Switch to List View
Image and Video Gallery
This is a searchable collection of scientific photos, illustrations, and videos. The images and videos in this gallery are licensed under Creative Commons Attribution Non-Commercial ShareAlike 3.0. This license lets you remix, tweak, and build upon this work non-commercially, as long as you credit and license your new creations under identical terms.
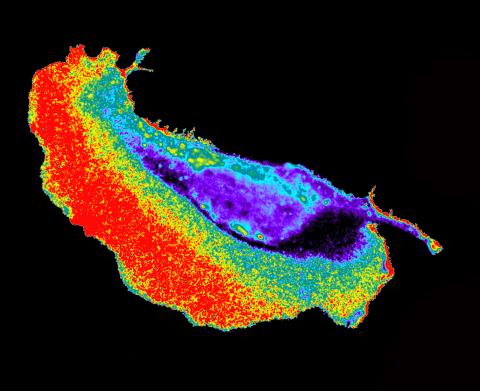
2452: Seeing signaling protein activation in cells 02
2452: Seeing signaling protein activation in cells 02
Cdc42, a member of the Rho family of small guanosine triphosphatase (GTPase) proteins, regulates multiple cell functions, including motility, proliferation, apoptosis, and cell morphology. In order to fulfill these diverse roles, the timing and location of Cdc42 activation must be tightly controlled. Klaus Hahn and his research group use special dyes designed to report protein conformational changes and interactions, here in living neutrophil cells. Warmer colors in this image indicate higher levels of activation. Cdc42 looks to be activated at cell protrusions.
Related to images 2451, 2453, and 2454.
Related to images 2451, 2453, and 2454.
Klaus Hahn, University of North Carolina, Chapel Hill Medical School
View Media
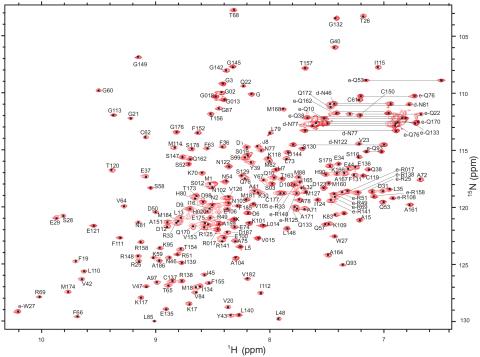
2299: 2-D NMR
2299: 2-D NMR
A two-dimensional NMR spectrum of a protein, in this case a 2D 1H-15N HSQC NMR spectrum of a 228 amino acid DNA/RNA-binding protein.
Dr. Xiaolian Gao's laboratory at the University of Houston
View Media
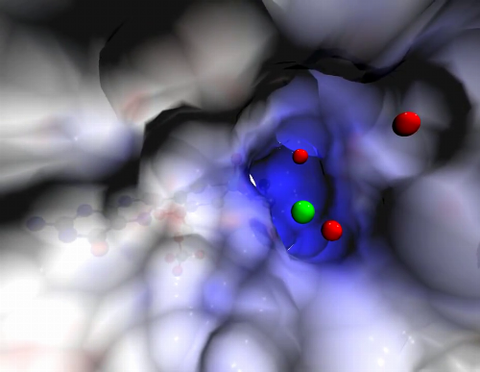
2746: Active site of sulfite oxidase
2746: Active site of sulfite oxidase
Sulfite oxidase is an enzyme that is essential for normal neurological development in children. This video shows the active site of the enzyme and its molybdenum cofactor visible as a faint ball-and-stick representation buried within the protein. The positively charged channel (blue) at the active site contains a chloride ion (green) and three water molecules (red). As the protein oscillates, one can see directly down the positively charged channel. At the bottom is the molybdenum atom of the active site (light blue) and its oxo group (red) that is transferred to sulfite to form sulfate in the catalytic reaction.
John Enemark, University of Arizona
View Media
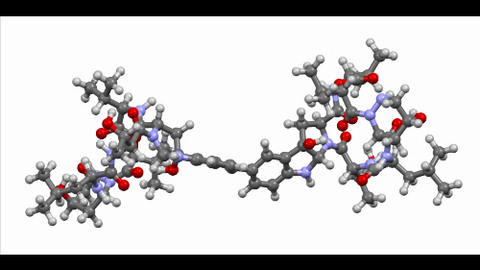
6851: Himastatin, 360-degree view
6851: Himastatin, 360-degree view
A 360-degree view of the molecule himastatin, which was first isolated from the bacterium Streptomyces himastatinicus. Himastatin shows antibiotic activity. The researchers who created this video developed a new, more concise way to synthesize himastatin so it can be studied more easily.
More information about the research that produced this video can be found in the Science paper “Total synthesis of himastatin” by D’Angelo et al.
Related to images 6848 and 6850.
More information about the research that produced this video can be found in the Science paper “Total synthesis of himastatin” by D’Angelo et al.
Related to images 6848 and 6850.
Mohammad Movassaghi, Massachusetts Institute of Technology.
View Media
6994: Respiratory droplet
6994: Respiratory droplet
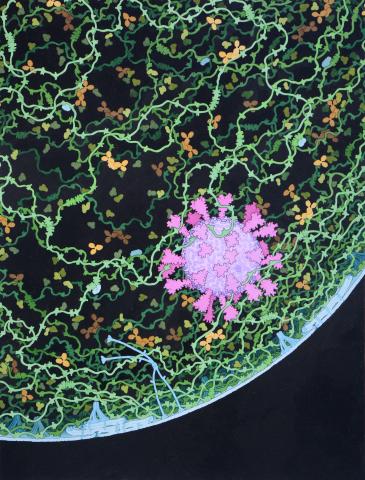
This painting shows a cross section of a small respiratory droplet, like the ones that are thought to transmit SARS-CoV-2, the virus that causes COVID-19. The virus is shown in pink, and the droplet is also filled with molecules that are present in the respiratory tract, including mucins (green), pulmonary surfactant proteins and lipids (blue), and antibodies (tan).
Amy Wu and Christine Zardecki, RCSB Protein Data Bank.
