Switch to List View
Image and Video Gallery
This is a searchable collection of scientific photos, illustrations, and videos. The images and videos in this gallery are licensed under Creative Commons Attribution Non-Commercial ShareAlike 3.0. This license lets you remix, tweak, and build upon this work non-commercially, as long as you credit and license your new creations under identical terms.
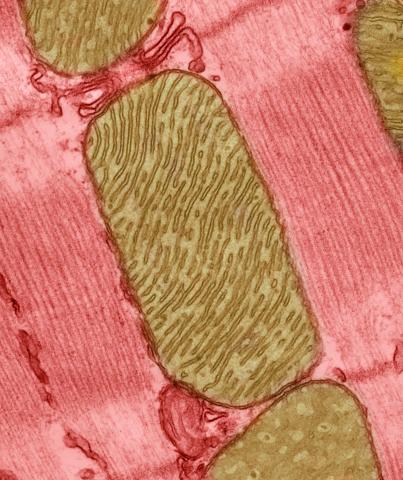
3664: Mitochondria from rat heart muscle cell_2
3664: Mitochondria from rat heart muscle cell_2
These mitochondria (brown) are from the heart muscle cell of a rat. Mitochondria have an inner membrane that folds in many places (and that appears here as striations). This folding vastly increases the surface area for energy production. Nearly all our cells have mitochondria. Related to image 3661.
National Center for Microscopy and Imaging Research
View Media

1287: Mitochondria
1287: Mitochondria
Bean-shaped mitochondria are cells' power plants. These organelles have their own DNA and replicate independently. The highly folded inner membranes are the site of energy generation.
Judith Stoffer
View Media
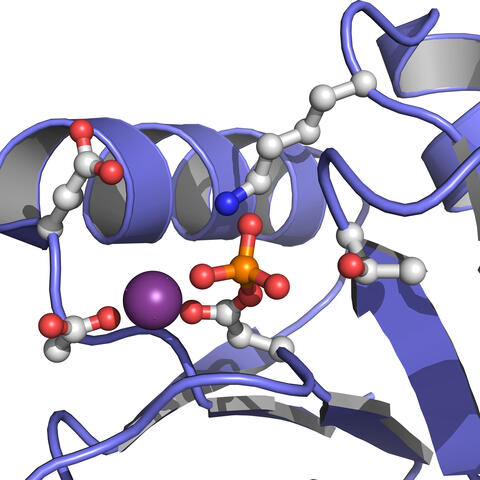
3412: Active Site of E. coli response regulator PhoB
3412: Active Site of E. coli response regulator PhoB
Active site of E. coli response regulator PhoB.
Ann Stock, Rutgers University
View Media
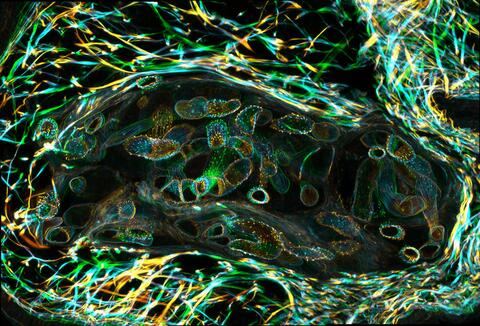
3627: Larvae from the parasitic worm that causes schistosomiasis
3627: Larvae from the parasitic worm that causes schistosomiasis
The parasitic worm that causes schistosomiasis hatches in water and grows up in a freshwater snail, as shown here. Once mature, the worm swims back into the water, where it can infect people through skin contact. Initially, an infected person might have a rash, itchy skin, or flu-like symptoms, but the real damage is done over time to internal organs.
This image was part of the Life: Magnified exhibit that ran from June 3, 2014, to January 21, 2015, at Dulles International Airport.
This image was part of the Life: Magnified exhibit that ran from June 3, 2014, to January 21, 2015, at Dulles International Airport.
