Switch to List View
Image and Video Gallery
This is a searchable collection of scientific photos, illustrations, and videos. The images and videos in this gallery are licensed under Creative Commons Attribution Non-Commercial ShareAlike 3.0. This license lets you remix, tweak, and build upon this work non-commercially, as long as you credit and license your new creations under identical terms.
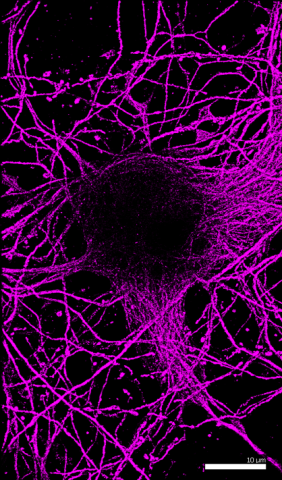
6890: Microtubules in hippocampal neurons
6890: Microtubules in hippocampal neurons
Microtubules (magenta) in neurons of the hippocampus, a part of the brain involved in learning and memory. Microtubules are strong, hollow fibers that provide structural support to cells. This image was captured using Stochastic Optical Reconstruction Microscopy (STORM).
Related to images 6889, 6891, and 6892.
Related to images 6889, 6891, and 6892.
Melike Lakadamyali, Perelman School of Medicine at the University of Pennsylvania.
View Media
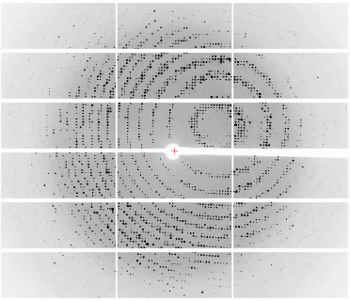
6765: X-ray diffraction pattern from a crystallized cefotaxime-CCD-1 complex
6765: X-ray diffraction pattern from a crystallized cefotaxime-CCD-1 complex
CCD-1 is an enzyme produced by the bacterium Clostridioides difficile that helps it resist antibiotics. Researchers crystallized complexes where a CCD-1 molecule and a molecule of the antibiotic cefotaxime were bound together. Then, they shot X-rays at the complexes to determine their structure—a process known as X-ray crystallography. This image shows the X-ray diffraction pattern of a complex.
Related to images 6764, 6766, and 6767.
Related to images 6764, 6766, and 6767.
Keith Hodgson, Stanford University.
View Media
1087: Natcher Building 07
1087: Natcher Building 07
NIGMS staff are located in the Natcher Building on the NIH campus.
Alisa Machalek, National Institute of General Medical Sciences
View Media
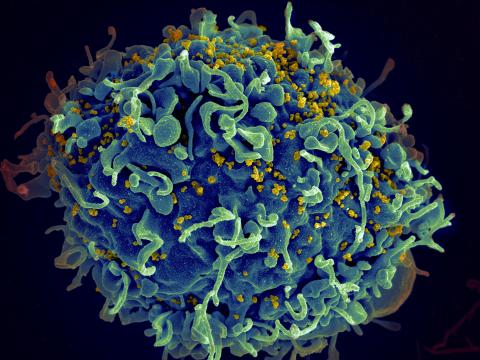
3638: HIV, the AIDS virus, infecting a human cell
3638: HIV, the AIDS virus, infecting a human cell
This human T cell (blue) is under attack by HIV (yellow), the virus that causes AIDS. The virus specifically targets T cells, which play a critical role in the body's immune response against invaders like bacteria and viruses.
This image was part of the Life: Magnified exhibit that ran from June 3, 2014, to January 21, 2015, at Dulles International Airport.
This image was part of the Life: Magnified exhibit that ran from June 3, 2014, to January 21, 2015, at Dulles International Airport.
Seth Pincus, Elizabeth Fischer, and Austin Athman, National Institute of Allergy and Infectious Diseases, National Institutes of Health
View Media
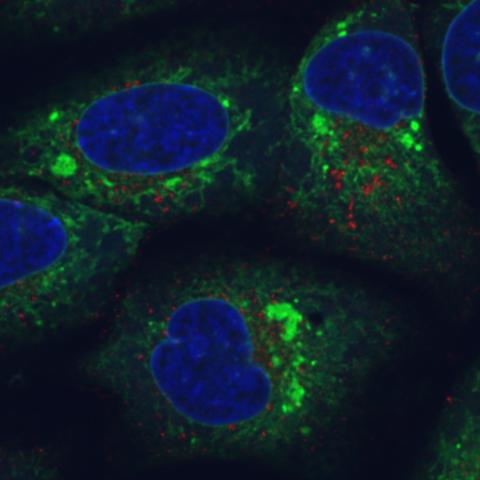
6773: Endoplasmic reticulum abnormalities
6773: Endoplasmic reticulum abnormalities
Human cells with the gene that codes for the protein FIT2 deleted. Green indicates an endoplasmic reticulum (ER) resident protein. The lack of FIT2 affected the structure of the ER and caused the resident protein to cluster in ER membrane aggregates, seen as large, bright-green spots. Red shows where the degradation of cell parts—called autophagy—is taking place, and the nucleus is visible in blue. This image was captured using a confocal microscope.
Michel Becuwe, Harvard University.
View Media
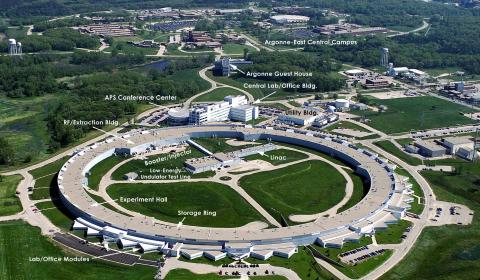
2358: Advanced Photon Source (APS) at Argonne National Lab
2358: Advanced Photon Source (APS) at Argonne National Lab
The intense X-rays produced by synchrotrons such as the Advanced Photon Source are ideally suited for protein structure determination. Using synchrotron X-rays and advanced computers scientists can determine protein structures at a pace unheard of decades ago.