View Media
2318: Gene silencing
2318: Gene silencing
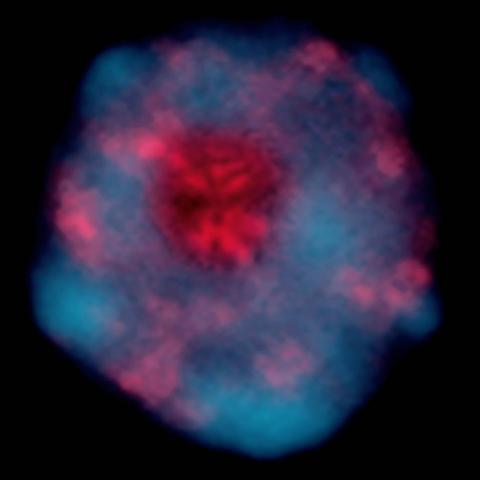
Pretty in pink, the enzyme histone deacetylase (HDA6) stands out against a background of blue-tinted DNA in the nucleus of an Arabidopsis plant cell. Here, HDA6 concentrates in the nucleolus (top center), where ribosomal RNA genes reside. The enzyme silences the ribosomal RNA genes from one parent while those from the other parent remain active. This chromosome-specific silencing of ribosomal RNA genes is an unusual phenomenon observed in hybrid plants.
Olga Pontes and Craig Pikaard, Washington University
View Media
2740: Early life of a protein
2740: Early life of a protein
This illustration represents the early life of a protein—specifically, apomyoglobin—as it is synthesized by a ribosome and emerges from the ribosomal tunnel, which contains the newly formed protein's conformation. The synthesis occurs in the complex swirl of the cell medium, filled with interactions among many molecules. Researchers in Silvia Cavagnero's laboratory are studying the structure and dynamics of newly made proteins and polypeptides using spectroscopic and biochemical techniques.
Silvia Cavagnero, University of Wisconsin, Madison
View Media
2343: Protein rv2844 from M. tuberculosis
2343: Protein rv2844 from M. tuberculosis
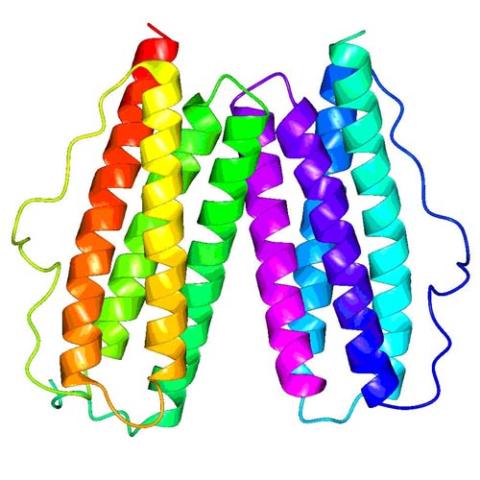
This crystal structure shows a conserved hypothetical protein from Mycobacterium tuberculosis. Only 12 other proteins share its sequence homology, and none has a known function. This structure indicates the protein may play a role in metabolic pathways. Featured as one of the August 2007 Protein Structure Initiative Structures of the Month.
Integrated Center for Structure and Function Innovation
View Media
3253: Pulsating response to stress in bacteria
3253: Pulsating response to stress in bacteria
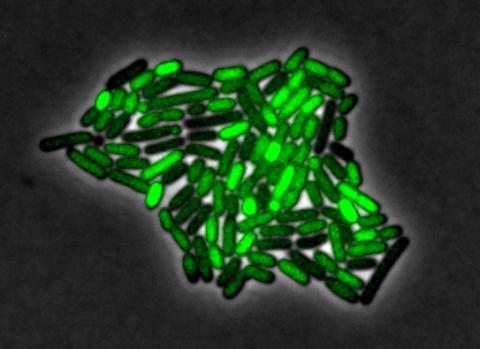
By attaching fluorescent proteins to the genetic circuit responsible for B. subtilis's stress response, researchers can observe the cells' pulses as green flashes. In response to a stressful environment like one lacking food, B. subtilis activates a large set of genes that help it respond to the hardship. Instead of leaving those genes on as previously thought, researchers discovered that the bacteria flip the genes on and off, increasing the frequency of these pulses with increasing stress. See entry 3254 for the related video.
Michael Elowitz, Caltech University
View Media
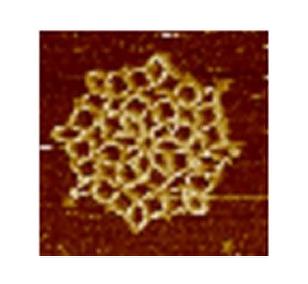

3724: Snowflake DNA origami
3724: Snowflake DNA origami
An atomic force microscopy image shows DNA folded into an intricate, computer-designed structure. The image is featured on Biomedical Beat blog post Cool Images: A Holiday-Themed Collection. For more background on DNA origami, see Cool Image: DNA Origami. See also related image 3690.
Hao Yan, Arizona State University
View Media
3415: X-ray co-crystal structure of Src kinase bound to a DNA-templated macrocycle inhibitor 3
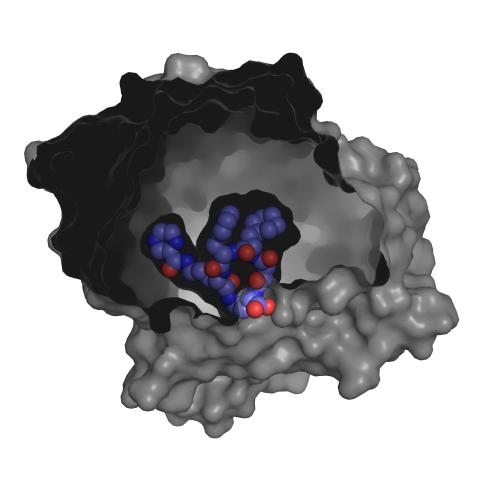
3415: X-ray co-crystal structure of Src kinase bound to a DNA-templated macrocycle inhibitor 3
X-ray co-crystal structure of Src kinase bound to a DNA-templated macrocycle inhibitor. Related to 3413, 3414, 3416, 3417, 3418, and 3419.
Markus A. Seeliger, Stony Brook University Medical School and David R. Liu, Harvard University
View Media
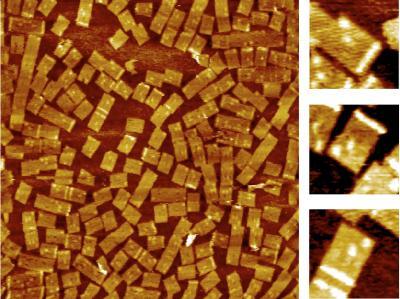

2455: Golden gene chips
2455: Golden gene chips
A team of chemists and physicists used nanotechnology and DNA's ability to self-assemble with matching RNA to create a new kind of chip for measuring gene activity. When RNA of a gene of interest binds to a DNA tile (gold squares), it creates a raised surface (white areas) that can be detected by a powerful microscope. This nanochip approach offers manufacturing and usage advantages over existing gene chips and is a key step toward detecting gene activity in a single cell. Featured in the February 20, 2008, issue of Biomedical Beat.