Bo Wang and Phillip A. Newmark, University of Illinois at Urbana-Champaign, 2013 FASEB BioArt winner
View Media
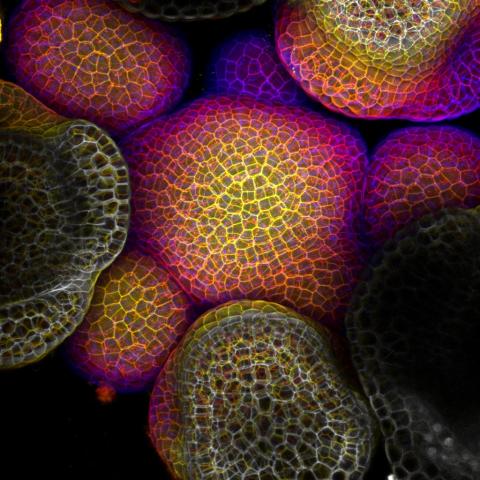
3606: Flower-forming cells in a small plant related to cabbage (Arabidopsis)
3606: Flower-forming cells in a small plant related to cabbage (Arabidopsis)
In plants, as in animals, stem cells can transform into a variety of different cell types. The stem cells at the growing tip of this Arabidopsis plant will soon become flowers. Arabidopsis is frequently studied by cellular and molecular biologists because it grows rapidly (its entire life cycle is only 6 weeks), produces lots of seeds, and has a genome that is easy to manipulate.
This image was part of the Life: Magnified exhibit that ran from June 3, 2014, to January 21, 2015, at Dulles International Airport.
This image was part of the Life: Magnified exhibit that ran from June 3, 2014, to January 21, 2015, at Dulles International Airport.
Arun Sampathkumar and Elliot Meyerowitz, California Institute of Technology
View Media
2739: Tetrapolar mitosis
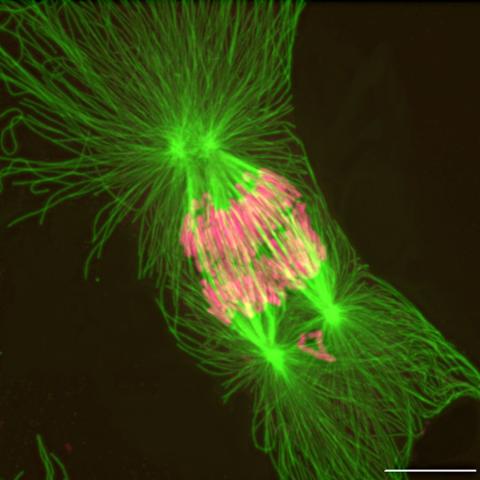
2739: Tetrapolar mitosis
This image shows an abnormal, tetrapolar mitosis. Chromosomes are highlighted pink. The cells shown are S3 tissue cultured cells from Xenopus laevis, African clawed frog.
Gary Gorbsky, Oklahoma Medical Research Foundation
View Media
2780: Arabidopsis leaf injected with a pathogen
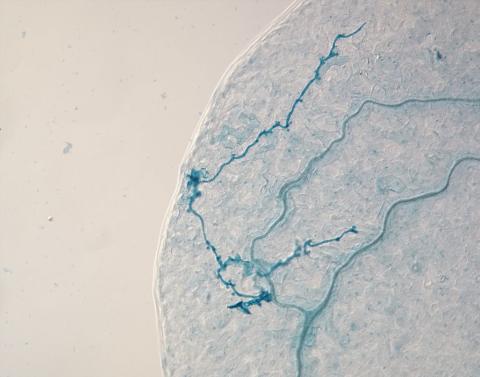
2780: Arabidopsis leaf injected with a pathogen
This is a magnified view of an Arabidopsis thaliana leaf eight days after being infected with the pathogen Hyaloperonospora arabidopsidis, which is closely related to crop pathogens that cause 'downy mildew' diseases. It is also more distantly related to the agent that caused the Irish potato famine. The veins of the leaf are light blue; in darker blue are the pathogen's hyphae growing through the leaf. The small round blobs along the length of the hyphae are called haustoria; each is invading a single plant cell to suck nutrients from the cell. Jeff Dangl and other NIGMS-supported researchers investigate how this pathogen and other like it use virulence mechanisms to suppress host defense and help the pathogens grow.
Jeff Dangl, University of North Carolina, Chapel Hill
View Media
1085: Natcher Building 05
1085: Natcher Building 05
NIGMS staff are located in the Natcher Building on the NIH campus.