Southeast Collaboratory for Structural Genomics
View Media
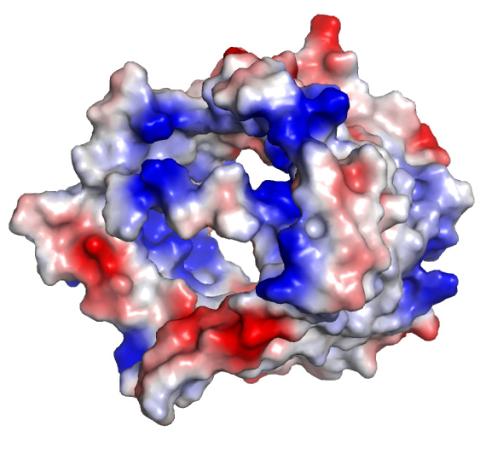
2494: VDAC-1 (3)
2494: VDAC-1 (3)
The structure of the pore-forming protein VDAC-1 from humans. This molecule mediates the flow of products needed for metabolism--in particular the export of ATP--across the outer membrane of mitochondria, the power plants for eukaryotic cells. VDAC-1 is involved in metabolism and the self-destruction of cells--two biological processes central to health.
Related to images 2491, 2495, and 2488.
Related to images 2491, 2495, and 2488.
Gerhard Wagner, Harvard Medical School
View Media

5815: Introduction to Genome Editing Using CRISPR/Cas9
5815: Introduction to Genome Editing Using CRISPR/Cas9
Genome editing using CRISPR/Cas9 is a rapidly expanding field of scientific research with emerging applications in disease treatment, medical therapeutics and bioenergy, just to name a few. This technology is now being used in laboratories all over the world to enhance our understanding of how living biological systems work, how to improve treatments for genetic diseases and how to develop energy solutions for a better future.
Janet Iwasa
View Media
6983: Genetic mosaicism in fruit flies
6983: Genetic mosaicism in fruit flies
Fat tissue from the abdomen of a genetically mosaic adult fruit fly. Genetic mosaicism means that the fly has cells with different genotypes even though it formed from a single zygote. This specific mosaicism results in accumulation of a critical fly adipokine (blue-green) within the fat tissue cells that have reduced expression a key nutrient sensing gene (in left panel). The dotted line shows the cells lacking the gene that is present and functioning in the rest of the cells. Nuclei are labelled in magenta. This image was captured using a confocal microscope and shows a maximum intensity projection of many slices.
Related to images 6982, 6984, and 6985.
Related to images 6982, 6984, and 6985.
Akhila Rajan, Fred Hutchinson Cancer Center
View Media
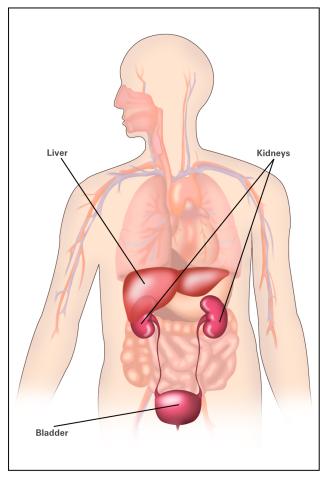
2497: Body toxins (with labels)
2497: Body toxins (with labels)
Body organs such as the liver and kidneys process chemicals and toxins. These "target" organs are susceptible to damage caused by these substances. See image 2496 for an unlabeled version of this illustration.
Crabtree + Company
View Media
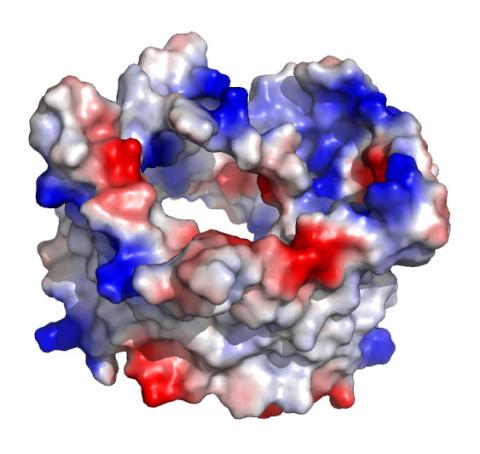
2488: VDAC-1 (1)
2488: VDAC-1 (1)
The structure of the pore-forming protein VDAC-1 from humans. This molecule mediates the flow of products needed for metabolism--in particular the export of ATP--across the outer membrane of mitochondria, the power plants for eukaryotic cells. VDAC-1 is involved in metabolism and the self-destruction of cells--two biological processes central to health.
Related to images 2491, 2494, and 2495.
Related to images 2491, 2494, and 2495.
Gerhard Wagner, Harvard Medical School
View Media
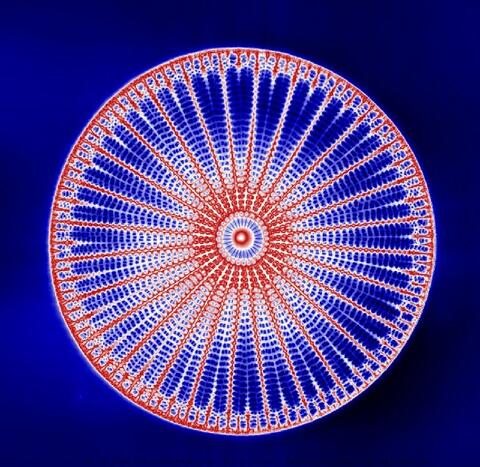

6902: Arachnoidiscus diatom
6902: Arachnoidiscus diatom
An Arachnoidiscus diatom with a diameter of 190µm. Diatoms are microscopic algae that have cell walls made of silica, which is the strongest known biological material relative to its density. In Arachnoidiscus, the cell wall is a radially symmetric pillbox-like shell composed of overlapping halves that contain intricate and delicate patterns. Sometimes, Arachnoidiscus is called “a wheel of glass.”
This image was taken with the orientation-independent differential interference contrast microscope.
This image was taken with the orientation-independent differential interference contrast microscope.
Michael Shribak, Marine Biological Laboratory/University of Chicago.
View Media
1157: Streptococcus bacteria
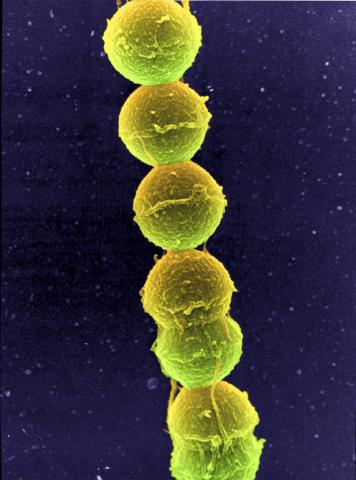
1157: Streptococcus bacteria
Image of Streptococcus, a type (genus) of spherical bacteria that can colonize the throat and back of the mouth. Stroptococci often occur in pairs or in chains, as shown here.