Hao Yan and Yonggang Ke, Arizona State University
View Media
3446: Biofilm blocking fluid flow
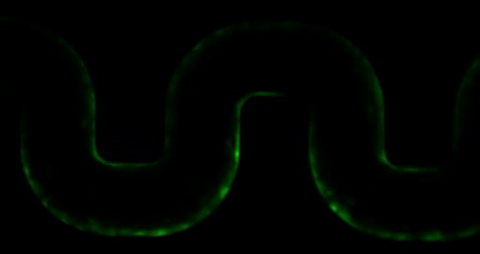
3446: Biofilm blocking fluid flow
This time-lapse movie shows that bacterial communities called biofilms can create blockages that prevent fluid flow in devices such as stents and catheters over a period of about 56 hours. This video was featured in a news release from Princeton University.
Bonnie Bassler, Princeton University
View Media
2588: Genetic patchworks
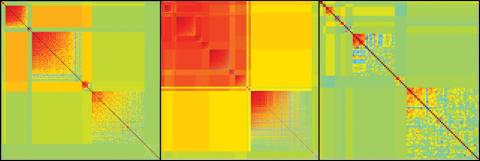
2588: Genetic patchworks
Each point in these colorful patchworks represents the correlation between two sleep-associated genes in fruit flies. Vibrant reds and oranges represent high and intermediate degrees of association between the genes, respectively. Genes in these areas show similar activity patterns in different fly lines. Cool blues represent gene pairs where one partner's activity is high and the other's is low. The green areas show pairs with activities that are not correlated. These quilt-like depictions help illustrate a recent finding that genes act in teams to influence sleep patterns.
Susan Harbison and Trudy Mackay, North Carolina State University
View Media
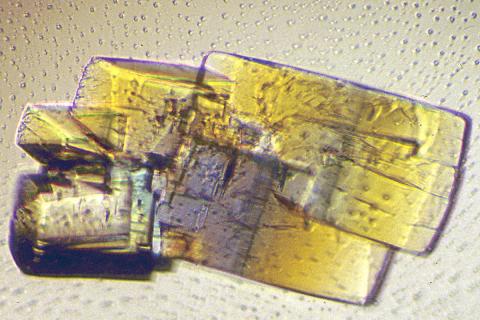

2402: RNase A (2)
2402: RNase A (2)
A crystal of RNase A protein created for X-ray crystallography, which can reveal detailed, three-dimensional protein structures.
Alex McPherson, University of California, Irvine
View Media

3530: Lorsch Swearing In
3530: Lorsch Swearing In
Jon Lorsch at his swearing in as NIGMS director in August 2013. Also shown are Francis Collins, NIH Director, and Judith Greenberg, former NIGMS Acting Director.
View Media
1316: Mitosis - interphase
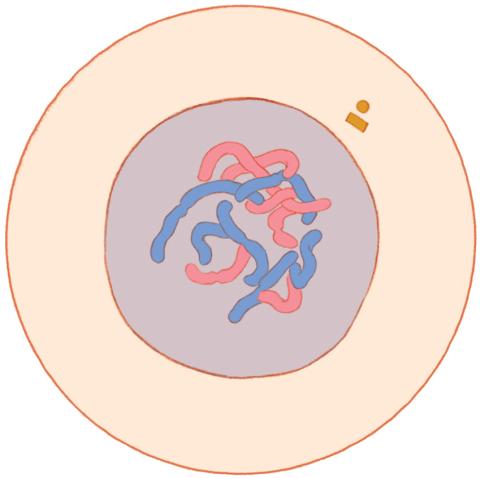
1316: Mitosis - interphase
A cell in interphase, at the start of mitosis: Chromosomes duplicate, and the copies remain attached to each other. Mitosis is responsible for growth and development, as well as for replacing injured or worn out cells throughout the body. For simplicity, mitosis is illustrated here with only six chromosomes.
Judith Stoffer
View Media
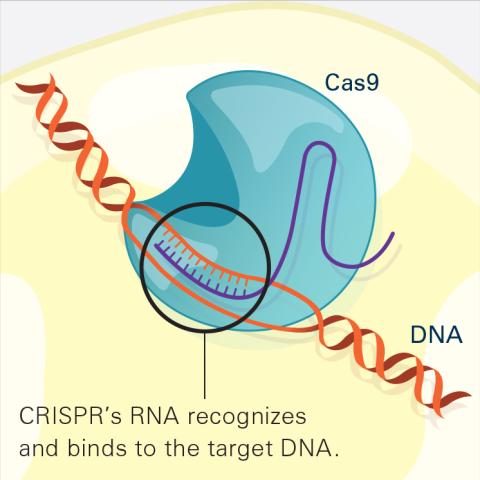
6486: CRISPR Illustration Frame 2
6486: CRISPR Illustration Frame 2
This illustration shows, in simplified terms, how the CRISPR-Cas9 system can be used as a gene-editing tool. The CRISPR system has two components joined together: a finely tuned targeting device (a small strand of RNA programmed to look for a specific DNA sequence) and a strong cutting device (an enzyme called Cas9 that can cut through a double strand of DNA). In this frame (2 of 4), the CRISPR machine locates the target DNA sequence once inserted into a cell.
For an explanation and overview of the CRISPR-Cas9 system, see the iBiology video, and find the full CRIPSR illustration here.
For an explanation and overview of the CRISPR-Cas9 system, see the iBiology video, and find the full CRIPSR illustration here.
National Institute of General Medical Sciences.
View Media
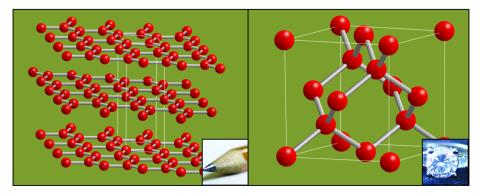
2507: Carbon building blocks (with examples)
2507: Carbon building blocks (with examples)
The arrangement of identical molecular components can make a dramatic difference. For example, carbon atoms can be arranged into dull graphite (left) or sparkly diamonds (right). See image 2506 for an illustration without examples.