Alisa Machalek, National Institute of General Medical Sciences
View Media

5815: Introduction to Genome Editing Using CRISPR/Cas9
5815: Introduction to Genome Editing Using CRISPR/Cas9
Genome editing using CRISPR/Cas9 is a rapidly expanding field of scientific research with emerging applications in disease treatment, medical therapeutics and bioenergy, just to name a few. This technology is now being used in laboratories all over the world to enhance our understanding of how living biological systems work, how to improve treatments for genetic diseases and how to develop energy solutions for a better future.
Janet Iwasa
View Media
3290: Three neurons and human ES cells
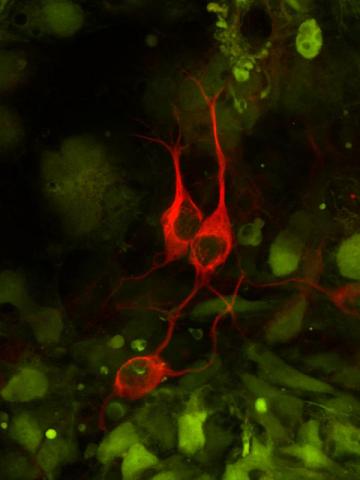
3290: Three neurons and human ES cells
The three neurons (red) visible in this image were derived from human embryonic stem cells. Undifferentiated stem cells are green here. Image and caption information courtesy of the California Institute for Regenerative Medicine.
Anirvan Ghosh lab, University of California, San Diego, via CIRM
View Media
3688: Brain cells in the hippocampus
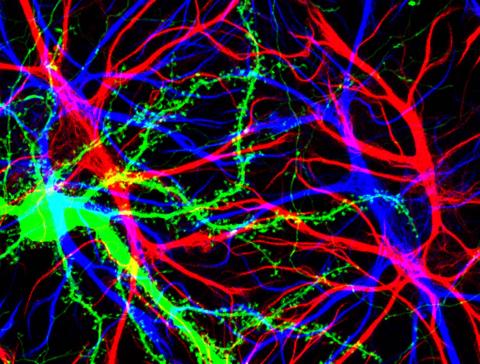
3688: Brain cells in the hippocampus
Hippocampal cells in culture with a neuron in green, showing hundreds of the small protrusions known as dendritic spines. The dendrites of other neurons are labeled in blue, and adjacent glial cells are shown in red.
Shelley Halpain, UC San Diego
View Media
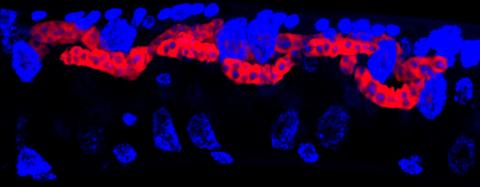
5779: Microsporidia in roundworm 3
5779: Microsporidia in roundworm 3
Many disease-causing microbes manipulate their host’s metabolism and cells for their own ends. Microsporidia—which are parasites closely related to fungi—infect and multiply inside animal cells, and take the rearranging of cells’ interiors to a new level. They reprogram animal cells such that the cells start to fuse, causing them to form long, continuous tubes. As shown in this image of the roundworm Caenorhabditis elegans, microsporidia (shown in red) have invaded the worm’s gut cells (the large blue dots are the cells' nuclei) and have instructed the cells to merge. The cell fusion enables the microsporidia to thrive and propagate in the expanded space. Scientists study microsporidia in worms to gain more insight into how these parasites manipulate their host cells. This knowledge might help researchers devise strategies to prevent or treat infections with microsporidia.
For more on the research into microsporidia, see this news release from the University of California San Diego. Related to images 5777 and 5778.
For more on the research into microsporidia, see this news release from the University of California San Diego. Related to images 5777 and 5778.