Tina Weatherby Carvalho, University of Hawaii at Manoa
View Media
3736: Transmission electron microscopy of myelinated axons with ECM between the axons
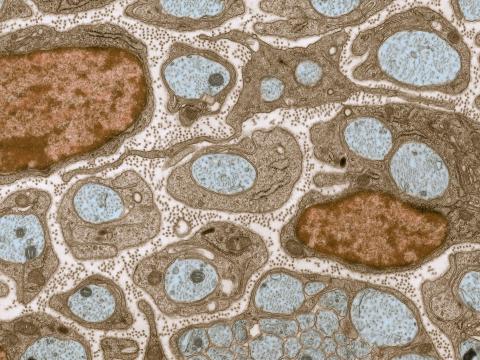
3736: Transmission electron microscopy of myelinated axons with ECM between the axons
The extracellular matrix (ECM) is most prevalent in connective tissues but also is present between the stems (axons) of nerve cells, as shown here. Blue-colored nerve cell axons are surrounded by brown-colored, myelin-supplying Schwann cells, which act like insulation around an electrical wire to help speed the transmission of electric nerve impulses down the axon. The ECM is pale pink. The tiny brown spots within it are the collagen fibers that are part of the ECM.
Tom Deerinck, National Center for Microscopy and Imaging Research (NCMIR)
View Media

3689: Computer sketch of bird-and-flower DNA origami
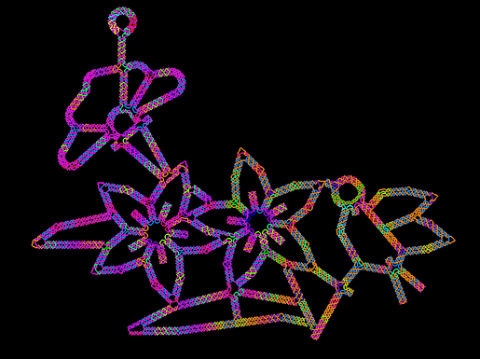
3689: Computer sketch of bird-and-flower DNA origami
A computer-generated sketch of a DNA origami folded into a flower-and-bird structure. See also related image 3690.
Hao Yan, Arizona State University
View Media
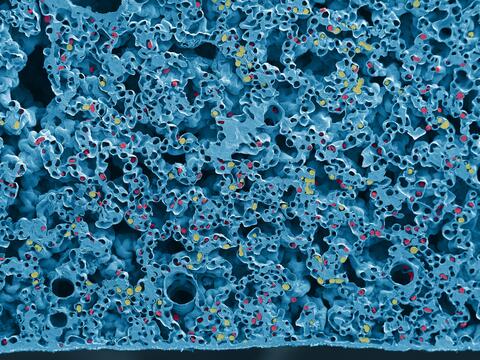
6389: Red and white blood cells in the lung
6556: Floral pattern in a mixture of two bacterial species, Acinetobacter baylyi and Escherichia coli, grown on a semi-solid agar for 72 hour
6556: Floral pattern in a mixture of two bacterial species, Acinetobacter baylyi and Escherichia coli, grown on a semi-solid agar for 72 hour
Floral pattern emerging as two bacterial species, motile Acinetobacter baylyi and non-motile Escherichia coli (green), are grown together for 72 hours on 0.5% agar surface from a small inoculum in the center of a Petri dish.
See 6557 for a photo of this process at 24 hours on 0.75% agar surface.
See 6553 for a photo of this process at 48 hours on 1% agar surface.
See 6555 for another photo of this process at 48 hours on 1% agar surface.
See 6550 for a video of this process.
See 6557 for a photo of this process at 24 hours on 0.75% agar surface.
See 6553 for a photo of this process at 48 hours on 1% agar surface.
See 6555 for another photo of this process at 48 hours on 1% agar surface.
See 6550 for a video of this process.
L. Xiong et al, eLife 2020;9: e48885
View Media
3253: Pulsating response to stress in bacteria
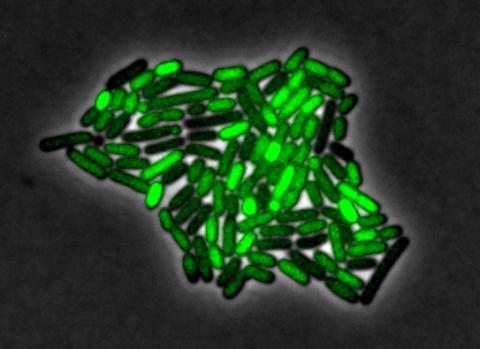
3253: Pulsating response to stress in bacteria
By attaching fluorescent proteins to the genetic circuit responsible for B. subtilis's stress response, researchers can observe the cells' pulses as green flashes. In response to a stressful environment like one lacking food, B. subtilis activates a large set of genes that help it respond to the hardship. Instead of leaving those genes on as previously thought, researchers discovered that the bacteria flip the genes on and off, increasing the frequency of these pulses with increasing stress. See entry 3254 for the related video.
Michael Elowitz, Caltech University
View Media
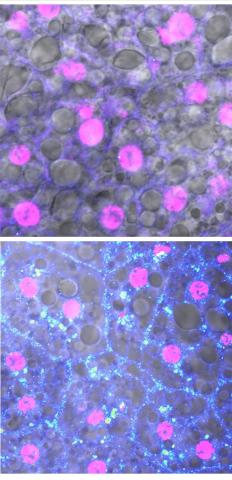
6984: Fruit fly starvation leads to adipokine accumulation
6984: Fruit fly starvation leads to adipokine accumulation
Adult Drosophila abdominal fat tissue showing cell nuclei labelled in magenta. The upper panel is from well-fed flies, and the lower panel is from flies that have been deprived of food for 4 hours. Starvation results in the accumulation of a key adipokine—a fat hormone (blue-green dots).
Related to images 6982, 6983, and 6985.
Related to images 6982, 6983, and 6985.