Crabtree + Company
View Media
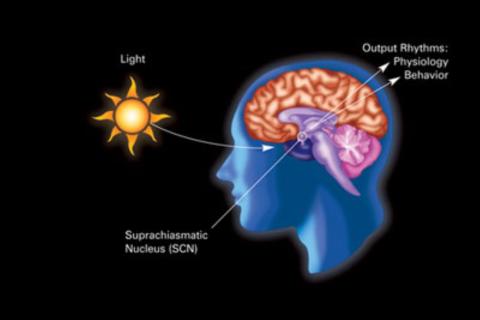
2841: Circadian rhythm
2841: Circadian rhythm
The human body keeps time with a master clock called the suprachiasmatic nucleus or SCN. Situated inside the brain, it's a tiny sliver of tissue about the size of a grain of rice, located behind the eyes. It sits quite close to the optic nerve, which controls vision, and this means that the SCN "clock" can keep track of day and night. The SCN helps control sleep by coordinating the actions of billions of miniature "clocks" throughout the body. These aren't actually clocks, but rather are ensembles of genes inside clusters of cells that switch on and off in a regular, 24-hour cycle in our physiological day.
Crabtree + Company
View Media
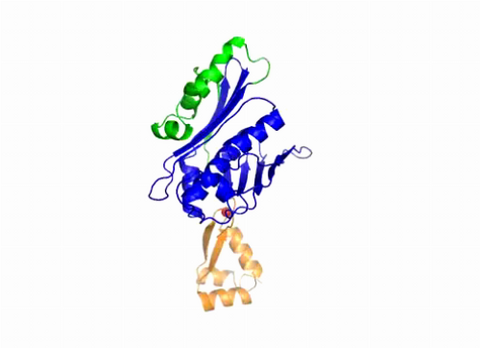
3402: Hsp33 Heat Shock Protein Inactive to Active
3402: Hsp33 Heat Shock Protein Inactive to Active
When the heat shock protein hsp33 is folded, it is inactive and contains a zinc ion, stabilizing the redox sensitive domain (orange). In the presence of an environmental stressor, the protein releases the zinc ion, which leads to the unfolding of the redox domain. This unfolding causes the chaperone to activate by reaching out its "arm" (green) to protect other proteins.
Dana Reichmann, University of Michigan
View Media
6791: Yeast cells entering mitosis
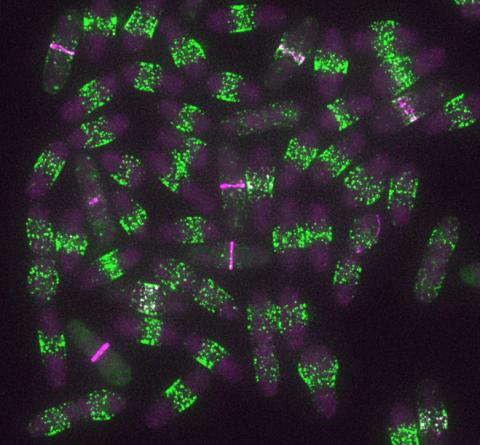
6791: Yeast cells entering mitosis
Yeast cells entering mitosis, also known as cell division. The green and magenta dots are two proteins that play important roles in mitosis. They show where the cells will split. This image was captured using wide-field microscopy with deconvolution.
Related to images 6792, 6793, 6794, 6797, 6798, and videos 6795 and 6796.
Related to images 6792, 6793, 6794, 6797, 6798, and videos 6795 and 6796.
Alaina Willet, Kathy Gould’s lab, Vanderbilt University.
View Media
3614: Birth of a yeast cell
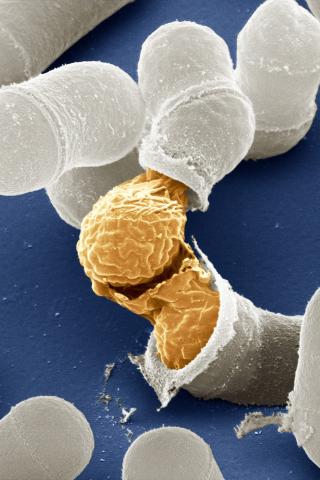
3614: Birth of a yeast cell
Yeast make bread, beer, and wine. And like us, yeast can reproduce sexually. A mother and father cell fuse and create one large cell that contains four offspring. When environmental conditions are favorable, the offspring are released, as shown here. Yeast are also a popular study subject for scientists. Research on yeast has yielded vast knowledge about basic cellular and molecular biology as well as about myriad human diseases, including colon cancer and various metabolic disorders.
This image was part of the Life: Magnified exhibit that ran from June 3, 2014, to January 21, 2015, at Dulles International Airport.
This image was part of the Life: Magnified exhibit that ran from June 3, 2014, to January 21, 2015, at Dulles International Airport.
Juergen Berger, Max Planck Institute for Developmental Biology, and Maria Langegger, Friedrich Miescher Laboratory of the Max Planck Society, Germany
View Media
3509: Neuron with labeled synapses
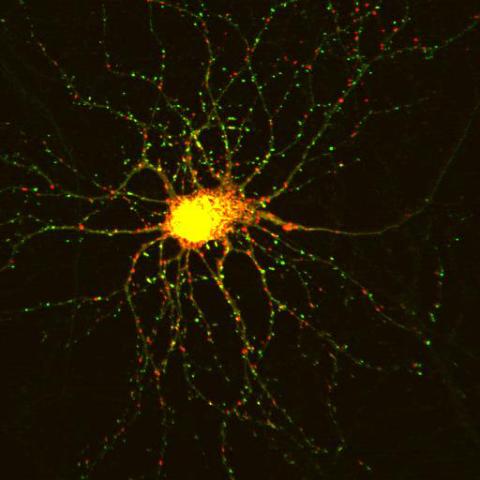
3509: Neuron with labeled synapses
In this image, recombinant probes known as FingRs (Fibronectin Intrabodies Generated by mRNA display) were expressed in a cortical neuron, where they attached fluorescent proteins to either PSD95 (green) or Gephyrin (red). PSD-95 is a marker for synaptic strength at excitatory postsynaptic sites, and Gephyrin plays a similar role at inhibitory postsynaptic sites. Thus, using FingRs it is possible to obtain a map of synaptic connections onto a particular neuron in a living cell in real time.
Don Arnold and Richard Roberts, University of Southern California.
View Media
1330: Mitosis - prophase
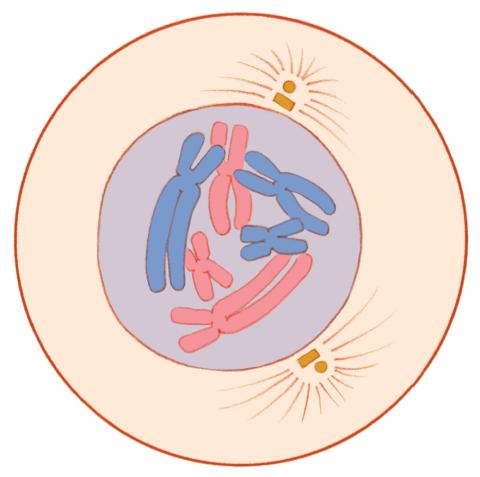
1330: Mitosis - prophase
A cell in prophase, near the start of mitosis: In the nucleus, chromosomes condense and become visible. In the cytoplasm, the spindle forms. Mitosis is responsible for growth and development, as well as for replacing injured or worn out cells throughout the body. For simplicity, mitosis is illustrated here with only six chromosomes.
