Switch to List View
Image and Video Gallery
This is a searchable collection of scientific photos, illustrations, and videos. The images and videos in this gallery are licensed under Creative Commons Attribution Non-Commercial ShareAlike 3.0. This license lets you remix, tweak, and build upon this work non-commercially, as long as you credit and license your new creations under identical terms.
3584: Rotavirus structure
3584: Rotavirus structure
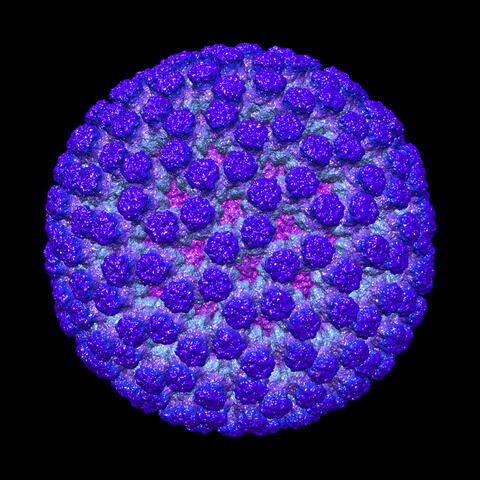
This image shows a computer-generated, three-dimensional map of the rotavirus structure. This virus infects humans and other animals and causes severe diarrhea in infants and young children. By the age of five, almost every child in the world has been infected with this virus at least once. Scientists have found a vaccine against rotavirus, so in the United States there are very few fatalities, but in developing countries and in places where the vaccine is unavailable, this virus is responsible for more than 200,000 deaths each year.
The rotavirus comprises three layers: the outer, middle and inner layers. On infection, the outer layer is removed, leaving behind a "double-layered particle." Researchers have studied the structure of this double-layered particle with a transmission electron microscope. Many images of the virus at a magnification of ~50,000x were acquired, and computational analysis was used to combine the individual particle images into a three-dimensional reconstruction.
The image was rendered by Melody Campbell (PhD student at TSRI). Work that led to the 3D map was published in Campbell et al. Movies of ice-embedded particles enhance resolution in electron cryo-microscopy. Structure. 2012;20(11):1823-8. PMCID: PMC3510009.
This image was part of the Life: Magnified exhibit that ran from June 3, 2014, to January 21, 2015, at Dulles International Airport.
The rotavirus comprises three layers: the outer, middle and inner layers. On infection, the outer layer is removed, leaving behind a "double-layered particle." Researchers have studied the structure of this double-layered particle with a transmission electron microscope. Many images of the virus at a magnification of ~50,000x were acquired, and computational analysis was used to combine the individual particle images into a three-dimensional reconstruction.
The image was rendered by Melody Campbell (PhD student at TSRI). Work that led to the 3D map was published in Campbell et al. Movies of ice-embedded particles enhance resolution in electron cryo-microscopy. Structure. 2012;20(11):1823-8. PMCID: PMC3510009.
This image was part of the Life: Magnified exhibit that ran from June 3, 2014, to January 21, 2015, at Dulles International Airport.
Bridget Carragher, The Scripps Research Institute, La Jolla, CA
View Media
6614: Los ritmos circadianos y el núcleo supraquiasmático
6614: Los ritmos circadianos y el núcleo supraquiasmático
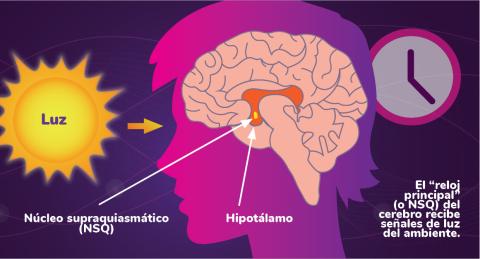
Los ritmos circadianos son cambios físicos, mentales y de comportamiento que siguen un ciclo de 24 horas. Los ritmos circadianos se ven influenciados por la luz y están regulados por el núcleo supraquiasmático del cerebro, a veces denominado el reloj principal.
Vea 6613 para la versión en inglés de esta infografía.
Vea 6613 para la versión en inglés de esta infografía.
NIGMS
View Media
6661: Zebrafish embryo showing vasculature
6661: Zebrafish embryo showing vasculature
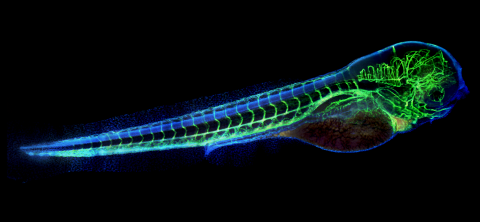
A zebrafish embryo. The blue areas are cell bodies, the green lines are blood vessels, and the red glow is blood. This image was created by stitching together five individual images captured with a hyperspectral multipoint confocal fluorescence microscope that was developed at the Eliceiri Lab.
Kevin Eliceiri, University of Wisconsin-Madison.
View Media

6752: Petri dish
6752: Petri dish
The white circle in this image is a Petri dish, named for its inventor, Julius Richard Petri. These dishes are one of the most common pieces of equipment in biology labs, where researchers use them to grow cells.
H. Robert Horvitz and Dipon Ghosh, Massachusetts Institute of Technology.
View Media
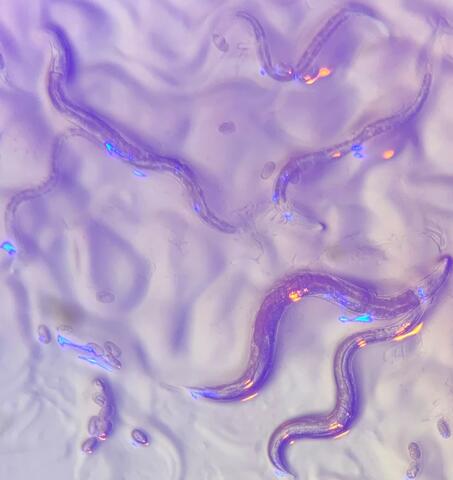
6750: C. elegans with blue and yellow lights in the background
6750: C. elegans with blue and yellow lights in the background
These microscopic roundworms, called Caenorhabditis elegans, lack eyes and the opsin proteins used by visual systems to detect colors. However, researchers found that the worms can still sense the color of light in a way that enables them to avoid pigmented toxins made by bacteria. This image was captured using a stereo microscope.