Keir Balla and Emily Troemel, University of California San Diego
View Media
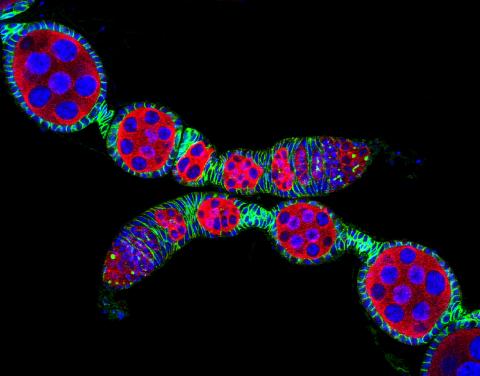
5772: Confocal microscopy image of two Drosophila ovarioles
5772: Confocal microscopy image of two Drosophila ovarioles
Ovarioles in female insects are tubes in which egg cells (called oocytes) form at one end and complete their development as they reach the other end of the tube. This image, taken with a confocal microscope, shows ovarioles in a very popular lab animal, the fruit fly Drosophila. The basic structure of ovarioles supports very rapid egg production, with some insects (like termites) producing several thousand eggs per day. Each insect ovary typically contains four to eight ovarioles, but this number varies widely depending on the insect species.
Scientists use insect ovarioles, for example, to study the basic processes that help various insects, including those that cause disease (like some mosquitos and biting flies), reproduce very quickly.
Scientists use insect ovarioles, for example, to study the basic processes that help various insects, including those that cause disease (like some mosquitos and biting flies), reproduce very quickly.
2004 Olympus BioScapes Competition
View Media
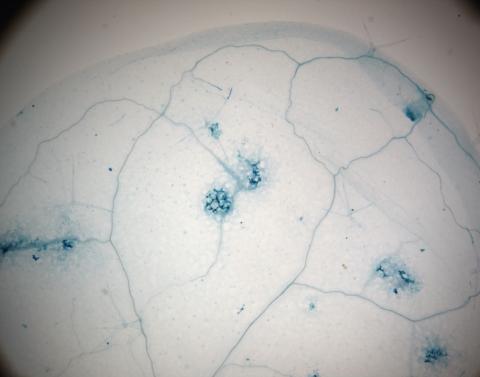
2781: Disease-resistant Arabidopsis leaf
2781: Disease-resistant Arabidopsis leaf
This is a magnified view of an Arabidopsis thaliana leaf a few days after being exposed to the pathogen Hyaloperonospora arabidopsidis. The plant from which this leaf was taken is genetically resistant to the pathogen. The spots in blue show areas of localized cell death where infection occurred, but it did not spread. Compare this response to that shown in Image 2782. Jeff Dangl has been funded by NIGMS to study the interactions between pathogens and hosts that allow or suppress infection.
Jeff Dangl, University of North Carolina, Chapel Hill
View Media
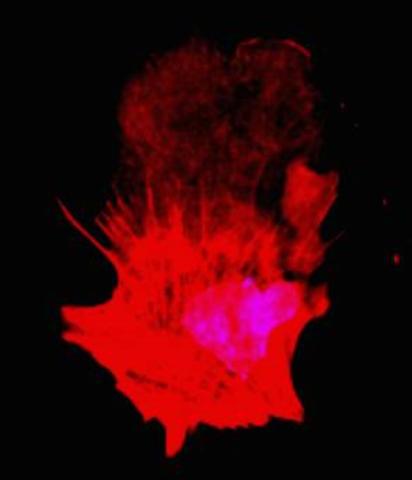
3332: Polarized cells- 01
3332: Polarized cells- 01
Cells move forward with lamellipodia and filopodia supported by networks and bundles of actin filaments. Proper, controlled cell movement is a complex process. Recent research has shown that an actin-polymerizing factor called the Arp2/3 complex is the key component of the actin polymerization engine that drives amoeboid cell motility. ARPC3, a component of the Arp2/3 complex, plays a critical role in actin nucleation. In this photo, the ARPC3+/+ fibroblast cells were fixed and stained with Alexa 546 phalloidin for F-actin (red) and DAPI to visualize the nucleus (blue). ARPC3+/+ fibroblast cells with lamellipodia leading edge. Related to images 3328, 3329, 3330, 3331, and 3333.