Judith Stoffer
View Media

6550: Time-lapse video of floral pattern in a mixture of two bacterial species, Acinetobacter baylyi and Escherichia coli, grown on a semi-solid agar for 24 hours
6550: Time-lapse video of floral pattern in a mixture of two bacterial species, Acinetobacter baylyi and Escherichia coli, grown on a semi-solid agar for 24 hours
This time-lapse video shows the emergence of a flower-like pattern in a mixture of two bacterial species, motile Acinetobacter baylyi and non-motile Escherichia coli (green), that are grown together for 24 hours on 0.75% agar surface from a small inoculum in the center of a Petri dish.
See 6557 for a photo of this process at 24 hours on 0.75% agar surface.
See 6553 for a photo of this process at 48 hours on 1% agar surface.
See 6555 for another photo of this process at 48 hours on 1% agar surface.
See 6556 for a photo of this process at 72 hours on 0.5% agar surface.
See 6557 for a photo of this process at 24 hours on 0.75% agar surface.
See 6553 for a photo of this process at 48 hours on 1% agar surface.
See 6555 for another photo of this process at 48 hours on 1% agar surface.
See 6556 for a photo of this process at 72 hours on 0.5% agar surface.
L. Xiong et al, eLife 2020;9: e48885
View Media

2361: Chromium X-ray source
2361: Chromium X-ray source
In the determination of protein structures by X-ray crystallography, this unique soft (l = 2.29Å) X-ray source is used to collect anomalous scattering data from protein crystals containing light atoms such as sulfur, calcium, zinc and phosphorous. These data can be used to image the protein.
The Southeast Collaboratory for Structural Genomics
View Media
3628: Skin cancer cells (squamous cell carcinoma)
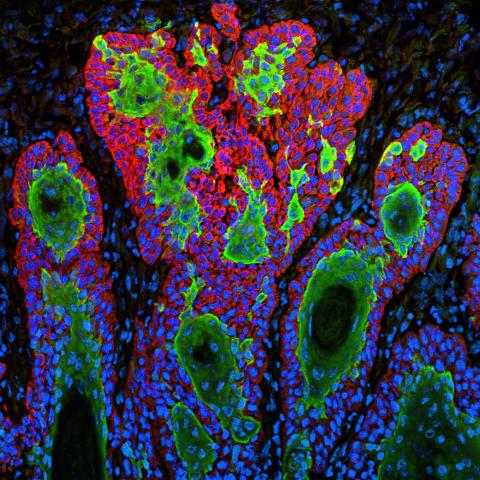
3628: Skin cancer cells (squamous cell carcinoma)
This image shows the uncontrolled growth of cells in squamous cell carcinoma, the second most common form of skin cancer. If caught early, squamous cell carcinoma is usually not life-threatening.
This image was part of the Life: Magnified exhibit that ran from June 3, 2014, to January 21, 2015, at Dulles International Airport.
This image was part of the Life: Magnified exhibit that ran from June 3, 2014, to January 21, 2015, at Dulles International Airport.
Markus Schober and Elaine Fuchs, The Rockefeller University
View Media
3397: Myelinated axons 2
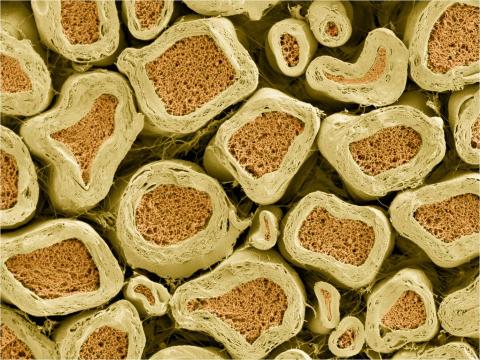
3397: Myelinated axons 2
Top view of myelinated axons in a rat spinal root. Myelin is a type of fat that forms a sheath around and thus insulates the axon to protect it from losing the electrical current needed to transmit signals along the axon. The axoplasm inside the axon is shown in pink. Related to 3396.
Tom Deerinck, National Center for Microscopy and Imaging Research (NCMIR)
View Media
6350: Aldolase
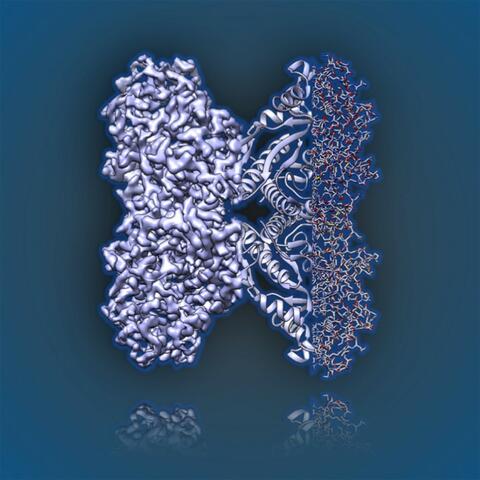
6350: Aldolase
2.5Å resolution reconstruction of rabbit muscle aldolase collected on a FEI/Thermo Fisher Titan Krios with energy filter and image corrector.
National Resource for Automated Molecular Microscopy http://nramm.nysbc.org/nramm-images/ Source: Bridget Carragher
View Media
5881: Zebrafish larva
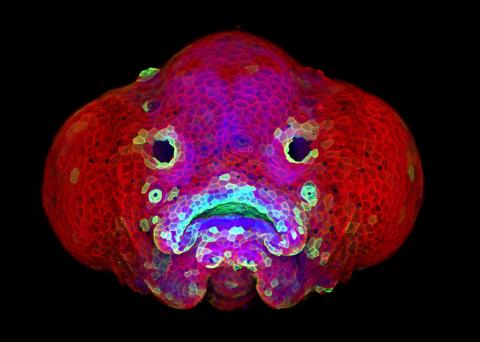
5881: Zebrafish larva
You are face to face with a 6-day-old zebrafish larva. What look like eyes will become nostrils, and the bulges on either side will become eyes. Scientists use fast-growing, transparent zebrafish to see body shapes form and organs develop over the course of just a few days. Images like this one help researchers understand how gene mutations can lead to facial abnormalities such as cleft lip and palate in people.
