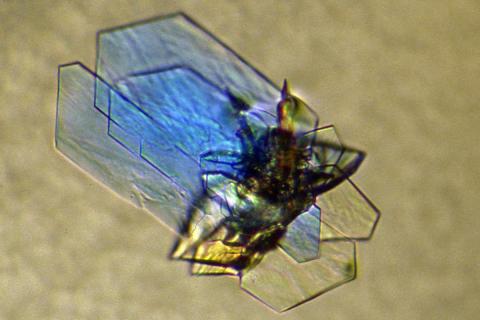

2410: DNase
2410: DNase
Crystals of DNase protein created for X-ray crystallography, which can reveal detailed, three-dimensional protein structures.
Alex McPherson, University of California, Irvine
View Media
3483: Chang Shan
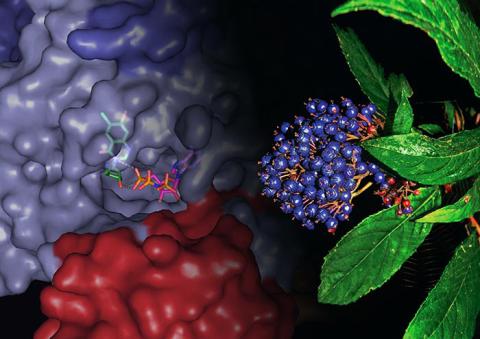
3483: Chang Shan
For thousands of years, Chinese herbalists have treated malaria using Chang Shan, a root extract from a type of hydrangea that grows in Tibet and Nepal. Recent studies have suggested Chang Shan can also reduce scar formation, treat multiple sclerosis and even slow cancer progression.
Paul Schimmel Lab, Scripps Research Institute
View Media
2506: Carbon building blocks
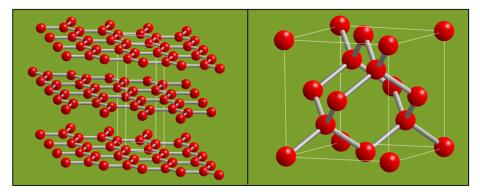
2506: Carbon building blocks
The arrangement of identical molecular components can make a dramatic difference. For example, carbon atoms can be arranged into dull graphite (left) or sparkly diamonds (right). See image 2507 for an illustration with examples.
Crabtree + Company
View Media
3617: Cells keep their shape with actin filaments and microtubules
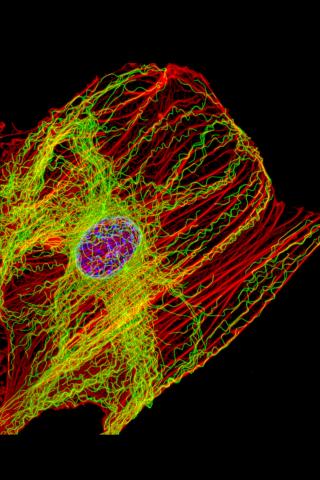
3617: Cells keep their shape with actin filaments and microtubules
This image shows a normal fibroblast, a type of cell that is common in connective tissue and frequently studied in research labs. This cell has a healthy skeleton composed of actin (red) and microtubles (green). Actin fibers act like muscles to create tension and microtubules act like bones to withstand compression.
This image was part of the Life: Magnified exhibit that ran from June 3, 2014, to January 21, 2015, at Dulles International Airport.
This image was part of the Life: Magnified exhibit that ran from June 3, 2014, to January 21, 2015, at Dulles International Airport.
James J. Faust and David G. Capco, Arizona State University
View Media
2330: Repairing DNA
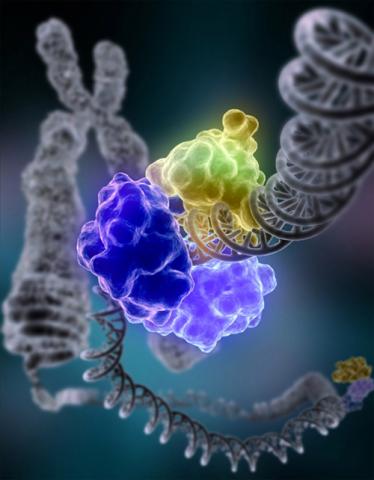
2330: Repairing DNA
Like a watch wrapped around a wrist, a special enzyme encircles the double helix to repair a broken strand of DNA. Without molecules that can mend such breaks, cells can malfunction, die, or become cancerous. Related to image 3493.
Tom Ellenberger, Washington University School of Medicine
View Media
7036: CRISPR Illustration
7036: CRISPR Illustration
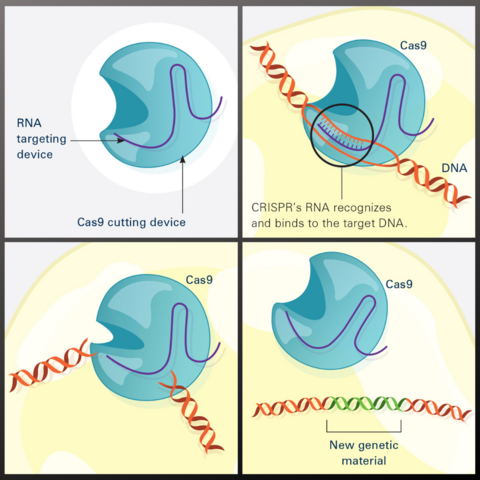
This illustration shows, in simplified terms, how the CRISPR-Cas9 system can be used as a gene-editing tool.