H. Robert Horvitz and Dipon Ghosh, Massachusetts Institute of Technology.
View Media

5815: Introduction to Genome Editing Using CRISPR/Cas9
5815: Introduction to Genome Editing Using CRISPR/Cas9
Genome editing using CRISPR/Cas9 is a rapidly expanding field of scientific research with emerging applications in disease treatment, medical therapeutics and bioenergy, just to name a few. This technology is now being used in laboratories all over the world to enhance our understanding of how living biological systems work, how to improve treatments for genetic diseases and how to develop energy solutions for a better future.
Janet Iwasa
View Media
5887: Plasma-Derived Membrane Vesicles
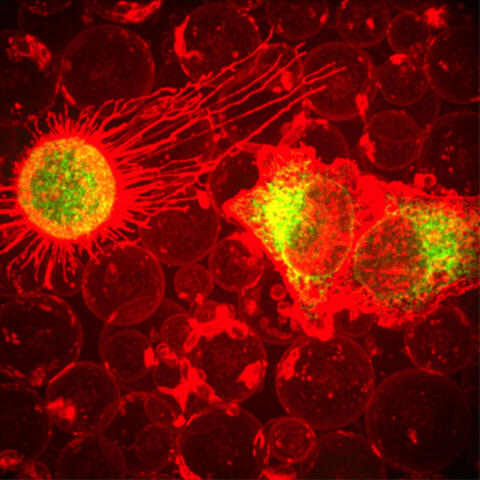
5887: Plasma-Derived Membrane Vesicles
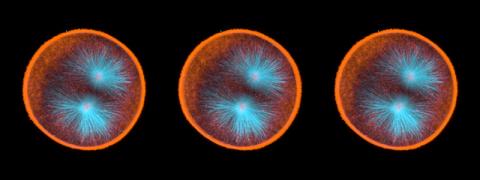
This fiery image doesn’t come from inside a bubbling volcano. Instead, it shows animal cells caught in the act of making bubbles, or blebbing. Some cells regularly pinch off parts of their membranes to produce bubbles filled with a mix of proteins and fats. The bubbles (red) are called plasma-derived membrane vesicles, or PMVs, and can travel to other parts of the body where they may aid in cell-cell communication. The University of Texas, Austin, researchers responsible for this photo are exploring ways to use PMVs to deliver medicines to precise locations in the body.
This image, entered in the Biophysical Society’s 2017 Art of Science Image contest, used two-channel spinning disk confocal fluorescence microscopy. It was also featured in the NIH Director’s Blog in May 2017.
This image, entered in the Biophysical Society’s 2017 Art of Science Image contest, used two-channel spinning disk confocal fluorescence microscopy. It was also featured in the NIH Director’s Blog in May 2017.
Jeanne Stachowiak, University of Texas at Austin
View Media
6993: RNA polymerase
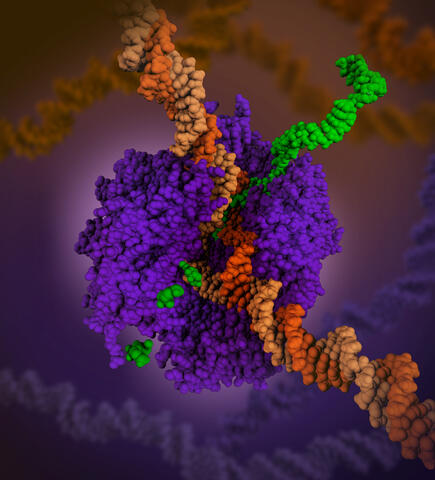
6993: RNA polymerase
RNA polymerase (purple) is a complex enzyme at the heart of transcription. During this process, the enzyme unwinds the DNA double helix and uses one strand (darker orange) as a template to create the single-stranded messenger RNA (green), later used by ribosomes for protein synthesis.
From the RNA polymerase II elongation complex of Saccharomyces cerevisiae (PDB entry 1I6H) as seen in PDB-101's What is a Protein? video.
From the RNA polymerase II elongation complex of Saccharomyces cerevisiae (PDB entry 1I6H) as seen in PDB-101's What is a Protein? video.
Amy Wu and Christine Zardecki, RCSB Protein Data Bank.
View Media
2740: Early life of a protein
2740: Early life of a protein
This illustration represents the early life of a protein—specifically, apomyoglobin—as it is synthesized by a ribosome and emerges from the ribosomal tunnel, which contains the newly formed protein's conformation. The synthesis occurs in the complex swirl of the cell medium, filled with interactions among many molecules. Researchers in Silvia Cavagnero's laboratory are studying the structure and dynamics of newly made proteins and polypeptides using spectroscopic and biochemical techniques.
Silvia Cavagnero, University of Wisconsin, Madison
View Media
2432: ARTS triggers apoptosis
2432: ARTS triggers apoptosis
Cell showing overproduction of the ARTS protein (red). ARTS triggers apoptosis, as shown by the activation of caspase-3 (green) a key tool in the cell's destruction. The nucleus is shown in blue. Image is featured in October 2015 Biomedical Beat blog post Cool Images: A Halloween-Inspired Cell Collection.
Hermann Steller, Rockefeller University
View Media
2314: Finding one bug
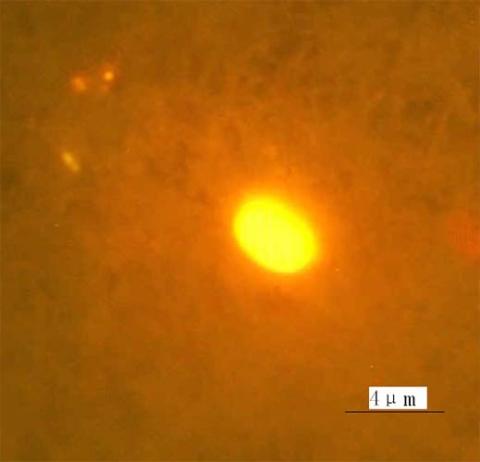
2314: Finding one bug
A nanometer-sized biosensor can detect a single deadly bacterium in tainted ground beef. How? Researchers attached nanoparticles, each packed with thousands of dye molecules, to an antibody that recognizes the microbe E. coli O157:H7. When the nanoball-antibody combo comes into contact with the E. coli bacterium, it glows. Here is the transition, a single bacterial cell glows brightly when it encounters nanoparticle-antibody biosensors, each packed with thousands of dye molecules.