Akhila Rajan, Fred Hutchinson Cancer Center
View Media
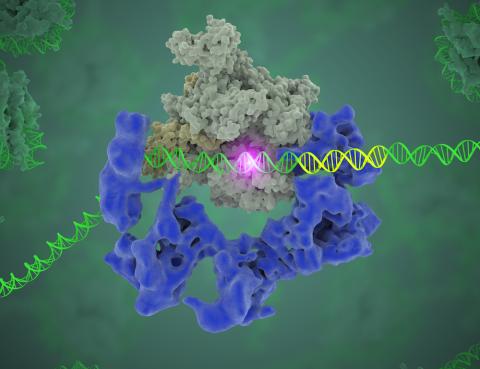
3766: TFIID complex binds DNA to start gene transcription
3766: TFIID complex binds DNA to start gene transcription
Gene transcription is a process by which the genetic information encoded in DNA is transcribed into RNA. It's essential for all life and requires the activity of proteins, called transcription factors, that detect where in a DNA strand transcription should start. In eukaryotes (i.e., those that have a nucleus and mitochondria), a protein complex comprising 14 different proteins is responsible for sniffing out transcription start sites and starting the process. This complex, called TFIID, represents the core machinery to which an enzyme, named RNA polymerase, can bind to and read the DNA and transcribe it to RNA. Scientists have used cryo-electron microscopy (cryo-EM) to visualize the TFIID-RNA polymerase-DNA complex in unprecedented detail. In this illustration, TFIID (blue) contacts the DNA and recruits the RNA polymerase (gray) for gene transcription. The start of the transcribed gene is shown with a flash of light. To learn more about the research that has shed new light on gene transcription, see this news release from Berkeley Lab. Related to video 5730.
Eva Nogales, Berkeley Lab
View Media
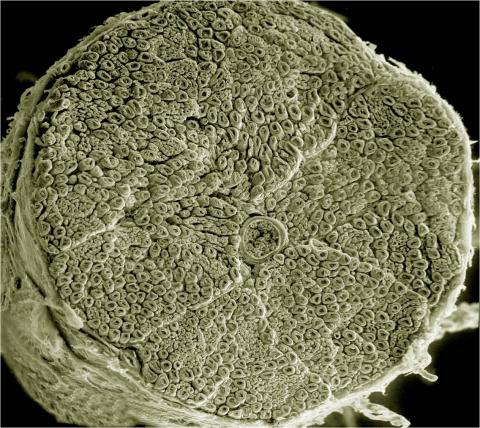
3387: NCMIR human spinal nerve
3387: NCMIR human spinal nerve
Spinal nerves are part of the peripheral nervous system. They run within the spinal column to carry nerve signals to and from all parts of the body. The spinal nerves enable all the movements we do, from turning our heads to wiggling our toes, control the movements of our internal organs, such as the colon and the bladder, as well as allow us to feel touch and the location of our limbs.
Tom Deerinck, National Center for Microscopy and Imaging Research (NCMIR)
View Media
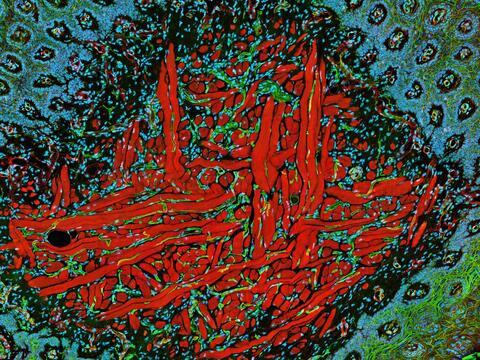
5811: NCMIR Tongue 2
5811: NCMIR Tongue 2
Microscopy image of a tongue. One in a series of two, see image 5810
National Center for Microscopy and Imaging Research (NCMIR)
View Media
1056: Skin cross-section
1056: Skin cross-section
Cross-section of skin anatomy shows layers and different tissue types.
National Institutes of Health Medical Arts
View Media
3332: Polarized cells- 01
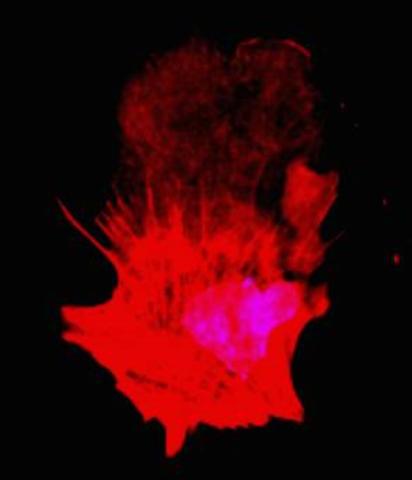
3332: Polarized cells- 01
Cells move forward with lamellipodia and filopodia supported by networks and bundles of actin filaments. Proper, controlled cell movement is a complex process. Recent research has shown that an actin-polymerizing factor called the Arp2/3 complex is the key component of the actin polymerization engine that drives amoeboid cell motility. ARPC3, a component of the Arp2/3 complex, plays a critical role in actin nucleation. In this photo, the ARPC3+/+ fibroblast cells were fixed and stained with Alexa 546 phalloidin for F-actin (red) and DAPI to visualize the nucleus (blue). ARPC3+/+ fibroblast cells with lamellipodia leading edge. Related to images 3328, 3329, 3330, 3331, and 3333.
Rong Li and Praveen Suraneni, Stowers Institute for Medical Research
View Media
3637: Purkinje cells are one of the main cell types in the brain
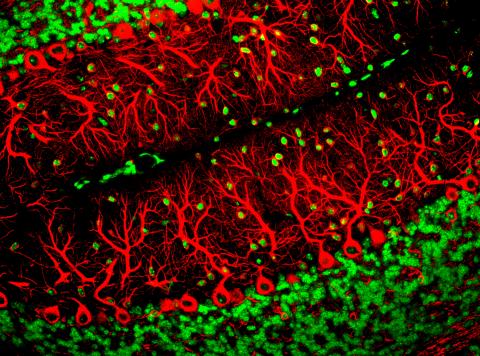
3637: Purkinje cells are one of the main cell types in the brain
This image captures Purkinje cells (red), one of the main types of nerve cell found in the brain. These cells have elaborate branching structures called dendrites that receive signals from other nerve cells.
This image was part of the Life: Magnified exhibit that ran from June 3, 2014, to January 21, 2015, at Dulles International Airport.
This image was part of the Life: Magnified exhibit that ran from June 3, 2014, to January 21, 2015, at Dulles International Airport.
Yinghua Ma and Timothy Vartanian, Cornell University, Ithaca, N.Y.