Frame 1 shows the two components of the CRISPR system: a strong cutting device (an enzyme called Cas9 that can cut through a double strand of DNA), and a finely tuned targeting device (a small strand of RNA programmed to look for a specific DNA sequence).
In frame 2, the CRISPR machine locates the target DNA sequence once inserted into a cell.
In frame 3, the Cas9 enzyme cuts both strands of the DNA.
Frame 4 shows a repaired DNA strand with new genetic material that researchers can introduce, which the cell automatically incorporates into the gap when it repairs the broken DNA.
For an explanation and overview of the CRISPR-Cas9 system, see the iBiology video.
Download the individual frames: Frame 1, Frame 2, Frame 3, and Frame 4.
Frame 1 shows the two components of the CRISPR system: a strong cutting device (an enzyme called Cas9 that can cut through a double strand of DNA), and a finely tuned targeting device (a small strand of RNA programmed to look for a specific DNA sequence).
In frame 2, the CRISPR machine locates the target DNA sequence once inserted into a cell.
In frame 3, the Cas9 enzyme cuts both strands of the DNA.
Frame 4 shows a repaired DNA strand with new genetic material that researchers can introduce, which the cell automatically incorporates into the gap when it repairs the broken DNA.
For an explanation and overview of the CRISPR-Cas9 system, see the iBiology video.
Download the individual frames: Frame 1, Frame 2, Frame 3, and Frame 4.
National Institute of General Medical Sciences.
View Media
3292: Centrioles anchor cilia in planaria
3292: Centrioles anchor cilia in planaria
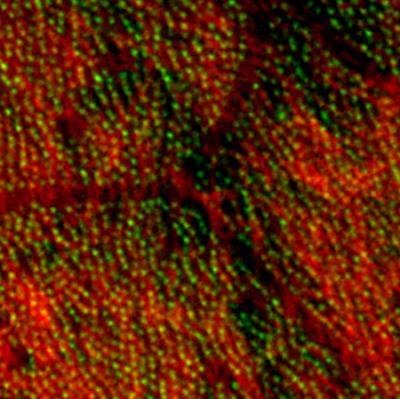
Centrioles (green) anchor cilia (red), which project on the surface of pharynx cells of the freshwater planarian Schmidtea mediterranea. Centrioles require cellular structures called centrosomes for assembly in other animal species, but this flatworm known for its regenerative ability was unexpectedly found to lack centrosomes. From a Stowers University news release.
Juliette Azimzadeh, University of California, San Francisco
View Media
3426: Regeneration of Mouse Ears
3426: Regeneration of Mouse Ears
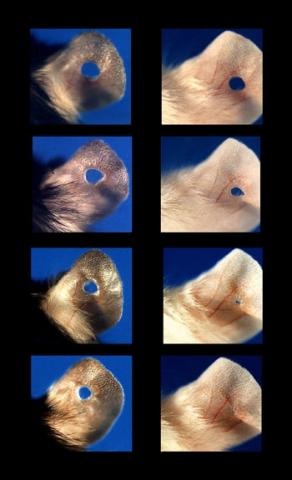
Normal mice, like the B6 breed pictured on the left, develop scars when their ears are pierced. The Murphy Roths Large (MRL) mice pictured on the right can grow back lost ear tissue thanks to an inactive version of the p21 gene. When researchers knocked out that same gene in other mouse breeds, their ears also healed completely without scarring. Journal Article: Clark, L.D., Clark, R.K. and Heber-Katz, E. 1998. A new murine model for mammalian wound repair and regeneration. Clin Immunol Immunopathol 88: 35-45.
Ellen Heber-Katz, The Wistar Institute
View Media

2714: Stretch detectors
2714: Stretch detectors
Muscles stretch and contract when we walk, and skin splits open and knits back together when we get a paper cut. To study these contractile forces, researchers built a three-dimensional scaffold that mimics tissue in an organism. Researchers poured a mixture of cells and elastic collagen over microscopic posts in a dish. Then they studied how the cells pulled and released the posts as they formed a web of tissue. To measure forces between posts, the researchers developed a computer model. Their findings--which show that contractile forces vary throughout the tissue--could have a wide range of medical applications.
Christopher Chen, University of Pennsylvania
View Media
3792: Nucleolus subcompartments spontaneously self-assemble 3
3792: Nucleolus subcompartments spontaneously self-assemble 3
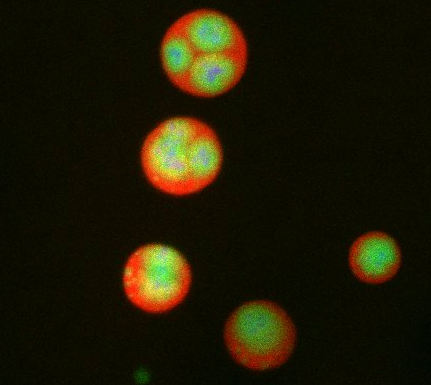
What looks a little like distant planets with some mysterious surface features are actually assemblies of proteins normally found in the cell's nucleolus, a small but very important protein complex located in the cell's nucleus. It forms on the chromosomes at the location where the genes for the RNAs are that make up the structure of the ribosome, the indispensable cellular machine that makes proteins from messenger RNAs.