This image won a 2016 FASEB BioArt award. In addition, NIH Director Francis Collins featured this on his blog on January 26, 2017.
This image won a 2016 FASEB BioArt award. In addition, NIH Director Francis Collins featured this on his blog on January 26, 2017.
Oscar Ruiz and George Eisenhoffer, University of Texas MD Anderson Cancer Center, Houston
View Media
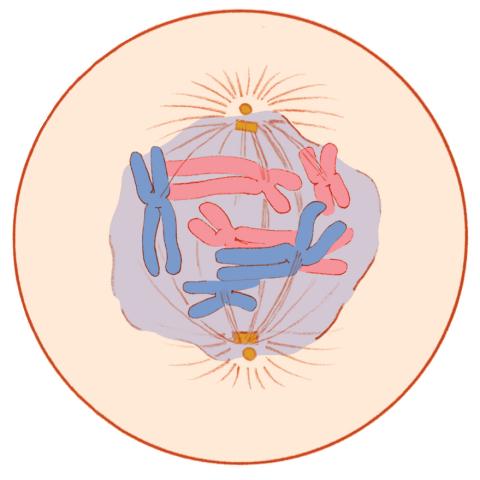
1331: Mitosis - prometaphase
1331: Mitosis - prometaphase
A cell in prometaphase during mitosis: The nuclear membrane breaks apart, and the spindle starts to interact with the chromosomes. Mitosis is responsible for growth and development, as well as for replacing injured or worn out cells throughout the body. For simplicity, mitosis is illustrated here with only six chromosomes.
Judith Stoffer
View Media
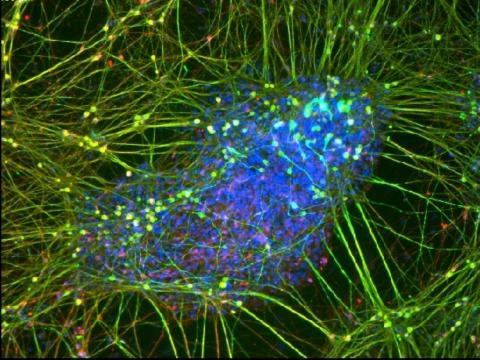
3264: Peripheral nerve cell derived from ES cells
3264: Peripheral nerve cell derived from ES cells
A peripheral nerve cell made from human embryonic stem cell-derived neural crest stem cells. The nucleus is shown in blue, and nerve cell proteins peripherin and beta-tubulin (Tuj1) are shown in green and red, respectively. Related to image 3263.
Stephen Dalton, University of Georgia
View Media
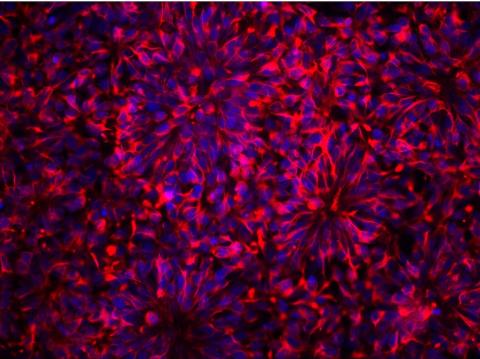
3284: Neurons from human ES cells
3284: Neurons from human ES cells
These neural precursor cells were derived from human embryonic stem cells. The neural cell bodies are stained red, and the nuclei are blue. Image and caption information courtesy of the California Institute for Regenerative Medicine.
Xianmin Zeng lab, Buck Institute for Age Research, via CIRM
View Media
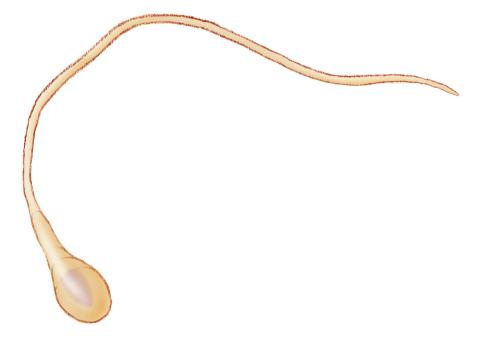
1293: Sperm cell
6503: Arabidopsis Thaliana: Flowers Spring to Life
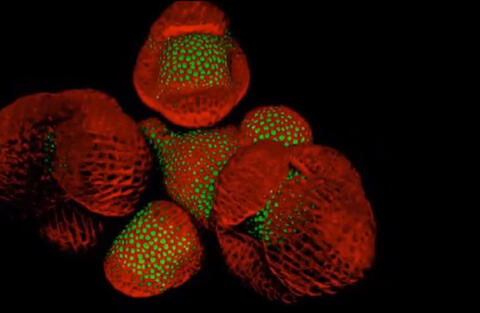
6503: Arabidopsis Thaliana: Flowers Spring to Life
This image capture shows how a single gene, STM, plays a starring role in plant development. This gene acts like a molecular fountain of youth, keeping cells ever-young until it’s time to grow up and commit to making flowers and other plant parts. Because of its ease of use and low cost, Arabidopsis is a favorite model for scientists to learn the basic principles driving tissue growth and regrowth for humans as well as the beautiful plants outside your window. Image captured from video Watch Flowers Spring to Life, featured in the NIH Director's Blog: Watch Flowers Spring to Life.
Nathanaёl Prunet NIH Support: National Institute of General Medical Sciences
View Media
3429: Enzyme transition states
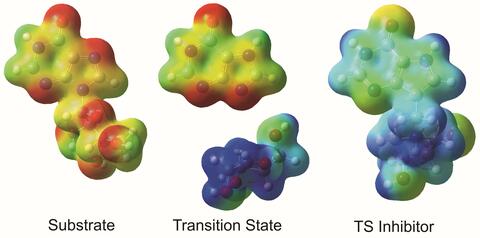
3429: Enzyme transition states
The molecule on the left is an electrostatic potential map of the van der Waals surface of the transition state for human purine nucleoside phosphorylase. The colors indicate the electron density at any position of the molecule. Red indicates electron-rich regions with negative charge and blue indicates electron-poor regions with positive charge. The molecule on the right is called DADMe-ImmH. It is a chemically stable analogue of the transition state on the left. It binds to the enzyme millions of times tighter than the substrate. This inhibitor is in human clinical trials for treating patients with gout. This image appears in Figure 4, Schramm, V.L. (2011) Annu. Rev. Biochem. 80:703-732.