Weihong Tan, University of Florida in Gainesville
View Media
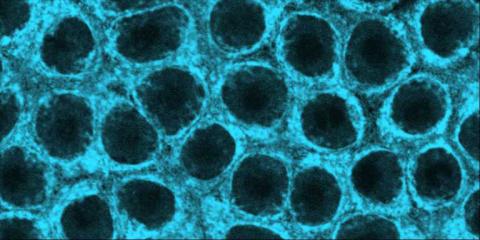
2324: Movements of myosin
2324: Movements of myosin
Inside the fertilized egg cell of a fruit fly, we see a type of myosin (related to the protein that helps muscles contract) made to glow by attaching a fluorescent protein. After fertilization, the myosin proteins are distributed relatively evenly near the surface of the embryo. The proteins temporarily vanish each time the cells' nuclei--initially buried deep in the cytoplasm--divide. When the multiplying nuclei move to the surface, they shift the myosin, producing darkened holes. The glowing myosin proteins then gather, contract, and start separating the nuclei into their own compartments.
Victoria Foe, University of Washington
View Media
2426: Zinc finger
2426: Zinc finger
The structure of a gene-regulating zinc finger protein bound to DNA.
Jeremy M. Berg, National Institute of General Medical Sciences
View Media
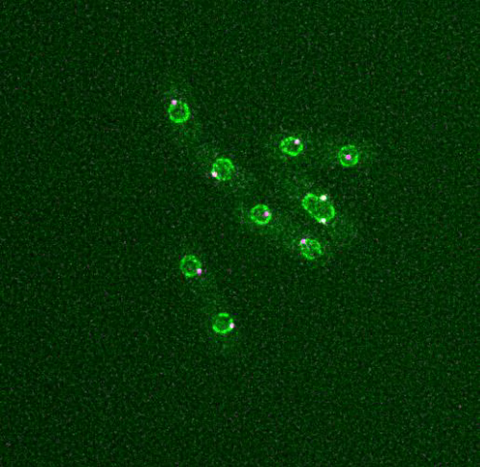
6795: Dividing yeast cells with nuclear envelopes and spindle pole bodies
6795: Dividing yeast cells with nuclear envelopes and spindle pole bodies
Time-lapse video of yeast cells undergoing cell division. Nuclear envelopes are shown in green, and spindle pole bodies, which help pull apart copied genetic information, are shown in magenta. This video was captured using wide-field microscopy with deconvolution.
Related to images 6791, 6792, 6793, 6794, 6797, 6798, and video 6796.
Related to images 6791, 6792, 6793, 6794, 6797, 6798, and video 6796.
Alaina Willet, Kathy Gould’s lab, Vanderbilt University.
View Media

6903: Young squids
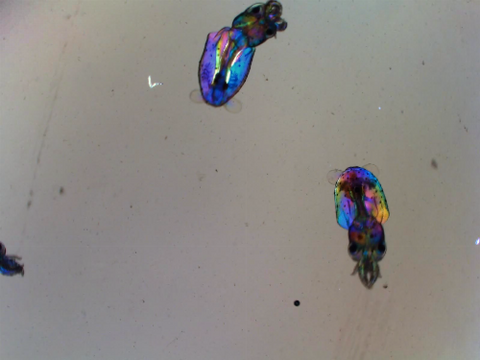
6903: Young squids
Real-time movie of young squids. Squids are often used as research organisms due to having the largest nervous system of any invertebrate, complex behaviors like instantaneous camouflage, and other unique traits.
This video was taken with polychromatic polarization microscope, as described in the Scientific Reports paper “Polychromatic Polarization Microscope: Bringing Colors to a Colorless World” by Shribak. The color is generated by interaction of white polarized light with the squid’s transparent soft tissue. The tissue works as a living tunable spectral filter, and the transmission band depends on the molecular orientation. When the young squid is moving, the tissue orientation changes, and its color shifts accordingly.
This video was taken with polychromatic polarization microscope, as described in the Scientific Reports paper “Polychromatic Polarization Microscope: Bringing Colors to a Colorless World” by Shribak. The color is generated by interaction of white polarized light with the squid’s transparent soft tissue. The tissue works as a living tunable spectral filter, and the transmission band depends on the molecular orientation. When the young squid is moving, the tissue orientation changes, and its color shifts accordingly.
Michael Shribak, Marine Biological Laboratory/University of Chicago.
View Media
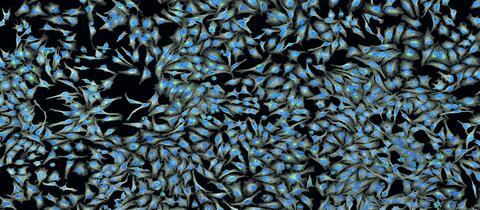
5761: A panorama view of cells
5761: A panorama view of cells
This photograph shows a panoramic view of HeLa cells, a cell line many researchers use to study a large variety of important research questions. The cells' nuclei containing the DNA are stained in blue and the cells' cytoskeletons in gray.
Tom Deerinck, National Center for Microscopy and Imaging Research
View Media
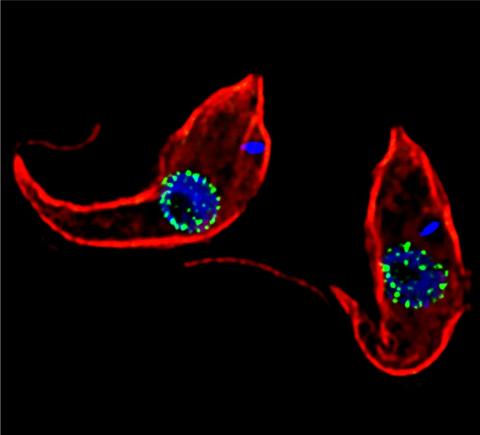
3765: Trypanosoma brucei, the cause of sleeping sickness
3765: Trypanosoma brucei, the cause of sleeping sickness
Trypanosoma brucei is a single-cell parasite that causes sleeping sickness in humans. Scientists have been studying trypanosomes for some time because of their negative effects on human and also animal health, especially in sub-Saharan Africa. Moreover, because these organisms evolved on a separate path from those of animals and plants more than a billion years ago, researchers study trypanosomes to find out what traits they may harbor that are common to or different from those of other eukaryotes (i.e., those organisms having a nucleus and mitochondria). This image shows the T. brucei cell membrane in red, the DNA in the nucleus and kinetoplast (a structure unique to protozoans, including trypanosomes, which contains mitochondrial DNA) in blue and nuclear pore complexes (which allow molecules to pass into or out of the nucleus) in green. Scientists have found that the trypanosome nuclear pore complex has a unique mechanism by which it attaches to the nuclear envelope. In addition, the trypanosome nuclear pore complex differs from those of other eukaryotes because its components have a near-complete symmetry, and it lacks almost all of the proteins that in other eukaryotes studied so far are required to assemble the pore.