However, how the nucleolus grows and maintains its structure has puzzled scientists for some time. It turns out that even though it looks like a simple liquid blob, it's rather well-organized, consisting of three distinct layers: the fibrillar center, where the RNA polymerase is active; the dense fibrillar component, which is enriched in the protein fibrillarin; and the granular component, which contains a protein called nucleophosmin. Researchers have now discovered that this multilayer structure of the nucleolus arises from differences in how the proteins in each compartment mix with water and with each other. These differences let the proteins readily separate from each other into the three nucleolus compartments.
This photo of nucleolus proteins in the eggs of a commonly used lab animal, the frog Xenopus laevis, shows each of the nucleolus compartments (the granular component is shown in red, the fibrillarin in yellow-green, and the fibrillar center in blue). The researchers have found that these compartments spontaneously fuse with each other on encounter without mixing with the other compartments.
For more details on this research, see this press release from Princeton. Related to video 3789, video 3791 and image 3793.
However, how the nucleolus grows and maintains its structure has puzzled scientists for some time. It turns out that even though it looks like a simple liquid blob, it's rather well-organized, consisting of three distinct layers: the fibrillar center, where the RNA polymerase is active; the dense fibrillar component, which is enriched in the protein fibrillarin; and the granular component, which contains a protein called nucleophosmin. Researchers have now discovered that this multilayer structure of the nucleolus arises from differences in how the proteins in each compartment mix with water and with each other. These differences let the proteins readily separate from each other into the three nucleolus compartments.
This photo of nucleolus proteins in the eggs of a commonly used lab animal, the frog Xenopus laevis, shows each of the nucleolus compartments (the granular component is shown in red, the fibrillarin in yellow-green, and the fibrillar center in blue). The researchers have found that these compartments spontaneously fuse with each other on encounter without mixing with the other compartments.
For more details on this research, see this press release from Princeton. Related to video 3789, video 3791 and image 3793.
Nilesh Vaidya, Princeton University
View Media

6609: 3D reconstruction of the Golgi apparatus in a pancreas cell
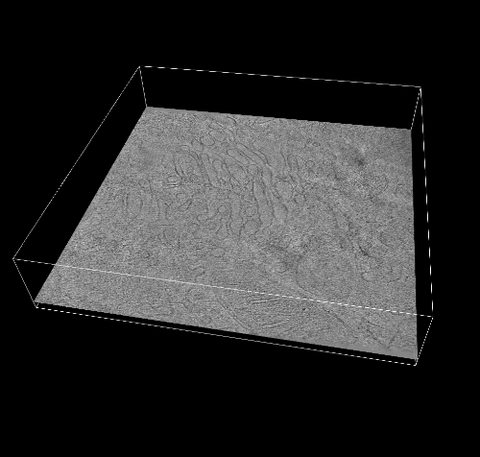
6609: 3D reconstruction of the Golgi apparatus in a pancreas cell
Researchers used cryo-electron tomography (cryo-ET) to capture images of a rat pancreas cell that were then compiled and color-coded to produce a 3D reconstruction. Visible features include the folded sacs of the Golgi apparatus (copper), transport vesicles (medium-sized dark-blue circles), microtubules (neon-green rods), a mitochondria membrane (pink), ribosomes (small pale-yellow circles), endoplasmic reticulum (aqua), and lysosomes (large yellowish-green circles). See 6606 for a still image from the video.
Xianjun Zhang, University of Southern California.
View Media
2601: Mouse liver labeled with fluorescent probe
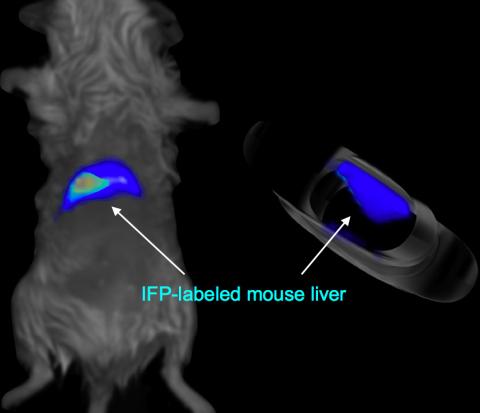
2601: Mouse liver labeled with fluorescent probe
A mouse liver glows after being tagged with specially designed infrared-fluorescent protein (IFP). Since its discovery in 1962, green fluorescent protein (GFP) has become an invaluable resource in biomedical imaging. But because of its short wavelength, the light that makes GFP glow doesn't penetrate far in whole animals. So University of California, San Diego cell biologist Roger Tsien--who shared the 2008 Nobel Prize in chemistry for groundbreaking work with GFP--made infrared-fluorescent proteins (IFPs) that shine under longer-wavelength light, allowing whole-body imaging in small animals.
Xiaokun Shu, University of California, San Diego
View Media
1272: Cytoskeleton
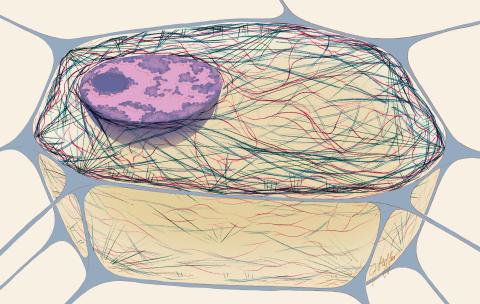
1272: Cytoskeleton
The three fibers of the cytoskeleton--microtubules in blue, intermediate filaments in red, and actin in green--play countless roles in the cell.