View Media
3571: HIV-1 virus in the colon
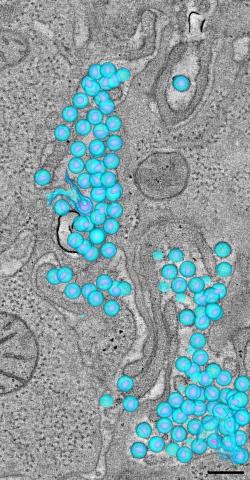
3571: HIV-1 virus in the colon
A tomographic reconstruction of the colon shows the location of large pools of HIV-1 virus particles (in blue) located in the spaces between adjacent cells. The purple objects within each sphere represent the conical cores that are one of the structural hallmarks of the HIV virus.
Mark Ladinsky, California Institute of Technology
View Media
3404: Normal vascular development in frog embryos
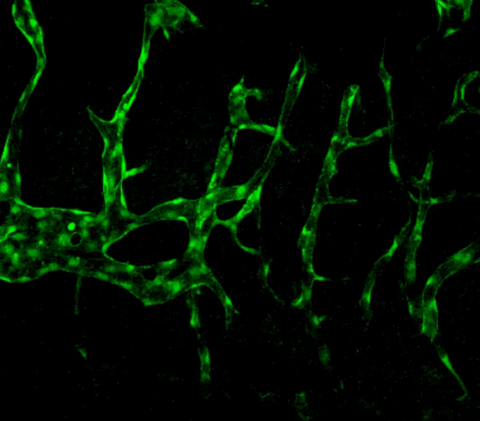
3404: Normal vascular development in frog embryos
Hye Ji Cha, University of Texas at Austin
View Media
2636: Computer model of cell membrane
2636: Computer model of cell membrane
A computer model of the cell membrane, where the plasma membrane is red, endoplasmic reticulum is yellow, and mitochondria are blue. This image relates to a July 27, 2009 article in Computing Life.
Bridget Wilson, University of New Mexico
View Media
2528: A drug's life in the body (with labels)
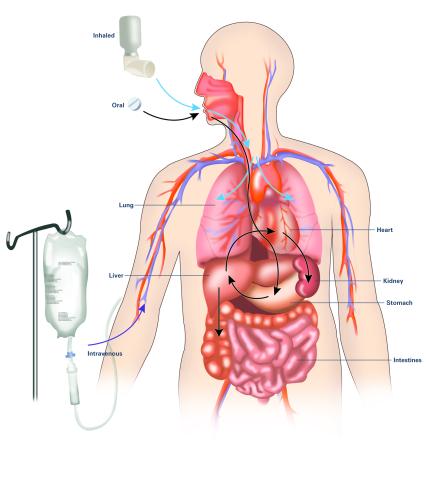
2528: A drug's life in the body (with labels)
A drug's life in the body. Medicines taken by mouth (oral) pass through the liver before they are absorbed into the bloodstream. Other forms of drug administration bypass the liver, entering the blood directly. See 2527 for an unlabeled version of this illustration. Featured in Medicines By Design.
Crabtree + Company
View Media

1090: Natcher Building 10
1090: Natcher Building 10
NIGMS staff are located in the Natcher Building on the NIH campus.
Alisa Machalek, National Institute of General Medical Sciences
View Media
2428: Colorful cells
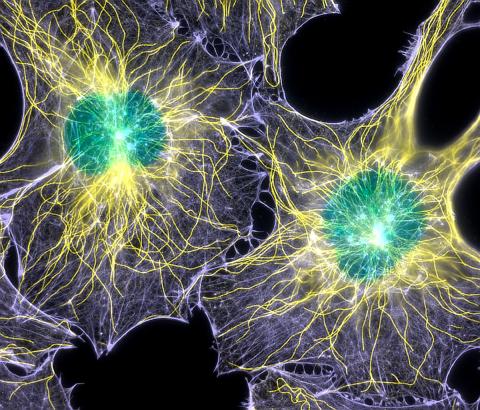
2428: Colorful cells
Actin (purple), microtubules (yellow), and nuclei (green) are labeled in these cells by immunofluorescence. This image won first place in the Nikon 2003 Small World photo competition.
Torsten Wittmann, Scripps Research Institute
View Media
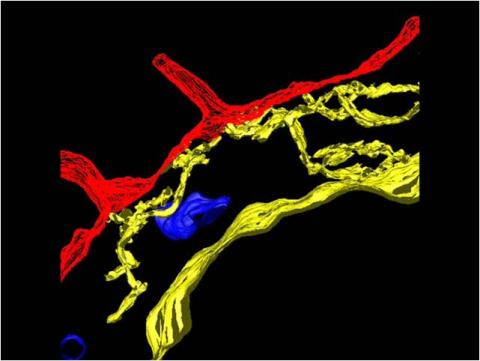
3615: An insect tracheal cell delivers air to muscles
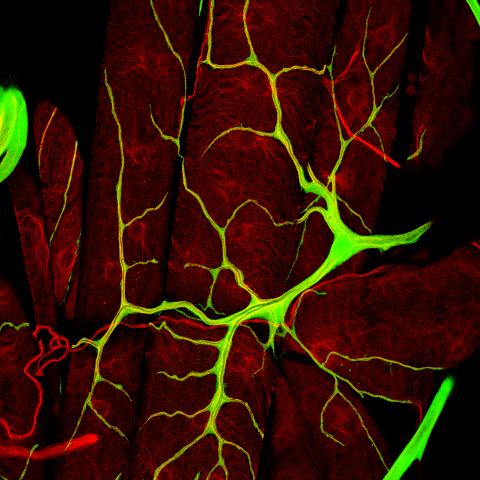
3615: An insect tracheal cell delivers air to muscles
Insects like the fruit fly use an elaborate network of branching tubes called trachea (green) to transport oxygen throughout their bodies. Fruit flies have been used in biomedical research for more than 100 years and remain one of the most frequently studied model organisms. They have a large percentage of genes in common with us, including hundreds of genes that are associated with human diseases.
This image was part of the Life: Magnified exhibit that ran from June 3, 2014, to January 21, 2015, at Dulles International Airport.
This image was part of the Life: Magnified exhibit that ran from June 3, 2014, to January 21, 2015, at Dulles International Airport.