Michael Rout, Rockefeller University
View Media
6553: Floral pattern in a mixture of two bacterial species, Acinetobacter baylyi and Escherichia coli, grown on a semi-solid agar for 48 hours (photo 1)
6553: Floral pattern in a mixture of two bacterial species, Acinetobacter baylyi and Escherichia coli, grown on a semi-solid agar for 48 hours (photo 1)
Floral pattern emerging as two bacterial species, motile Acinetobacter baylyi (red) and non-motile Escherichia coli (green), are grown together for 48 hours on 1% agar surface from a small inoculum in the center of a Petri dish.
See 6557 for a photo of this process at 24 hours on 0.75% agar surface.
See 6555 for another photo of this process at 48 hours on 1% agar surface.
See 6556 for a photo of this process at 72 hours on 0.5% agar surface.
See 6550 for a video of this process.
See 6557 for a photo of this process at 24 hours on 0.75% agar surface.
See 6555 for another photo of this process at 48 hours on 1% agar surface.
See 6556 for a photo of this process at 72 hours on 0.5% agar surface.
See 6550 for a video of this process.
L. Xiong et al, eLife 2020;9: e48885
View Media
3389: NCMIR Intestine-1
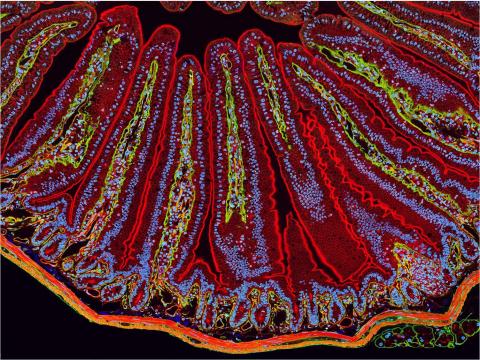
3389: NCMIR Intestine-1
The small intestine is where most of our nutrients from the food we eat are absorbed into the bloodstream. The walls of the intestine contain small finger-like projections called villi which increase the organ's surface area, enhancing nutrient absorption. It consists of the duodenum, which connects to the stomach, the jejenum and the ileum, which connects with the large intestine. Related to image 3390.
Tom Deerinck, National Center for Microscopy and Imaging Research (NCMIR)
View Media

1089: Natcher Building 09
1089: Natcher Building 09
NIGMS staff are located in the Natcher Building on the NIH campus.
Alisa Machalek, National Institute of General Medical Sciences
View Media
3580: V. Cholerae Biofilm
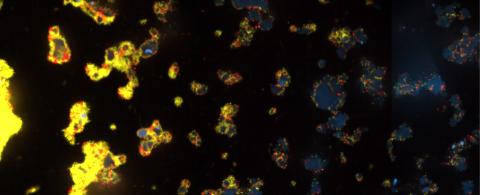
3580: V. Cholerae Biofilm
Industrious V. cholerae bacteria (yellow) tend to thrive in denser biofilms (left) while moochers (red) thrive in weaker biofilms (right). More information about the research behind this image can be found in a Biomedical Beat Blog posting from February 2014.
View Media
2771: Self-organizing proteins
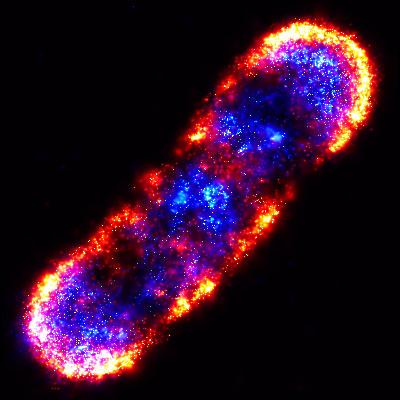
2771: Self-organizing proteins
Under the microscope, an E. coli cell lights up like a fireball. Each bright dot marks a surface protein that tells the bacteria to move toward or away from nearby food and toxins. Using a new imaging technique, researchers can map the proteins one at a time and combine them into a single image. This lets them study patterns within and among protein clusters in bacterial cells, which don't have nuclei or organelles like plant and animal cells. Seeing how the proteins arrange themselves should help researchers better understand how cell signaling works.
View Media
2541: Nucleotides make up DNA
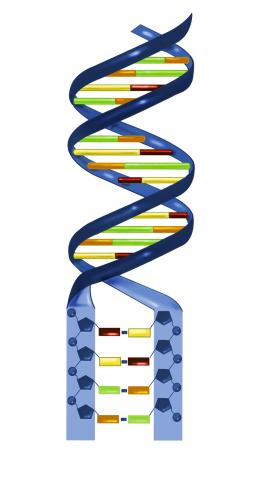
2541: Nucleotides make up DNA
DNA consists of two long, twisted chains made up of nucleotides. Each nucleotide contains one base, one phosphate molecule, and the sugar molecule deoxyribose. The bases in DNA nucleotides are adenine, thymine, cytosine, and guanine. See image 2542 for a labeled version of this illustration. Featured in The New Genetics.
Crabtree + Company
View Media
3624: Fibroblasts with nuclei in blue, energy factories in green and the actin cytoskeleton in red
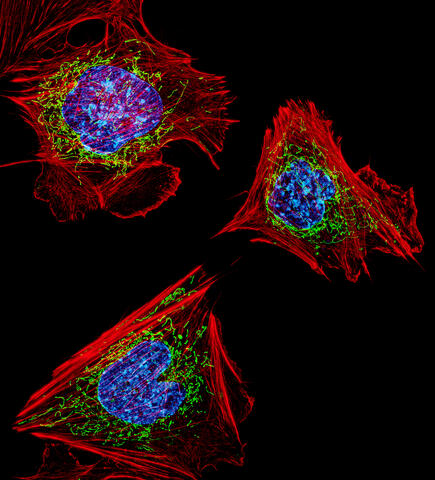
3624: Fibroblasts with nuclei in blue, energy factories in green and the actin cytoskeleton in red
The cells shown here are fibroblasts, one of the most common cells in mammalian connective tissue. These particular cells were taken from a mouse embryo. Scientists used them to test the power of a new microscopy technique that offers vivid views of the inside of a cell. The DNA within the nucleus (blue), mitochondria (green), and actin filaments in the cellular skeleton (red) are clearly visible.
This image was part of the Life: Magnified exhibit that ran from June 3, 2014, to January 21, 2015, at Dulles International Airport.
This image was part of the Life: Magnified exhibit that ran from June 3, 2014, to January 21, 2015, at Dulles International Airport.