Judith Stoffer
View Media
1160: Vibrio bacteria
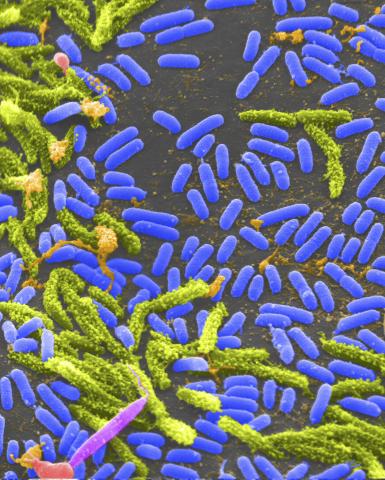
1160: Vibrio bacteria
Vibrio, a type (genus) of rod-shaped bacteria. Some Vibrio species cause cholera in humans.
Tina Weatherby Carvalho, University of Hawaii at Manoa
View Media
6346: Intasome
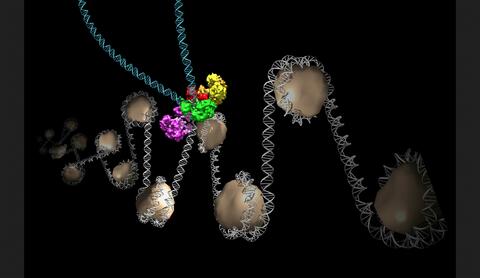
6346: Intasome
Salk researchers captured the structure of a protein complex called an intasome (center) that lets viruses similar to HIV establish permanent infection in their hosts. The intasome hijacks host genomic material, DNA (white) and histones (beige), and irreversibly inserts viral DNA (blue). The image was created by Jamie Simon and Dmitry Lyumkis. Work that led to the 3D map was published in: Ballandras-Colas A, Brown M, Cook NJ, Dewdney TG, Demeler B, Cherepanov P, Lyumkis D, & Engelman AN. (2016). Cryo-EM reveals a novel octameric integrase structure for ?-retroviral intasome function. Nature, 530(7590), 358—361
National Resource for Automated Molecular Microscopy http://nramm.nysbc.org/nramm-images/ Source: Bridget Carragher
View Media
2715: Glow-in-the-dark salamanders
2715: Glow-in-the-dark salamanders
These six-month-old axolotls, a kind of salamander, glow green and blue under ultraviolet light. That's because they were genetically modified to make harmless green fluorescent protein, or GFP. Like X-ray vision, GFP lets you see inside the axolotls as they hang out in their aquarium. GFP not only can reveal internal structures in living organisms, but it also can light up specific cells and even proteins within a cell. That allows scientists to identify and track things like cancer cells.
View Media
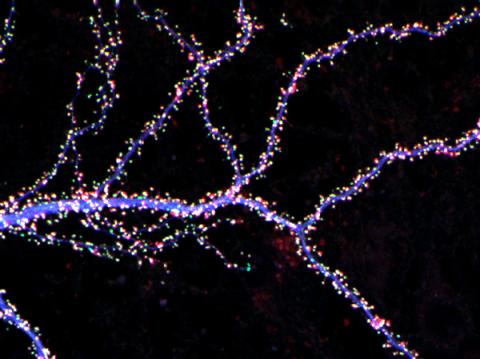
3686: Hippocampal neuron from rodent brain
3686: Hippocampal neuron from rodent brain
Hippocampal neuron from rodent brain with dendrites shown in blue. The hundreds of tiny magenta, green and white dots are the dendritic spines of excitatory synapses.
Shelley Halpain, UC San Diego
View Media

5758: Migrating pigment cells
5758: Migrating pigment cells
Pigment cells are cells that give skin its color. In fishes and amphibians, like frogs and salamanders, pigment cells are responsible for the characteristic skin patterns that help these organisms to blend into their surroundings or attract mates. The pigment cells are derived from neural crest cells, which are cells originating from the neural tube in the early embryo. This image shows neural crest cell-derived, migrating pigment cells in a salamander. Investigating pigment cell formation and migration in animals helps answer important fundamental questions about the factors that control pigmentation in the skin of animals, including humans. Related to images 5754, 5755, 5756 and 5757.
David Parichy, University of Washington
View Media
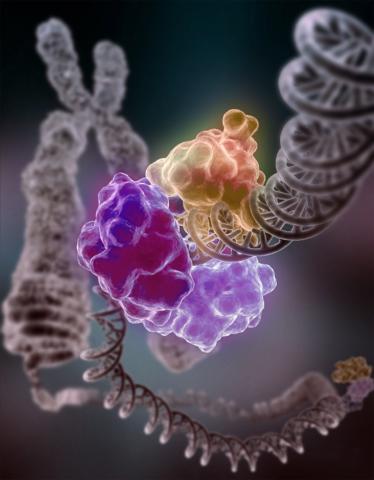
3493: Repairing DNA
3493: Repairing DNA
Like a watch wrapped around a wrist, a special enzyme encircles the double helix to repair a broken strand of DNA. Without molecules that can mend such breaks, cells can malfunction, die, or become cancerous. Related to image 2330.