Jayan Nair and Maria Leptin, European Molecular Biology Laboratory, Heidelberg, Germany
View Media
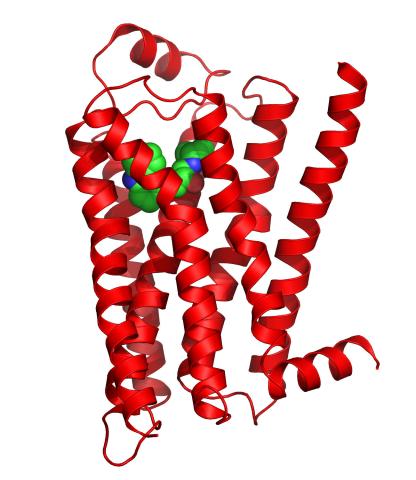
3358: Beta 2-adrenergic receptor
3358: Beta 2-adrenergic receptor
The receptor is shown bound to a partial inverse agonist, carazolol.
Raymond Stevens, The Scripps Research Institute
View Media
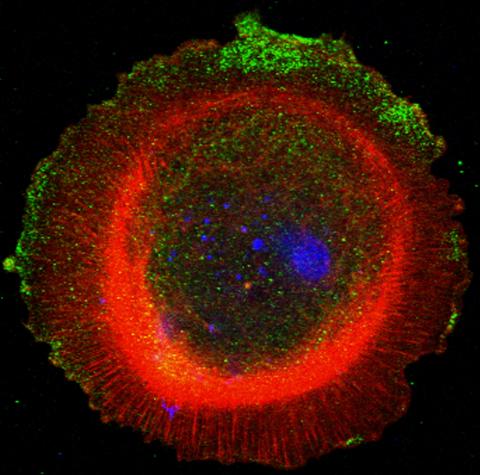
3330: mDia1 antibody staining-01
3330: mDia1 antibody staining-01
Cells move forward with lamellipodia and filopodia supported by networks and bundles of actin filaments. Proper, controlled cell movement is a complex process. Recent research has shown that an actin-polymerizing factor called the Arp2/3 complex is the key component of the actin polymerization engine that drives amoeboid cell motility. ARPC3, a component of the Arp2/3 complex, plays a critical role in actin nucleation. In this photo, the ARPC3+/+ fibroblast cells were fixed and stained with Alexa 546 phalloidin for F-actin (red), mDia1 (green), and DAPI to visualize the nucleus (blue). mDia1 is localized at the lamellipodia of ARPC3+/+ fibroblast cells. Related to images 3328, 3329, 3331, 3332, and 3333.
Rong Li and Praveen Suraneni, Stowers Institute for Medical Research
View Media
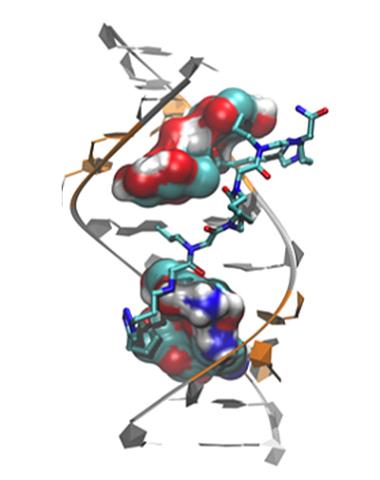
3573: Myotonic dystrophy type 2 genetic defect
3573: Myotonic dystrophy type 2 genetic defect
Scientists revealed a detailed image of the genetic defect that causes myotonic dystrophy type 2 and used that information to design drug candidates to counteract the disease.
Matthew Disney, Scripps Research Institute and Ilyas Yildirim, Northwestern University
View Media
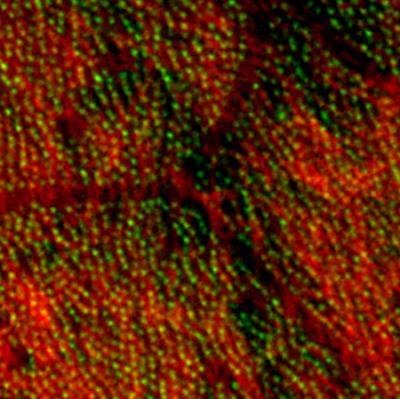
3292: Centrioles anchor cilia in planaria
3292: Centrioles anchor cilia in planaria
Centrioles (green) anchor cilia (red), which project on the surface of pharynx cells of the freshwater planarian Schmidtea mediterranea. Centrioles require cellular structures called centrosomes for assembly in other animal species, but this flatworm known for its regenerative ability was unexpectedly found to lack centrosomes. From a Stowers University news release.
Juliette Azimzadeh, University of California, San Francisco
View Media
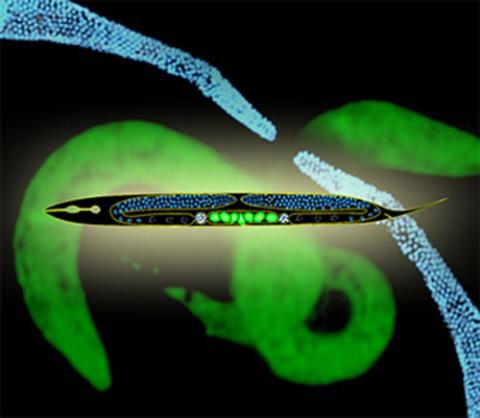
2333: Worms and human infertility
2333: Worms and human infertility
This montage of tiny, transparent C. elegans--or roundworms--may offer insight into understanding human infertility. Researchers used fluorescent dyes to label the worm cells and watch the process of sex cell division, called meiosis, unfold as nuclei (blue) move through the tube-like gonads. Such visualization helps the scientists identify mechanisms that enable these roundworms to reproduce successfully. Because meiosis is similar in all sexually reproducing organisms, what the scientists learn could apply to humans.