Dylan Burnette, NICHD
View Media
3421: Structure of Glutamate Dehydrogenase
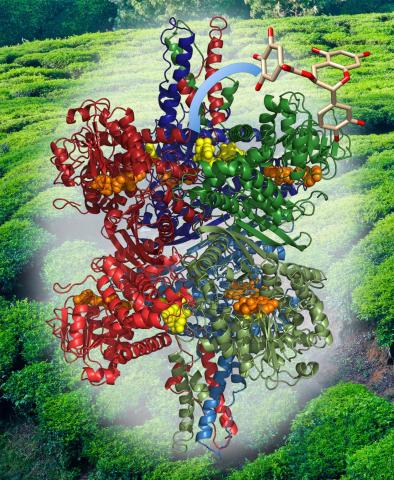
3421: Structure of Glutamate Dehydrogenase
Some children are born with a mutation in a regulatory site on this enzyme that causes them to over-secrete insulin when they consume protein. We found that a compound from green tea (shown in the stick figure and by the yellow spheres on the enzyme) is able to block this hyperactivity when given to animals with this disorder.
Judy Coyle, Donald Danforth Plant Science Center
View Media
3758: Dengue virus membrane protein structure
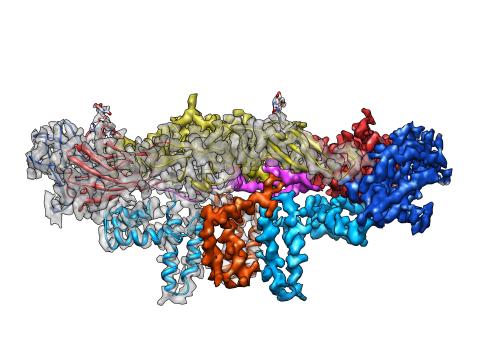
3758: Dengue virus membrane protein structure
Dengue virus is a mosquito-borne illness that infects millions of people in the tropics and subtropics each year. Like many viruses, dengue is enclosed by a protective membrane. The proteins that span this membrane play an important role in the life cycle of the virus. Scientists used cryo-EM to determine the structure of a dengue virus at a 3.5-angstrom resolution to reveal how the membrane proteins undergo major structural changes as the virus matures and infects a host. The image shows a side view of the structure of a protein composed of two smaller proteins, called E and M. Each E and M contributes two molecules to the overall protein structure (called a heterotetramer), which is important for assembling and holding together the viral membrane, i.e., the shell that surrounds the genetic material of the dengue virus. The dengue protein's structure has revealed some portions in the protein that might be good targets for developing medications that could be used to combat dengue virus infections. For more on cryo-EM see the blog post Cryo-Electron Microscopy Reveals Molecules in Ever Greater Detail. You can watch a rotating view of the dengue virus surface structure in video 3748.
Hong Zhou, UCLA
View Media
5772: Confocal microscopy image of two Drosophila ovarioles
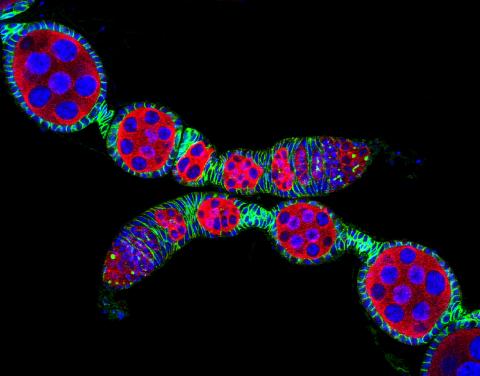
5772: Confocal microscopy image of two Drosophila ovarioles
Ovarioles in female insects are tubes in which egg cells (called oocytes) form at one end and complete their development as they reach the other end of the tube. This image, taken with a confocal microscope, shows ovarioles in a very popular lab animal, the fruit fly Drosophila. The basic structure of ovarioles supports very rapid egg production, with some insects (like termites) producing several thousand eggs per day. Each insect ovary typically contains four to eight ovarioles, but this number varies widely depending on the insect species.
Scientists use insect ovarioles, for example, to study the basic processes that help various insects, including those that cause disease (like some mosquitos and biting flies), reproduce very quickly.
Scientists use insect ovarioles, for example, to study the basic processes that help various insects, including those that cause disease (like some mosquitos and biting flies), reproduce very quickly.
2004 Olympus BioScapes Competition
View Media
3483: Chang Shan
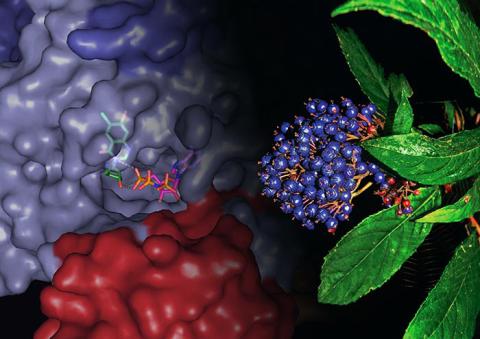
3483: Chang Shan
For thousands of years, Chinese herbalists have treated malaria using Chang Shan, a root extract from a type of hydrangea that grows in Tibet and Nepal. Recent studies have suggested Chang Shan can also reduce scar formation, treat multiple sclerosis and even slow cancer progression.
Paul Schimmel Lab, Scripps Research Institute
View Media
3414: X-ray co-crystal structure of Src kinase bound to a DNA-templated macrocycle inhibitor 2
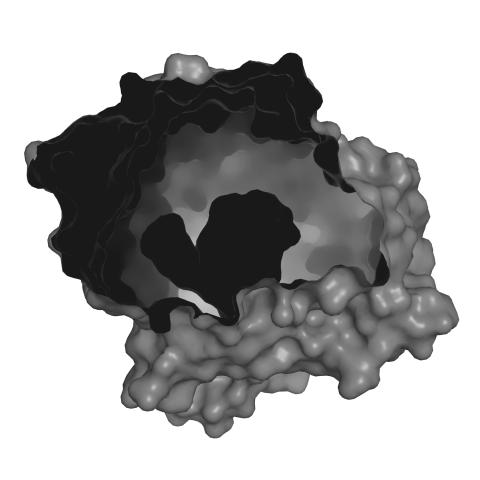
3414: X-ray co-crystal structure of Src kinase bound to a DNA-templated macrocycle inhibitor 2
X-ray co-crystal structure of Src kinase bound to a DNA-templated macrocycle inhibitor. Related to 3413, 3415, 3416, 3417, 3418, and 3419.