Tom Ellenberger, Washington University School of Medicine
View Media

2385: Heat shock protein complex from Methanococcus jannaschii
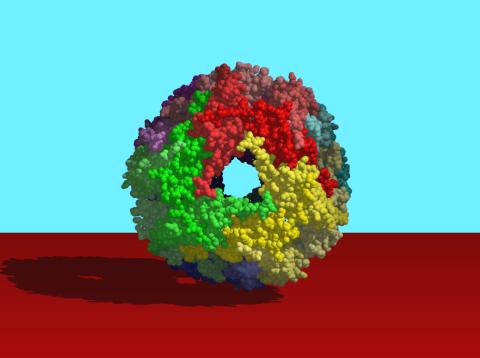
2385: Heat shock protein complex from Methanococcus jannaschii
Model based on X-ray crystallography of the structure of a small heat shock protein complex from the bacteria, Methanococcus jannaschii. Methanococcus jannaschii is an organism that lives at near boiling temperature, and this protein complex helps it cope with the stress of high temperature. Similar complexes are produced in human cells when they are "stressed" by events such as burns, heart attacks, or strokes. The complexes help cells recover from the stressful event.
Berkeley Structural Genomics Center, PSI-1
View Media
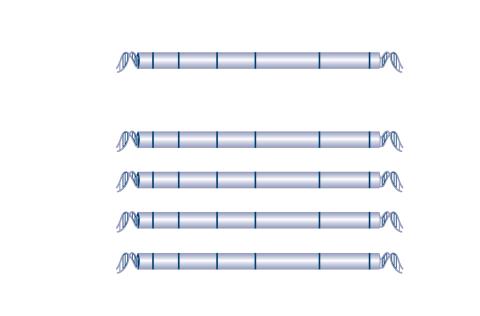
2566: Haplotypes
2566: Haplotypes
Haplotypes are combinations of gene variants that are likely to be inherited together within the same chromosomal region. In this example, an original haplotype (top) evolved over time to create three newer haplotypes that each differ by a few nucleotides (red). See image 2567 for a labeled version of this illustration. Featured in The New Genetics.
Crabtree + Company
View Media
2364: High-throughput protein structure determination pipeline
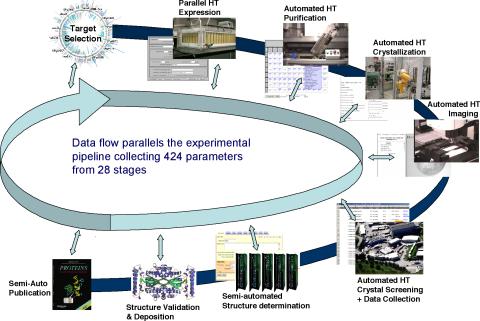
2364: High-throughput protein structure determination pipeline
This slide shows the technologies that the Joint Center for Structural Genomics developed for going from gene to structure and how the technologies have been integrated into a high-throughput pipeline, including all of the steps from target selection, parallel expression, protein purification, automated crystallization trials, automated crystal screening, structure determination, validation, and publication.
Joint Center for Structural Genomics
View Media
2337: Beta2-adrenergic receptor protein
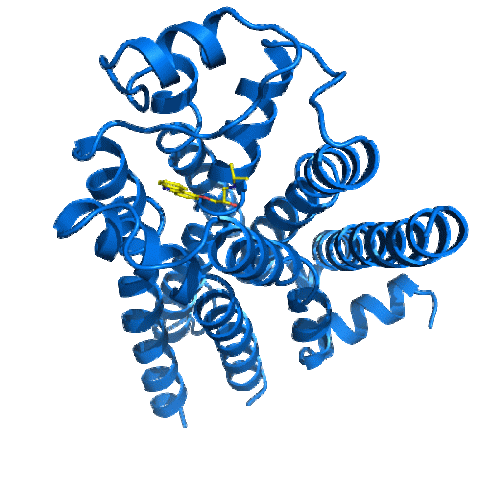
2337: Beta2-adrenergic receptor protein
Crystal structure of the beta2-adrenergic receptor protein. This is the first known structure of a human G protein-coupled receptor, a large family of proteins that control critical bodily functions and the action of about half of today's pharmaceuticals. Featured as one of the November 2007 Protein Structure Initiative Structures of the Month.
The Stevens Laboratory, The Scripps Research Institute
View Media
3613: Abnormal, spiky fibroblast
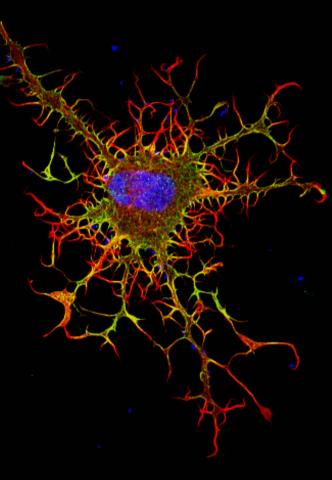
3613: Abnormal, spiky fibroblast
This is a fibroblast, a connective tissue cell that plays an important role in wound healing. Normal fibroblasts have smooth edges. In contrast, this spiky cell is missing a protein that is necessary for proper construction of the cell's skeleton. Its jagged shape makes it impossible for the cell to move normally. In addition to compromising wound healing, abnormal cell movement can lead to birth defects, faulty immune function, and other health problems.
This image was part of the Life: Magnified exhibit that ran from June 3, 2014, to January 21, 2015, at Dulles International Airport.
This image was part of the Life: Magnified exhibit that ran from June 3, 2014, to January 21, 2015, at Dulles International Airport.
Praveen Suraneni, Stowers Institute for Medical Research, Kansas City, Mo.