Vern Schramm, Albert Einstein College of Medicine of Yeshiva University
View Media
6570: Stress Response in Cells
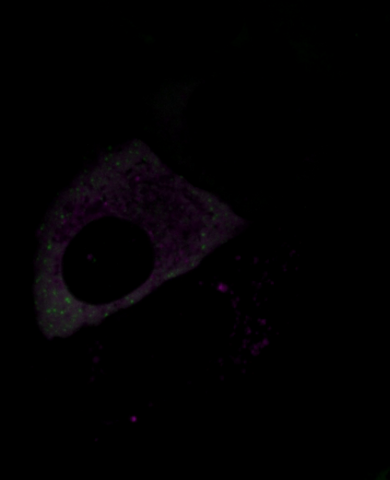
6570: Stress Response in Cells
Two highly stressed osteosarcoma cells are shown with a set of green droplet-like structures followed by a second set of magenta droplets. These droplets are composed of fluorescently labeled stress-response proteins, either G3BP or UBQLN2 (Ubiquilin-2). Each protein is undergoing a fascinating process, called phase separation, in which a non-membrane bound compartment of the cytoplasm emerges with a distinct environment from the surrounding cytoplasm. Subsequently, the proteins fuse with like proteins to form larger droplets, in much the same way that raindrops merge on a car’s windshield.
Julia F. Riley and Carlos A. Castañeda, Syracuse University
View Media
6569: Cryo-electron tomography of a Caulobacter bacterium
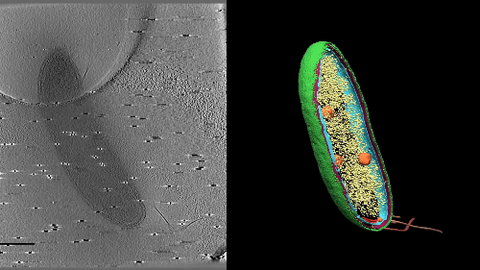
6569: Cryo-electron tomography of a Caulobacter bacterium
3D image of Caulobacter bacterium with various components highlighted: cell membranes (red and blue), protein shell (green), protein factories known as ribosomes (yellow), and storage granules (orange).
Peter Dahlberg, Stanford University.
View Media
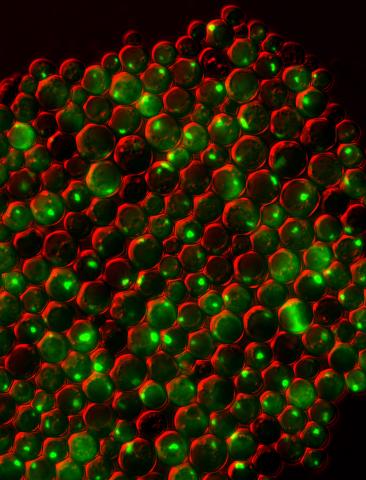
3550: Protein clumping in zinc-deficient yeast cells
3550: Protein clumping in zinc-deficient yeast cells
The green spots in this image are clumps of protein inside yeast cells that are deficient in both zinc and a protein called Tsa1 that prevents clumping. Protein clumping plays a role in many diseases, including Parkinson's and Alzheimer's, where proteins clump together in the brain. Zinc deficiency within a cell can cause proteins to mis-fold and eventually clump together. Normally, in yeast, Tsa1 codes for so-called "chaperone proteins" which help proteins in stressed cells, such as those with a zinc deficiency, fold correctly. The research behind this image was published in 2013 in the Journal of Biological Chemistry.
Colin MacDiarmid and David Eide, University of Wisconsin--Madison
View Media
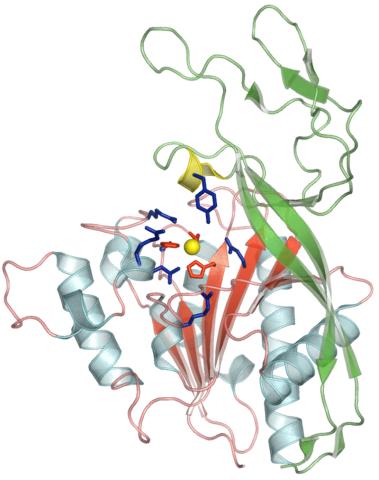
2352: Human aspartoacylase
2352: Human aspartoacylase
Model of aspartoacylase, a human enzyme involved in brain metabolism.
Center for Eukaryotic Structural Genomics, PSI
View Media
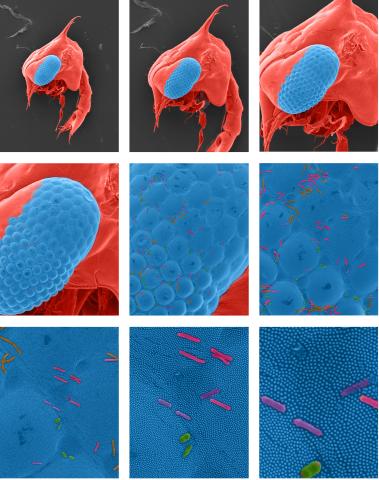
1251: Crab larva eye
1251: Crab larva eye
Colorized scanning electron micrographs progressively zoom in on the eye of a crab larva. In the higher-resolution frames, bacteria are visible on the eye.
Tina Weatherby Carvalho, University of Hawaii at Manoa
View Media
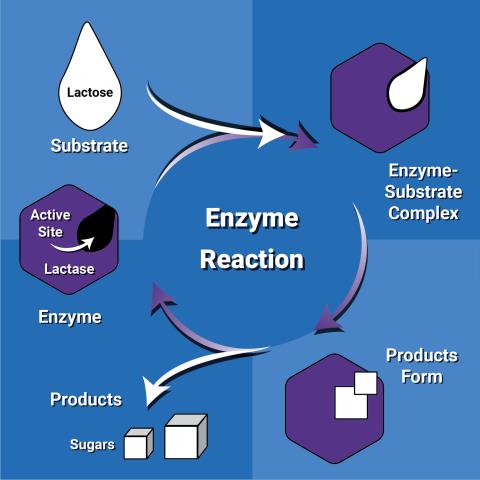
6604: Enzyme reaction
6604: Enzyme reaction
Enzymes speed up chemical reactions by reducing the amount of energy needed for the reactions. The substrate (lactose) binds to the active site of the enzyme (lactase) and is converted into products (sugars).
NIGMS
View Media

3479: Electrode probe on mouse Huntington's muscle cell
3479: Electrode probe on mouse Huntington's muscle cell
Using an electrode, researchers apply an electrical pulse onto a piece of muscle tissue affected by Huntington's disease.