Markus A. Seeliger, Stony Brook University Medical School and David R. Liu, Harvard University
View Media
2342: Protein from E. faecalis
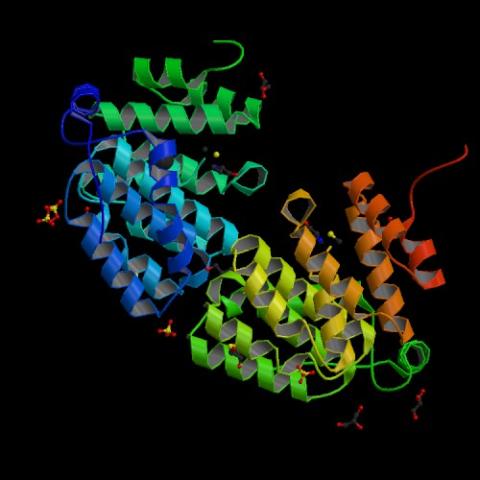
2342: Protein from E. faecalis
X-ray structure of a DNA repair enzyme superfamily representative from the human gastrointestinal bacterium Enterococcus faecalis. European scientists used this structure to generate homologous structures. Featured as the May 2007 Protein Structure Initiative Structure of the Month.
Midwest Center for Structural Genomics
View Media

2735: Network Map
2735: Network Map
This network map shows the overlap (green) between the long QT syndrome (yellow) and epilepsy (blue) protein-interaction neighborhoods located within the human interactome. Researchers have learned to integrate genetic, cellular and clinical information to find out why certain medicines can trigger fatal heart arrhythmias. Featured in Computing Life magazine.
Seth Berger, Mount Sinai School of Medicine
View Media
2784: Microtubule dynamics in real time
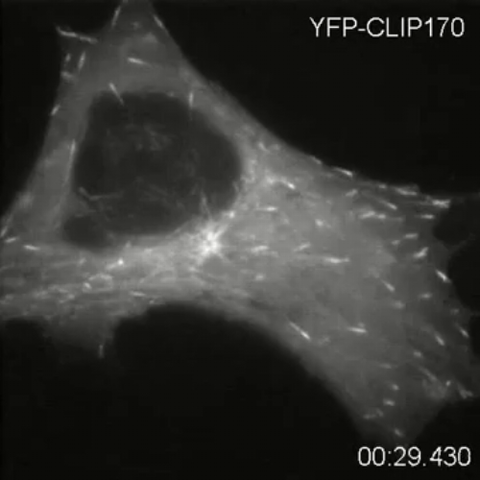
2784: Microtubule dynamics in real time
Cytoplasmic linker protein (CLIP)-170 is a microtubule plus-end-tracking protein that regulates microtubule dynamics and links microtubule ends to different intracellular structures. In this movie, the gene for CLIP-170 has been fused with green fluorescent protein (GFP). When the protein is expressed in cells, the activities can be monitored in real time. Here, you can see CLIP-170 streaming towards the edges of the cell.
Gary Borisy, Marine Biology Laboratory
View Media
3363: Dopamine D3 receptor
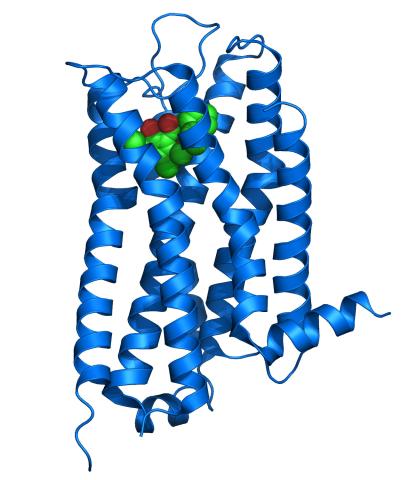
3363: Dopamine D3 receptor
The receptor is shown bound to an antagonist, eticlopride
Raymond Stevens, The Scripps Research Institute
View Media
6606: Cryo-ET cross-section of the Golgi apparatus
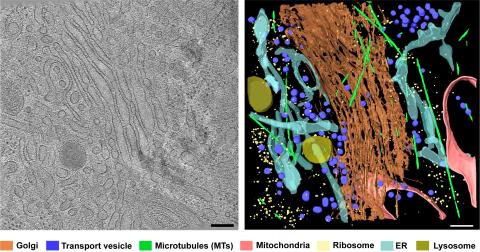
6606: Cryo-ET cross-section of the Golgi apparatus
On the left, a cross-section slice of a rat pancreas cell captured using cryo-electron tomography (cryo-ET). On the right, a 3D, color-coded version of the image highlighting cell structures. Visible features include the folded sacs of the Golgi apparatus (copper), transport vesicles (medium-sized dark-blue circles), microtubules (neon green), ribosomes (small pale-yellow circles), and lysosomes (large yellowish-green circles). Black line (bottom right of the left image) represents 200 nm. This image is a still from video 6609.
Xianjun Zhang, University of Southern California.
View Media
2553: Alternative splicing (with labels)
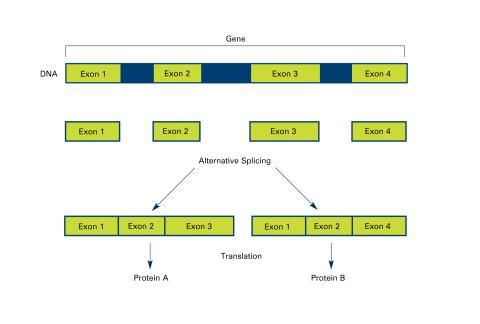
2553: Alternative splicing (with labels)
Arranging exons in different patterns, called alternative splicing, enables cells to make different proteins from a single gene. Featured in The New Genetics.
See image 2552 for an unlabeled version of this illustration.
See image 2552 for an unlabeled version of this illustration.
Crabtree + Company
View Media

6589: Cell-like compartments emerging from scrambled frog eggs 3
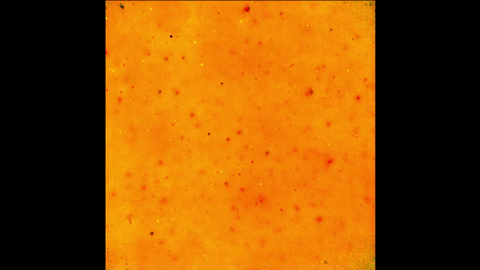
6589: Cell-like compartments emerging from scrambled frog eggs 3
Cell-like compartments spontaneously emerge from scrambled frog eggs. Endoplasmic reticulum (red) and microtubules (green) are visible. Video created using epifluorescence microscopy.
For more photos of cell-like compartments from frog eggs view: 6584, 6585, 6586, 6591, 6592, and 6593.