Abby Dernburg, Lawrence Berkeley National Laboratory
View Media
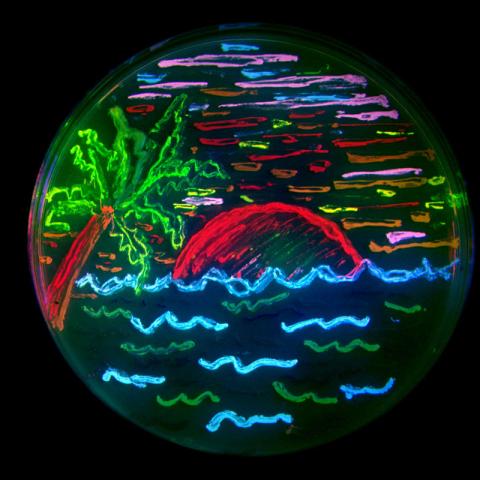

3492: Glowing bacteria make a pretty postcard
3492: Glowing bacteria make a pretty postcard
This tropical scene, reminiscent of a postcard from Key West, is actually a petri dish containing an artistic arrangement of genetically engineered bacteria. The image showcases eight of the fluorescent proteins created in the laboratory of the late Roger Y. Tsien, a cell biologist at the University of California, San Diego. Tsien, along with Osamu Shimomura of the Marine Biology Laboratory and Martin Chalfie of Columbia University, share the 2008 Nobel Prize in chemistry for their work on green fluorescent protein-a naturally glowing molecule from jellyfish that has become a powerful tool for studying molecules inside living cells.
Nathan C. Shaner, The Scintillon Institute
View Media
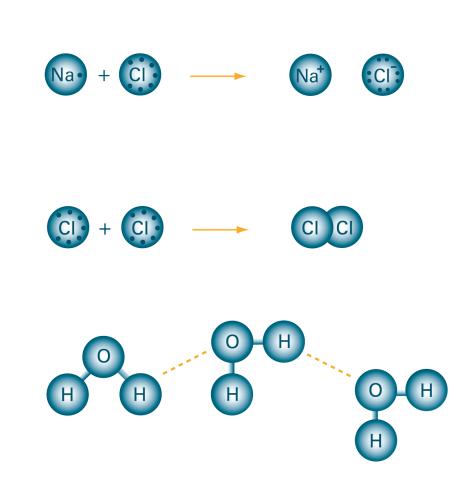
2519: Bond types
2519: Bond types
Ionic and covalent bonds hold molecules, like sodium chloride and chlorine gas, together. Hydrogen bonds among molecules, notably involving water, also play an important role in biology. See image 2520 for a labeled version of this illustration. Featured in The Chemistry of Health.
Crabtree + Company
View Media
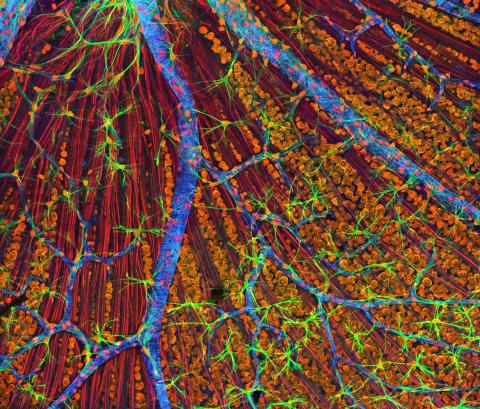
3309: Mouse Retina
3309: Mouse Retina
A genetic disorder of the nervous system, neurofibromatosis causes tumors to form on nerves throughout the body, including a type of tumor called an optic nerve glioma that can result in childhood blindness. The image was used to demonstrate the unique imaging capabilities of one of our newest (at the time) laser scanning microscopes and is of a wildtype (normal) mouse retina in the optic fiber layer. This layer is responsible for relaying information from the retina to the brain and was fluorescently stained to reveal the distribution of glial cells (green), DNA and RNA in the cell bodies of the retinal ganglion neurons (orange) and their optic nerve fibers (red), and actin in endothelial cells surrounding a prominent branching blood vessel (blue). By studying the microscopic structure of normal and diseased retina and optic nerves, we hope to better understand the altered biology of the tissues in these tumors with the prospects of developing therapeutic interventions.
Tom Deerinck, NCMIR
View Media
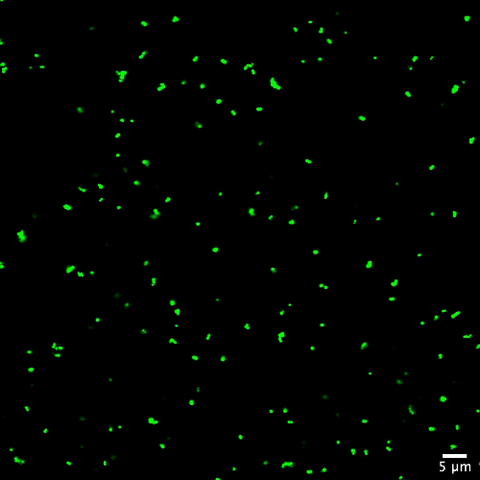
6805: Staphylococcus aureus aggregating upon contact with synovial fluid
6805: Staphylococcus aureus aggregating upon contact with synovial fluid
Staphylococcus aureus bacteria (green) grouping together upon contact with synovial fluid—a viscous substance found in joints. The formation of groups can help protect the bacteria from immune system defenses and from antibiotics, increasing the likelihood of an infection. This video is a 1-hour time lapse and was captured using a confocal laser scanning microscope.
More information about the research that produced this video can be found in the Journal of Bacteriology paper "In Vitro Staphylococcal Aggregate Morphology and Protection from Antibiotics Are Dependent on Distinct Mechanisms Arising from Postsurgical Joint Components and Fluid Motion" by Staats et al.
Related to images 6803 and 6804.
More information about the research that produced this video can be found in the Journal of Bacteriology paper "In Vitro Staphylococcal Aggregate Morphology and Protection from Antibiotics Are Dependent on Distinct Mechanisms Arising from Postsurgical Joint Components and Fluid Motion" by Staats et al.
Related to images 6803 and 6804.
Paul Stoodley, The Ohio State University.
View Media
1251: Crab larva eye
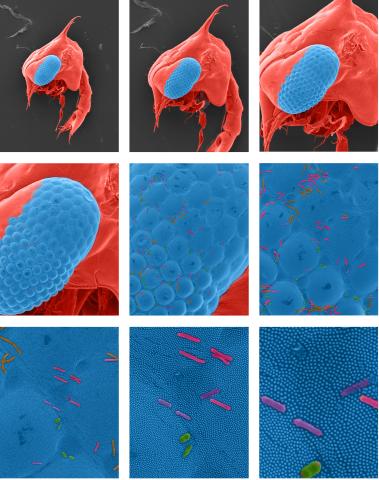
1251: Crab larva eye
Colorized scanning electron micrographs progressively zoom in on the eye of a crab larva. In the higher-resolution frames, bacteria are visible on the eye.