View Media
3574: Cytonemes in developing fruit fly cells
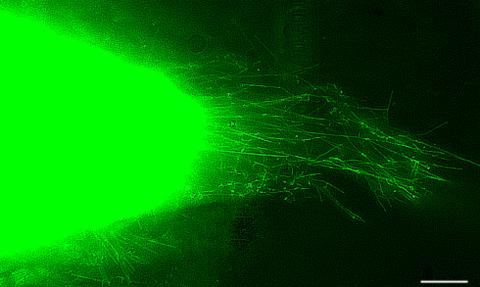
3574: Cytonemes in developing fruit fly cells
Scientists have long known that multicellular organisms use biological molecules produced by one cell and sensed by another to transmit messages that, for instance, guide proper development of organs and tissues. But it's been a puzzle as to how molecules dumped out into the fluid-filled spaces between cells can precisely home in on their targets. Using living tissue from fruit flies, a team led by Thomas Kornberg of the University of California, San Francisco, has shown that typical cells in animals can talk to each other via long, thin cell extensions called cytonemes (Latin for "cell threads") that may span the length of 50 or 100 cells. The point of contact between a cytoneme and its target cell acts as a communications bridge between the two cells.
Sougata Roy, University of California, San Francisco
View Media
2381: dUTP pyrophosphatase from M. tuberculosis
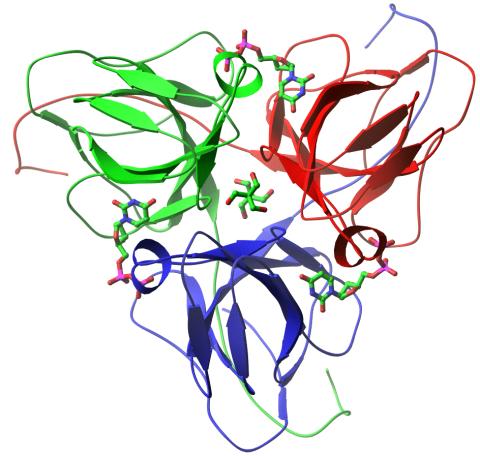
2381: dUTP pyrophosphatase from M. tuberculosis
Model of an enzyme, dUTP pyrophosphatase, from Mycobacterium tuberculosis. Drugs targeted to this enzyme might inhibit the replication of the bacterium that causes most cases of tuberculosis.
Mycobacterium Tuberculosis Center, PSI
View Media
6992: Molecular view of glutamatergic synapse
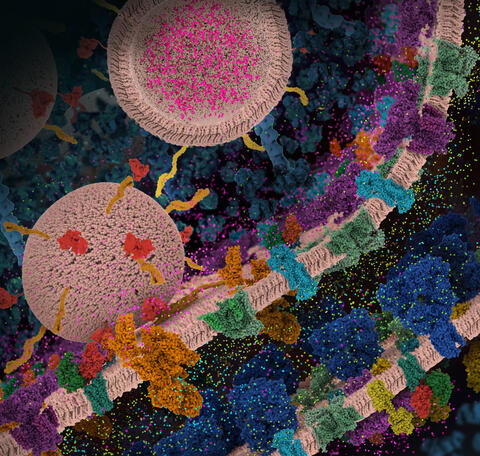
6992: Molecular view of glutamatergic synapse
This illustration highlights spherical pre-synaptic vesicles that carry the neurotransmitter glutamate. The presynaptic and postsynaptic membranes are shown with proteins relevant for transmitting and modulating the neuronal signal.
PDB 101’s Opioids and Pain Signaling video explains how glutamatergic synapses are involved in the process of pain signaling.
PDB 101’s Opioids and Pain Signaling video explains how glutamatergic synapses are involved in the process of pain signaling.
Amy Wu and Christine Zardecki, RCSB Protein Data Bank.
View Media
6753: Fruit fly nurse cells during egg development
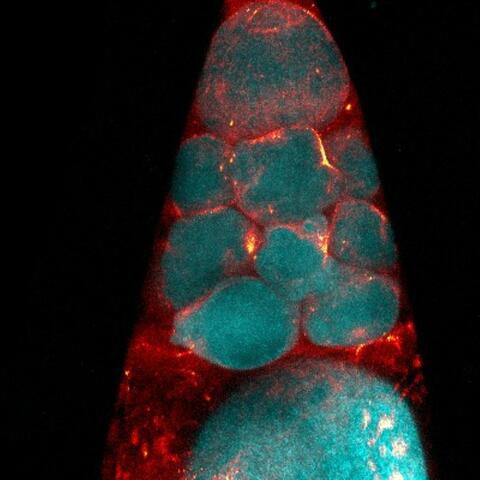
6753: Fruit fly nurse cells during egg development
In many animals, the egg cell develops alongside sister cells. These sister cells are called nurse cells in the fruit fly (Drosophila melanogaster), and their job is to “nurse” an immature egg cell, or oocyte. Toward the end of oocyte development, the nurse cells transfer all their contents into the oocyte in a process called nurse cell dumping. This process involves significant shape changes on the part of the nurse cells (blue), which are powered by wavelike activity of the protein myosin (red). This image was captured using a confocal laser scanning microscope. Related to video 6754.
Adam C. Martin, Massachusetts Institute of Technology.