Rong Li and Praveen Suraneni, Stowers Institute for Medical Research
View Media
3254: Pulsating response to stress in bacteria - video
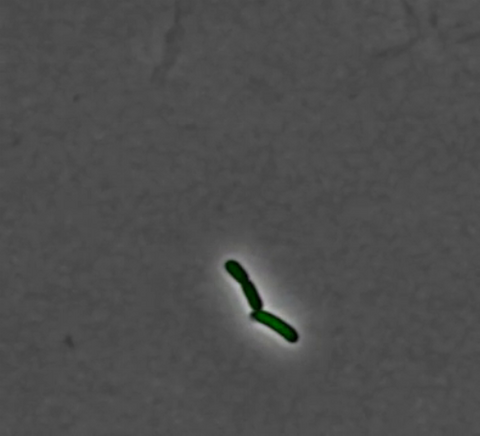
3254: Pulsating response to stress in bacteria - video
By attaching fluorescent proteins to the genetic circuit responsible for B. subtilis's stress response, researchers can observe the cells' pulses as green flashes. This video shows flashing cells as they multiply over the course of more than 12 hours. In response to a stressful environment like one lacking food, B. subtilis activates a large set of genes that help it respond to the hardship. Instead of leaving those genes on as previously thought, researchers discovered that the bacteria flip the genes on and off, increasing the frequency of these pulses with increasing stress. See entry 3253 for a related still image.
Michael Elowitz, Caltech University
View Media
3753: Coronavirus spike protein structure
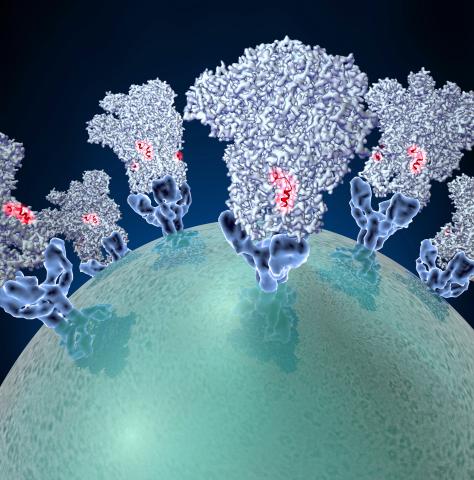
3753: Coronavirus spike protein structure
Coronaviruses are enveloped viruses responsible for 30 percent of mild respiratory infections and atypical deadly pneumonia in humans worldwide. These deadly pneumonia include those caused by infections with severe acute respiratory syndrome coronavirus (SARS-CoV) and Middle East respiratory syndrome coronavirus (MERS-CoV). The coronavirus spike glycoprotein mediates virus entry into cells and represents an important therapeutic target. The illustration shows a viral membrane decorated with spike glycoproteins; highlighted in red is a potential neutralization site, which is a protein sequence that might be used as a target for vaccines to combat viruses such as MERS-CoV and other coronaviruses.
Melody Campbell, UCSF
View Media
2649: Endoplasmic reticulum
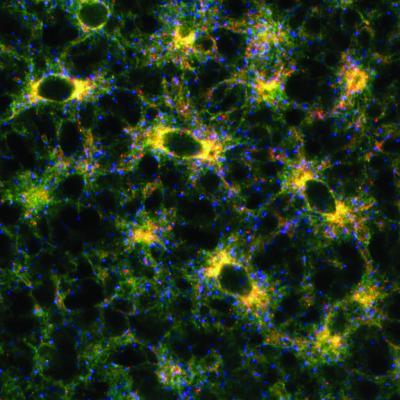
2649: Endoplasmic reticulum
Fluorescent markers show the interconnected web of tubes and compartments in the endoplasmic reticulum. The protein atlastin helps build and maintain this critical part of cells. The image is from a July 2009 news release.
Andrea Daga, Eugenio Medea Scientific Institute (Conegliano, Italy)
View Media
2528: A drug's life in the body (with labels)
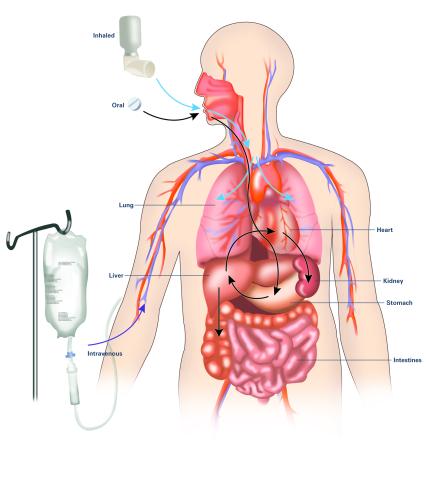
2528: A drug's life in the body (with labels)
A drug's life in the body. Medicines taken by mouth (oral) pass through the liver before they are absorbed into the bloodstream. Other forms of drug administration bypass the liver, entering the blood directly. See 2527 for an unlabeled version of this illustration. Featured in Medicines By Design.