Grigor Varuzhanyan and Andrew A. Voss, California State Polytechnic University
View Media
6790: Cell division and cell death
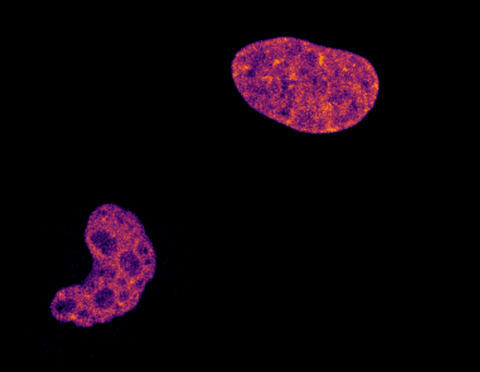
6790: Cell division and cell death
Two cells over a 2-hour period. The one on the bottom left goes through programmed cell death, also known as apoptosis. The one on the top right goes through cell division, also called mitosis. This video was captured using a confocal microscope.
Dylan T. Burnette, Vanderbilt University School of Medicine.
View Media
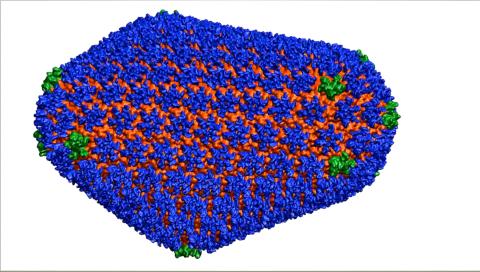

3477: HIV Capsid
3477: HIV Capsid
This image is a computer-generated model of the approximately 4.2 million atoms of the HIV capsid, the shell that contains the virus' genetic material. Scientists determined the exact structure of the capsid and the proteins that it's made of using a variety of imaging techniques and analyses. They then entered these data into a supercomputer that produced the atomic-level image of the capsid. This structural information could be used for developing drugs that target the capsid, possibly leading to more effective therapies. Related to image 6601.
Juan R. Perilla and the Theoretical and Computational Biophysics Group, University of Illinois at Urbana-Champaign
View Media
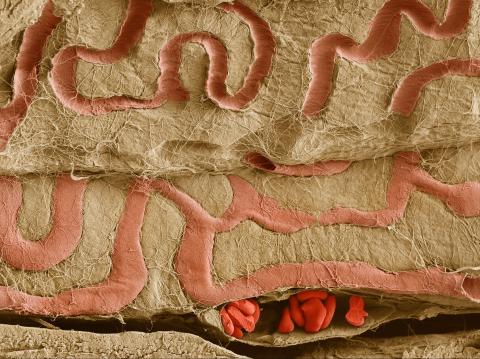
3739: Scanning electron microscopy of the ECM on the surface of a calf muscle
3739: Scanning electron microscopy of the ECM on the surface of a calf muscle
This image shows the extracellular matrix (ECM) on the surface of a soleus (lower calf) muscle in light brown and blood vessels in pink. Near the bottom of the photo, a vessel is opened up to reveal red blood cells. Scientists know less about the ECM in muscle than in other tissues, but it's increasingly clear that the ECM is critical to muscle function, and disruption of the ECM has been associated with many muscle disorders. The ECM in muscles stores and releases growth factors, suggesting that it might play a role in cellular communication.
Tom Deerinck, National Center for Microscopy and Imaging Research (NCMIR)
View Media
3486: Apoptosis reversed
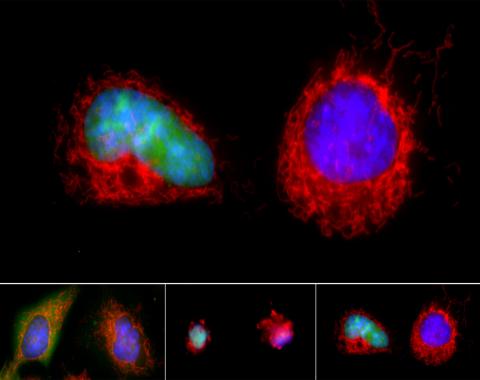
3486: Apoptosis reversed
Two healthy cells (bottom, left) enter into apoptosis (bottom, center) but spring back to life after a fatal toxin is removed (bottom, right; top).
Hogan Tang of the Denise Montell Lab, Johns Hopkins University School of Medicine
View Media
6748: Human retinal organoid
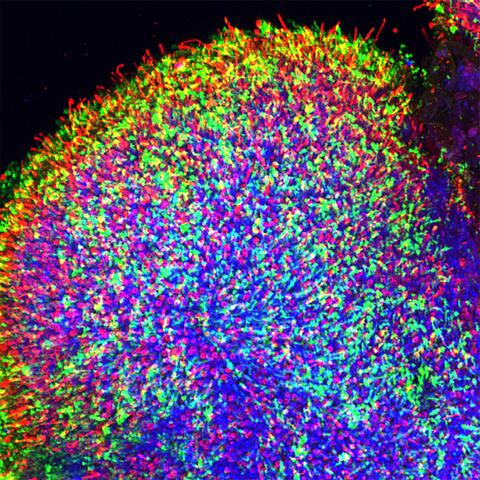
6748: Human retinal organoid
A replica of a human retina grown from stem cells. It shows rod photoreceptors (nerve cells responsible for dark vision) in green and red/green cones (nerve cells responsible for red and green color vision) in red. The cell nuclei are stained blue. This image was captured using a confocal microscope.
Kevin Eliceiri, University of Wisconsin-Madison.
View Media
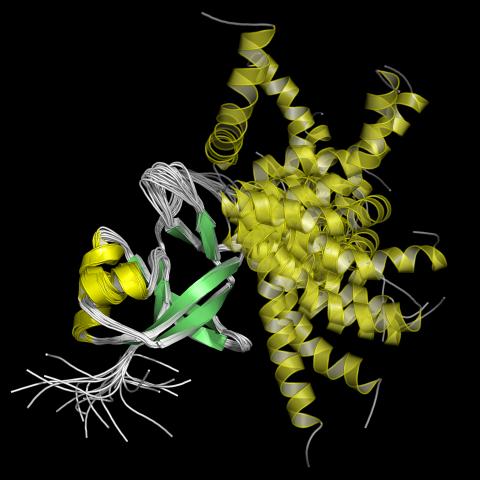
2339: Protein from Arabidopsis thaliana
2339: Protein from Arabidopsis thaliana
NMR solution structure of a plant protein that may function in host defense. This protein was expressed in a convenient and efficient wheat germ cell-free system. Featured as the June 2007 Protein Structure Initiative Structure of the Month.
Center for Eukaryotic Structural Genomics
View Media