View Media
3297: Four timepoints in gastrulation
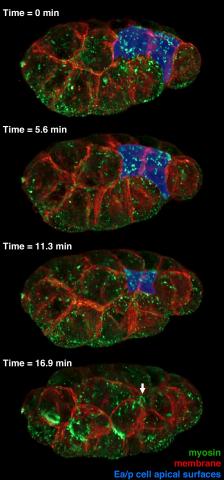
3297: Four timepoints in gastrulation
It has been said that gastrulation is the most important event in a person's life. This part of early embryonic development transforms a simple ball of cells and begins to define cell fate and the body axis. In a study published in Science magazine in March 2012, NIGMS grantee Bob Goldstein and his research group studied how contractions of actomyosin filaments in C. elegans and Drosophila embryos lead to dramatic rearrangements of cell and embryonic structure. This research is described in detail in the following article: "Triggering a Cell Shape Change by Exploiting Preexisting Actomyosin Contractions." In these images, myosin (green) and plasma membrane (red) are highlighted at four timepoints in gastrulation in the roundworm C. elegans. The blue highlights in the top three frames show how cells are internalized, and the site of closure around the involuting cells is marked with an arrow in the last frame. See related video 3334.
Bob Goldstein, University of North Carolina, Chapel Hill
View Media
2553: Alternative splicing (with labels)
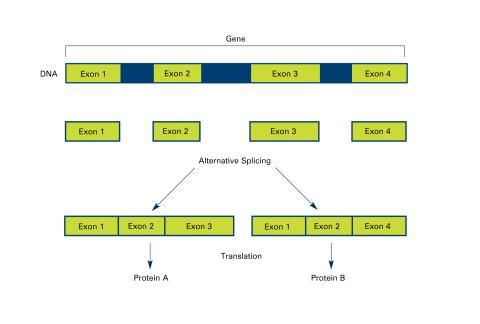
2553: Alternative splicing (with labels)
Arranging exons in different patterns, called alternative splicing, enables cells to make different proteins from a single gene. Featured in The New Genetics.
See image 2552 for an unlabeled version of this illustration.
See image 2552 for an unlabeled version of this illustration.
Crabtree + Company
View Media
7002: Plant resistosome
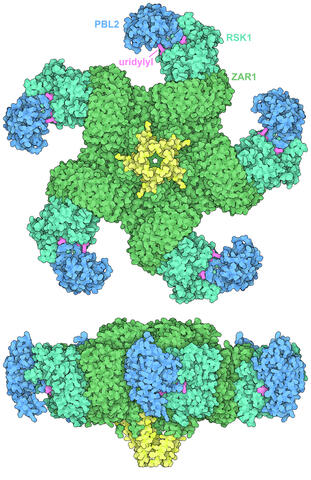
7002: Plant resistosome
The research organism Arabidopsis thaliana forms a large molecular machine called a resistosome to fight off infections. This illustration shows the top and side views of the fully-formed resistosome assembly (PDB entry 6J5T), composed of different proteins including one the plant uses as a decoy, PBL2 (dark blue), that gets uridylylated to begin the process of building the resistosome (uridylyl groups in magenta). Other proteins include RSK1 (turquoise) and ZAR1 (green) subunits. The ends of the ZAR1 subunits (yellow) form a funnel-like protrusion on one side of the assembly (seen in the side view). The funnel can carry out the critical protective function of the resistosome by inserting itself into the cell membrane to form a pore, which leads to a localized programmed cell death. The death of the infected cell helps protect the rest of the plant.
Amy Wu and Christine Zardecki, RCSB Protein Data Bank.
View Media
1276: Folding@Home
1276: Folding@Home
Stanford University scientist Vijay Pande decided to couple the power of computers with the help of the public. He initiated a project called Folding@Home, a so-called distributed computing project in which anyone who wants to can download a screensaver that performs protein-folding calculations when a computer is not in use. Folding@Home is modeled on a similar project called SETI@Home, which is used to search for extraterrestrial intelligence.
Judith Stoffer
View Media
3596: Heart rates time series image
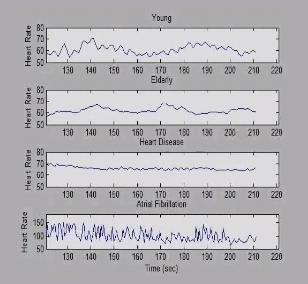
3596: Heart rates time series image
These time series show the heart rates of four different individuals. Automakers use steel scraps to build cars, construction companies repurpose tires to lay running tracks, and now scientists are reusing previously discarded medical data to better understand our complex physiology. Through a website called PhysioNet developed in part by Beth Israel Deaconess Medical Center cardiologist Ary Goldberger, scientists can access complete physiologic recordings, such as heart rate, respiration, brain activity and gait. They then can use free software to analyze the data and find patterns in it. The patterns could ultimately help health care professionals diagnose and treat health conditions like congestive heart failure, sleeping disorders, epilepsy and walking problems. PhysioNet is supported by NIH's National Institute of Biomedical Imaging and Bioengineering as well as by NIGMS.
Madalena Costa and Ary Goldberger, Beth Israel Deaconess Medical Center
View Media
6963: C. elegans trapped by carnivorous fungus
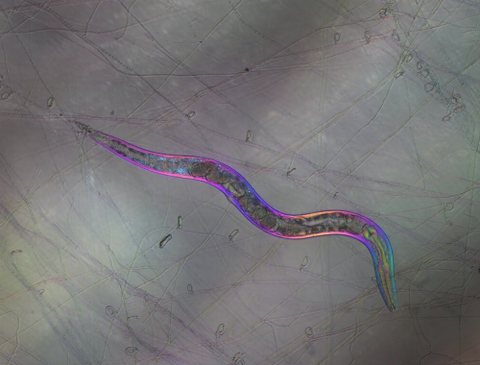
6963: C. elegans trapped by carnivorous fungus
Real-time footage of Caenorhabditis elegans, a tiny roundworm, trapped by a carnivorous fungus, Arthrobotrys dactyloides. This fungus makes ring traps in response to the presence of C. elegans. When a worm enters a ring, the trap rapidly constricts so that the worm cannot move away, and the fungus then consumes the worm. The size of the imaged area is 0.7mm x 0.9mm.