For videos of cell-like compartments from frog eggs view: 6587, 6588, and 6590.
Xianrui Cheng, Stanford University School of Medicine.
View Media

2683: GFP sperm
2683: GFP sperm
Fruit fly sperm cells glow bright green when they express the gene for green fluorescent protein (GFP).
View Media
3330: mDia1 antibody staining-01
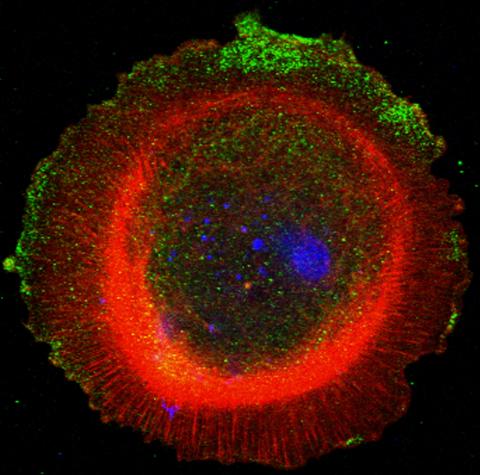
3330: mDia1 antibody staining-01
Cells move forward with lamellipodia and filopodia supported by networks and bundles of actin filaments. Proper, controlled cell movement is a complex process. Recent research has shown that an actin-polymerizing factor called the Arp2/3 complex is the key component of the actin polymerization engine that drives amoeboid cell motility. ARPC3, a component of the Arp2/3 complex, plays a critical role in actin nucleation. In this photo, the ARPC3+/+ fibroblast cells were fixed and stained with Alexa 546 phalloidin for F-actin (red), mDia1 (green), and DAPI to visualize the nucleus (blue). mDia1 is localized at the lamellipodia of ARPC3+/+ fibroblast cells. Related to images 3328, 3329, 3331, 3332, and 3333.
Rong Li and Praveen Suraneni, Stowers Institute for Medical Research
View Media
1278: Golgi theories
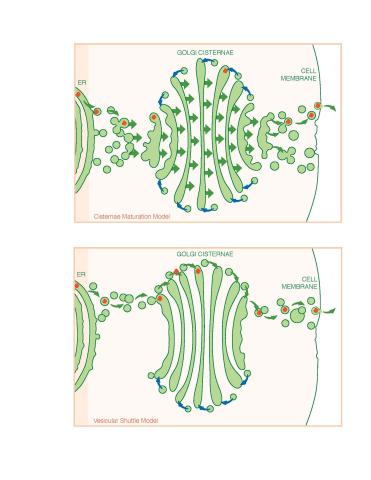
1278: Golgi theories
Two models for how material passes through the Golgi apparatus: the vesicular shuttle model and the cisternae maturation model.
Judith Stoffer
View Media
3478: DDR2 Receptors Attach to Collagen in Breast Tumor
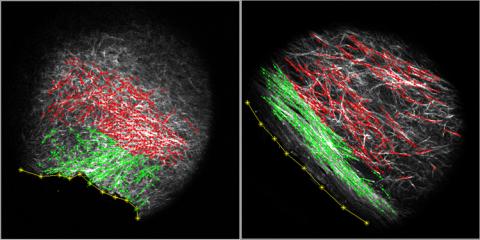
3478: DDR2 Receptors Attach to Collagen in Breast Tumor
On the left, the boundary of a breast tumor (yellow) attaches to collagen fibers that are closest to it (green) using DDR2. On the right, a tumor without DDR2 remains disconnected from the collagen.
Callie Corsa and Suzanne Ponik, Washington University School of Medicine in St. Louis
View Media

6796: Dividing yeast cells with spindle pole bodies and contractile rings
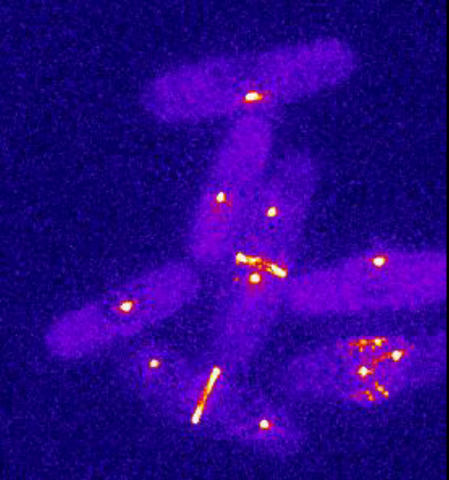
6796: Dividing yeast cells with spindle pole bodies and contractile rings
During cell division, spindle pole bodies (glowing dots) move toward the ends of yeast cells to separate copied genetic information. Contractile rings (glowing bands) form in cells’ middles and constrict to help them split. This time-lapse video was captured using wide-field microscopy with deconvolution.
Related to images 6791, 6792, 6793, 6794, 6797, 6798, and video 6795.
Related to images 6791, 6792, 6793, 6794, 6797, 6798, and video 6795.
Alaina Willet, Kathy Gould’s lab, Vanderbilt University.
View Media
2574: Simulation of uncontrolled avian flu outbreak
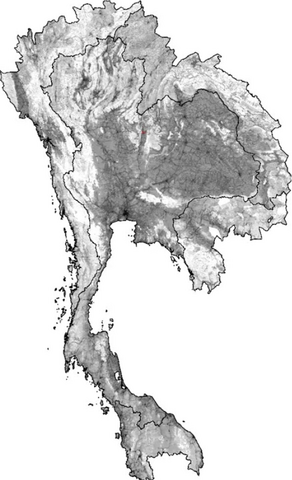
2574: Simulation of uncontrolled avian flu outbreak
This video simulation shows what an uncontrolled outbreak of transmissible avian flu among people living in Thailand might look like. Red indicates new cases while green indicates areas where the epidemic has finished. The video shows the spread of infection and recovery over 300 days in Thailand and neighboring countries.
Neil M. Ferguson, Imperial College London
View Media
2299: 2-D NMR
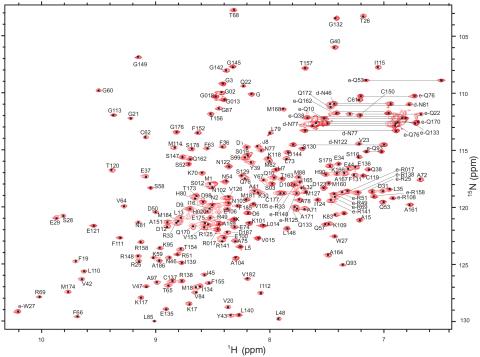
2299: 2-D NMR
A two-dimensional NMR spectrum of a protein, in this case a 2D 1H-15N HSQC NMR spectrum of a 228 amino acid DNA/RNA-binding protein.