Crabtree + Company
View Media
3408: Kluyveromyces polysporus Argonaute bound to guide RNA
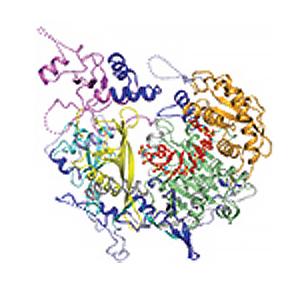
3408: Kluyveromyces polysporus Argonaute bound to guide RNA
A segment of siRNA, shown in red, guides a "slicer" protein called Argonaute (multi-colored twists and corkscrews) to the target RNA molecules.
Kotaro Nakanishi and David Weinberg, Massachusetts Institute of Technology
View Media
2534: Kinases
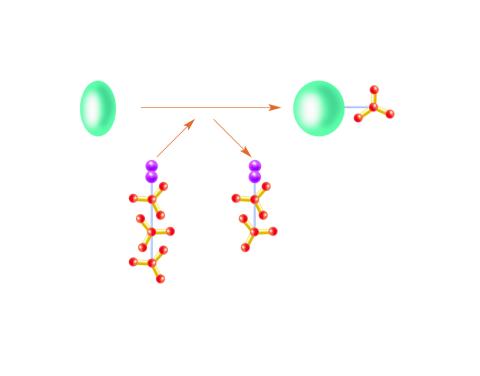
2534: Kinases
Kinases are enzymes that add phosphate groups (red-yellow structures) to proteins (green), assigning the proteins a code. In this reaction, an intermediate molecule called ATP (adenosine triphosphate) donates a phosphate group from itself, becoming ADP (adenosine diphosphate). See image 2535 for a labeled version of this illustration. Featured in Medicines By Design.
Crabtree + Company
View Media
3617: Cells keep their shape with actin filaments and microtubules
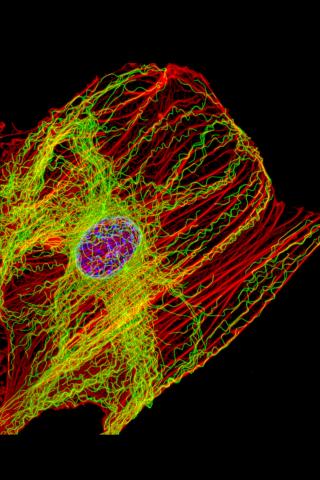
3617: Cells keep their shape with actin filaments and microtubules
This image shows a normal fibroblast, a type of cell that is common in connective tissue and frequently studied in research labs. This cell has a healthy skeleton composed of actin (red) and microtubles (green). Actin fibers act like muscles to create tension and microtubules act like bones to withstand compression.
This image was part of the Life: Magnified exhibit that ran from June 3, 2014, to January 21, 2015, at Dulles International Airport.
This image was part of the Life: Magnified exhibit that ran from June 3, 2014, to January 21, 2015, at Dulles International Airport.
James J. Faust and David G. Capco, Arizona State University
View Media
3557: Bioluminescent imaging in adult zebrafish - overhead view
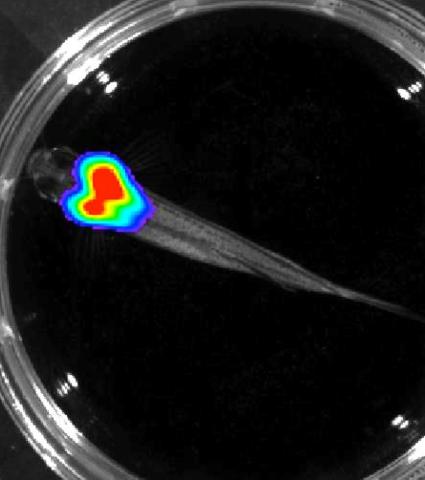
3557: Bioluminescent imaging in adult zebrafish - overhead view
Luciferase-based imaging enables visualization and quantification of internal organs and transplanted cells in live adult zebrafish. In this image, a cardiac muscle-restricted promoter drives firefly luciferase expression.