This video was obtained with a polychromatic polarizing microscope (PPM) in white light that shows the polychromatic birefringent image with hue corresponding to the slow axis orientation. More information about PPM can be found in the Scientific Reports paper “Polychromatic Polarization Microscope: Bringing Colors to a Colorless World” by Shribak.
This video was obtained with a polychromatic polarizing microscope (PPM) in white light that shows the polychromatic birefringent image with hue corresponding to the slow axis orientation. More information about PPM can be found in the Scientific Reports paper “Polychromatic Polarization Microscope: Bringing Colors to a Colorless World” by Shribak.
Michael Shribak, Marine Biological Laboratory/University of Chicago.
View Media

3530: Lorsch Swearing In
3530: Lorsch Swearing In
Jon Lorsch at his swearing in as NIGMS director in August 2013. Also shown are Francis Collins, NIH Director, and Judith Greenberg, former NIGMS Acting Director.
View Media
2684: Dicty fruit
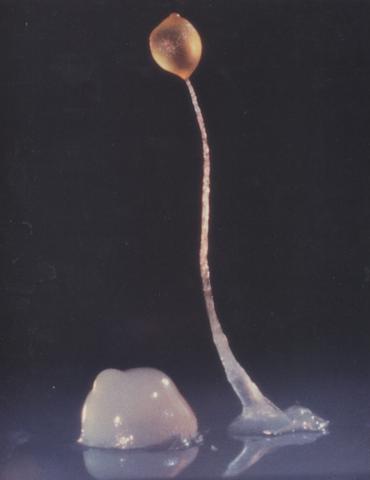
2684: Dicty fruit
Dictyostelium discoideum is a microscopic amoeba. A group of 100,000 form a mound as big as a grain of sand. Featured in The New Genetics.
View Media

3730: A molecular interaction network in yeast 1
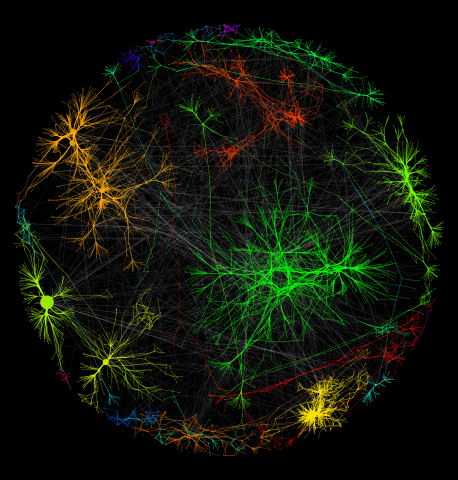
3730: A molecular interaction network in yeast 1
The image visualizes a part of the yeast molecular interaction network. The lines in the network represent connections among genes (shown as little dots) and different-colored networks indicate subnetworks, for instance, those in specific locations or pathways in the cell. Researchers use gene or protein expression data to build these networks; the network shown here was visualized with a program called Cytoscape. By following changes in the architectures of these networks in response to altered environmental conditions, scientists can home in on those genes that become central "hubs" (highly connected genes), for example, when a cell encounters stress. They can then further investigate the precise role of these genes to uncover how a cell's molecular machinery deals with stress or other factors. Related to images 3732 and 3733.
Keiichiro Ono, UCSD
View Media
2304: Bacteria working to eat
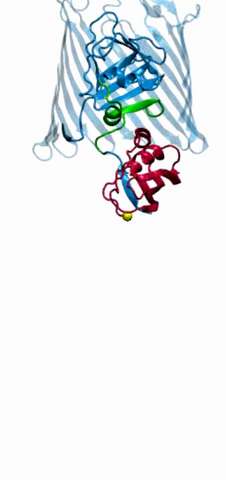
2304: Bacteria working to eat
Gram-negative bacteria perform molecular acrobatics just to eat. Because they're encased by two membranes, they must haul nutrients across both. To test one theory of how the bacteria manage this feat, researchers used computer simulations of two proteins involved in importing vitamin B12. Here, the protein (red) anchored in the inner membrane of bacteria tugs on a much larger protein (green and blue) in the outer membrane. Part of the larger protein unwinds, creating a pore through which the vitamin can pass.
Emad Tajkhorshid, University of Illinois at Urbana-Champaign
View Media
5765: Mitotic cell awaits chromosome alignment
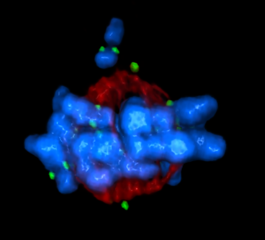
5765: Mitotic cell awaits chromosome alignment
During mitosis, spindle microtubules (red) attach to chromosome pairs (blue), directing them to the spindle equator. This midline alignment is critical for equal distribution of chromosomes in the dividing cell. Scientists are interested in how the protein kinase Plk1 (green) regulates this activity in human cells. Image is a volume projection of multiple deconvolved z-planes acquired with a Nikon widefield fluorescence microscope. This image was chosen as a winner of the 2016 NIH-funded research image call. Related to image 5766.
The research that led to this image was funded by NIGMS.
View Media
The research that led to this image was funded by NIGMS.