Tina Weatherby Carvalho, University of Hawaii at Manoa
View Media
3567: RSV-Infected Cell
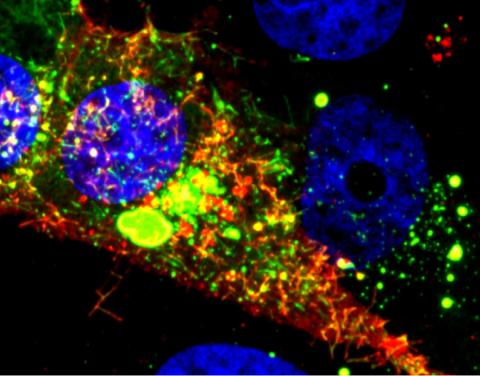
3567: RSV-Infected Cell
Viral RNA (red) in an RSV-infected cell. More information about the research behind this image can be found in a Biomedical Beat Blog posting from January 2014.
Eric Alonas and Philip Santangelo, Georgia Institute of Technology and Emory University
View Media
2693: Fruit fly in the pink
2693: Fruit fly in the pink
Fruit flies are a common model organism for basic medical research.
Crabtree + Company
View Media
1316: Mitosis - interphase
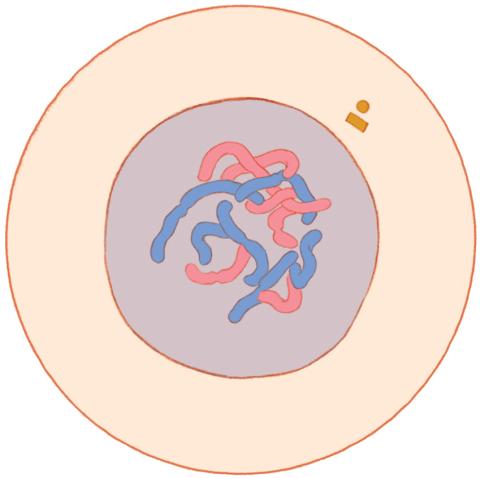
1316: Mitosis - interphase
A cell in interphase, at the start of mitosis: Chromosomes duplicate, and the copies remain attached to each other. Mitosis is responsible for growth and development, as well as for replacing injured or worn out cells throughout the body. For simplicity, mitosis is illustrated here with only six chromosomes.
Judith Stoffer
View Media
2382: PanB from M. tuberculosis (2)
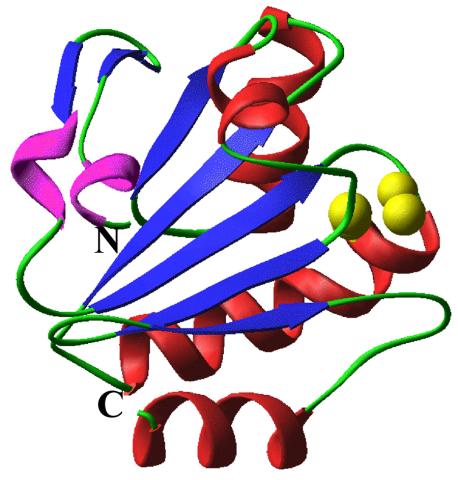
2382: PanB from M. tuberculosis (2)
Model of an enzyme, PanB, from Mycobacterium tuberculosis, the bacterium that causes most cases of tuberculosis. This enzyme is an attractive drug target.
Mycobacterium Tuberculosis Center, PSI-1
View Media
2540: Chromosome inside nucleus (with labels)
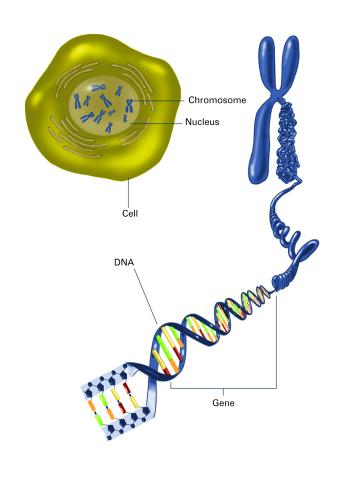
2540: Chromosome inside nucleus (with labels)
The long, stringy DNA that makes up genes is spooled within chromosomes inside the nucleus of a cell. (Note that a gene would actually be a much longer stretch of DNA than what is shown here.) See image 2539 for an unlabeled version of this illustration. Featured in The New Genetics.
Crabtree + Company
View Media

3718: A Bacillus subtilis biofilm grown in a Petri dish
3718: A Bacillus subtilis biofilm grown in a Petri dish
Bacterial biofilms are tightly knit communities of bacterial cells growing on, for example, solid surfaces, such as in water pipes or on teeth. Here, cells of the bacterium Bacillus subtilis have formed a biofilm in a laboratory culture. Researchers have discovered that the bacterial cells in a biofilm communicate with each other through electrical signals via specialized potassium ion channels to share resources, such as nutrients, with each other. This insight may help scientists to improve sanitation systems to prevent biofilms, which often resist common treatments, from forming and to develop better medicines to combat bacterial infections. See the Biomedical Beat blog post Bacterial Biofilms: A Charged Environment for more information.
Gürol Süel, UCSD
View Media
2754: Myosin V binding to actin
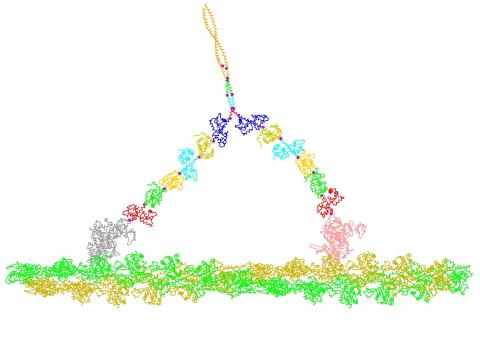
2754: Myosin V binding to actin
This simulation of myosin V binding to actin was created using the software tool Protein Mechanica. With Protein Mechanica, researchers can construct models using information from a variety of sources: crystallography, cryo-EM, secondary structure descriptions, as well as user-defined solid shapes, such as spheres and cylinders. The goal is to enable experimentalists to quickly and easily simulate how different parts of a molecule interact.
Simbios, NIH Center for Biomedical Computation at Stanford
View Media