For imagery of both the lateral and overhead view go to 3556.
For imagery of the lateral view go to 3558.
For more information about the illumated area go to 3559.
For imagery of both the lateral and overhead view go to 3556.
For imagery of the lateral view go to 3558.
For more information about the illumated area go to 3559.
Kenneth Poss, Duke University
View Media
1191: Mouse sperm sections
1191: Mouse sperm sections
This transmission electron micrograph shows sections of mouse sperm tails, or flagella.
Tina Weatherby Carvalho, University of Hawaii at Manoa
View Media
2454: Seeing signaling protein activation in cells 04
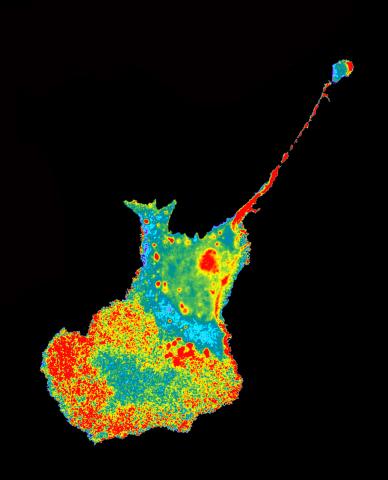
2454: Seeing signaling protein activation in cells 04
Cdc42, a member of the Rho family of small guanosine triphosphatase (GTPase) proteins, regulates multiple cell functions, including motility, proliferation, apoptosis, and cell morphology. In order to fulfill these diverse roles, the timing and location of Cdc42 activation must be tightly controlled. Klaus Hahn and his research group use special dyes designed to report protein conformational changes and interactions, here in living neutrophil cells. Warmer colors in this image indicate higher levels of activation. Cdc42 looks to be activated at cell protrusions.
Related to images 2451, 2452, and 2453.
Related to images 2451, 2452, and 2453.
Klaus Hahn, University of North Carolina, Chapel Hill Medical School
View Media
2364: High-throughput protein structure determination pipeline
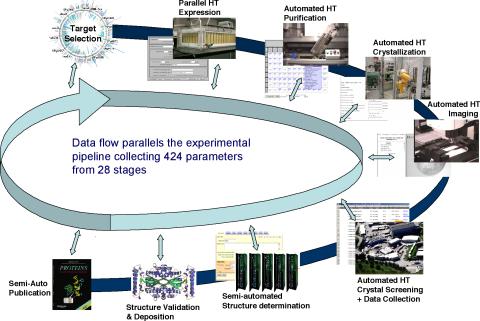
2364: High-throughput protein structure determination pipeline
This slide shows the technologies that the Joint Center for Structural Genomics developed for going from gene to structure and how the technologies have been integrated into a high-throughput pipeline, including all of the steps from target selection, parallel expression, protein purification, automated crystallization trials, automated crystal screening, structure determination, validation, and publication.
Joint Center for Structural Genomics
View Media
6999: HIV enzyme
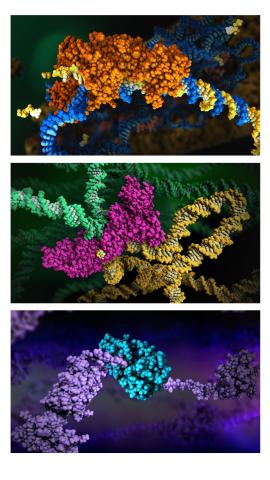
6999: HIV enzyme
These images model the molecular structures of three enzymes with critical roles in the life cycle of the human immunodeficiency virus (HIV). At the top, reverse transcriptase (orange) creates a DNA copy (yellow) of the virus's RNA genome (blue). In the middle image, integrase (magenta) inserts this DNA copy in the DNA genome (green) of the infected cell. At the bottom, much later in the viral life cycle, protease (turquoise) chops up a chain of HIV structural protein (purple) to generate the building blocks for making new viruses. See these enzymes in action on PDB 101’s video A Molecular View of HIV Therapy.
Amy Wu and Christine Zardecki, RCSB Protein Data Bank.