Dr. Xiaolian Gao's laboratory at the University of Houston
View Media
6995: Measles virus
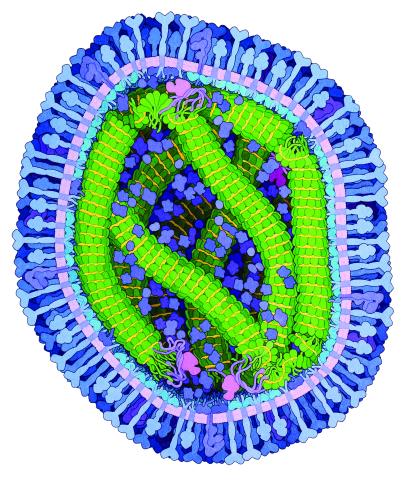
6995: Measles virus
A cross section of the measles virus in which six proteins work together to infect cells. The measles virus is extremely infectious; 9 out of 10 people exposed will contract the disease. Fortunately, an effective vaccine protects against infection.
For a zoomed-in look at the six important proteins, see Measles Virus Proteins.
For a zoomed-in look at the six important proteins, see Measles Virus Proteins.
Amy Wu and Christine Zardecki, RCSB Protein Data Bank.
View Media
1022: Lily mitosis 09
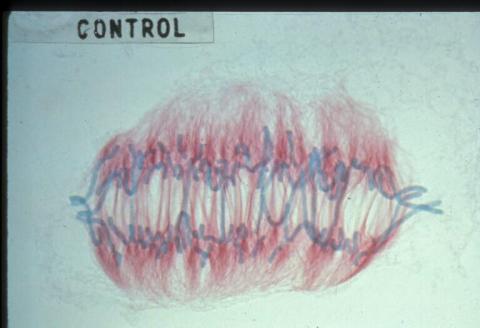
1022: Lily mitosis 09
A light microscope image of a cell from the endosperm of an African globe lily (Scadoxus katherinae). This is one frame of a time-lapse sequence that shows cell division in action. The lily is considered a good organism for studying cell division because its chromosomes are much thicker and easier to see than human ones. Staining shows microtubules in red and chromosomes in blue. Here, condensed chromosomes are clearly visible and are starting to separate to form two new cells.
Andrew S. Bajer, University of Oregon, Eugene
View Media
3268: Fluorescent E. coli bacteria
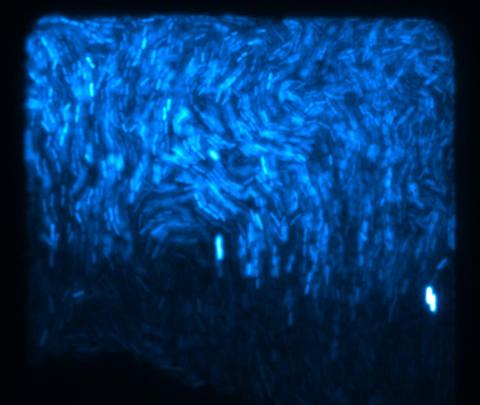
3268: Fluorescent E. coli bacteria
Bioengineers were able to coax bacteria to blink in unison on microfluidic chips. They called each blinking bacterial colony a biopixel. Thousands of fluorescent E. coli bacteria, shown here, make up a biopixel. Related to images 3265 and 3266. From a UC San Diego news release, "Researchers create living 'neon signs' composed of millions of glowing bacteria."
Jeff Hasty Lab, UC San Diego
View Media
3750: A dynamic model of the DNA helicase protein complex
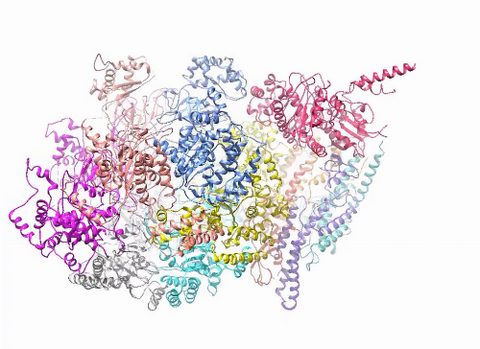
3750: A dynamic model of the DNA helicase protein complex
This short video shows a model of the DNA helicase in yeast. This DNA helicase has 11 proteins that work together to unwind DNA during the process of copying it, called DNA replication. Scientists used a technique called cryo-electron microscopy (cryo-EM), which allowed them to study the helicase structure in solution rather than in static crystals. Cryo-EM in combination with computer modeling therefore allows researchers to see movements and other dynamic changes in the protein. The cryo-EM approach revealed the helicase structure at much greater resolution than could be obtained before. The researchers think that a repeated motion within the protein as shown in the video helps it move along the DNA strand. To read more about DNA helicase and this proposed mechanism, see this news release by Brookhaven National Laboratory.
Huilin Li, Stony Brook University
View Media
6762: CCP enzyme
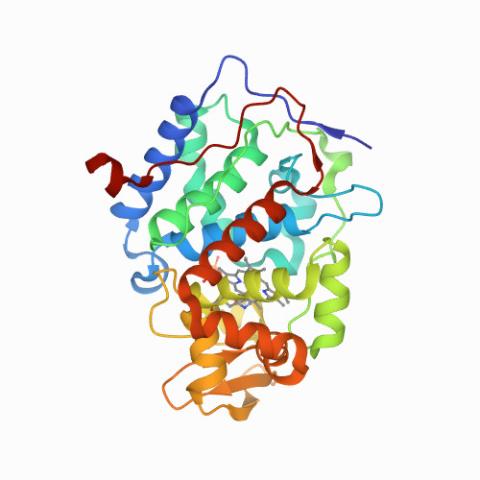
6762: CCP enzyme
The enzyme CCP is found in the mitochondria of baker’s yeast. Scientists study the chemical reactions that CCP triggers, which involve a water molecule, iron, and oxygen. This structure was determined using an X-ray free electron laser.
Protein Data Bank.
View Media
2763: Fused, dicentric chromosomes
2763: Fused, dicentric chromosomes
This fused chromosome has two functional centromeres, shown as two sets of red and green dots. Centromeres are DNA/protein complexes that are key to splitting the chromosomes evenly during cell division. When dicentric chromosomes like this one are formed in a person, fertility problems or other difficulties may arise. Normal chromosomes carrying a single centromere (one set of red and green dots) are also visible in this image.
Beth A. Sullivan, Duke University
View Media