View Media
6963: C. elegans trapped by carnivorous fungus
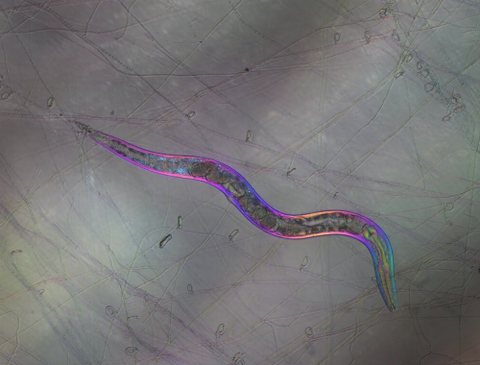
6963: C. elegans trapped by carnivorous fungus
Real-time footage of Caenorhabditis elegans, a tiny roundworm, trapped by a carnivorous fungus, Arthrobotrys dactyloides. This fungus makes ring traps in response to the presence of C. elegans. When a worm enters a ring, the trap rapidly constricts so that the worm cannot move away, and the fungus then consumes the worm. The size of the imaged area is 0.7mm x 0.9mm.
This video was obtained with a polychromatic polarizing microscope (PPM) in white light that shows the polychromatic birefringent image with hue corresponding to the slow axis orientation. More information about PPM can be found in the Scientific Reports paper “Polychromatic Polarization Microscope: Bringing Colors to a Colorless World” by Shribak.
This video was obtained with a polychromatic polarizing microscope (PPM) in white light that shows the polychromatic birefringent image with hue corresponding to the slow axis orientation. More information about PPM can be found in the Scientific Reports paper “Polychromatic Polarization Microscope: Bringing Colors to a Colorless World” by Shribak.
Michael Shribak, Marine Biological Laboratory/University of Chicago.
View Media
6985: Fruit fly brain responds to adipokines
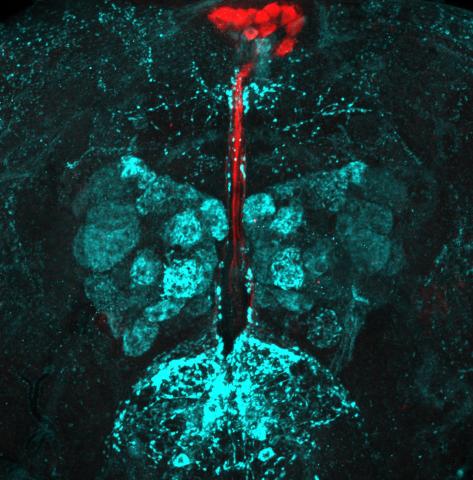
6985: Fruit fly brain responds to adipokines
Drosophila adult brain showing that an adipokine (fat hormone) generates a response from neurons (aqua) and regulates insulin-producing neurons (red).
Related to images 6982, 6983, and 6984.
Related to images 6982, 6983, and 6984.
Akhila Rajan, Fred Hutchinson Cancer Center
View Media
6553: Floral pattern in a mixture of two bacterial species, Acinetobacter baylyi and Escherichia coli, grown on a semi-solid agar for 48 hours (photo 1)
6553: Floral pattern in a mixture of two bacterial species, Acinetobacter baylyi and Escherichia coli, grown on a semi-solid agar for 48 hours (photo 1)
Floral pattern emerging as two bacterial species, motile Acinetobacter baylyi (red) and non-motile Escherichia coli (green), are grown together for 48 hours on 1% agar surface from a small inoculum in the center of a Petri dish.
See 6557 for a photo of this process at 24 hours on 0.75% agar surface.
See 6555 for another photo of this process at 48 hours on 1% agar surface.
See 6556 for a photo of this process at 72 hours on 0.5% agar surface.
See 6550 for a video of this process.
See 6557 for a photo of this process at 24 hours on 0.75% agar surface.
See 6555 for another photo of this process at 48 hours on 1% agar surface.
See 6556 for a photo of this process at 72 hours on 0.5% agar surface.
See 6550 for a video of this process.
L. Xiong et al, eLife 2020;9: e48885
View Media

3475: Automated Worm Sorter - 4
3475: Automated Worm Sorter - 4
Georgia Tech associate professor Hang Lu holds a microfluidic chip that is part of a system that uses artificial intelligence and cutting-edge image processing to automatically examine large number of nematodes used for genetic research.
Georgia Tech/Gary Meek
View Media
5770: EM of yeast cell division
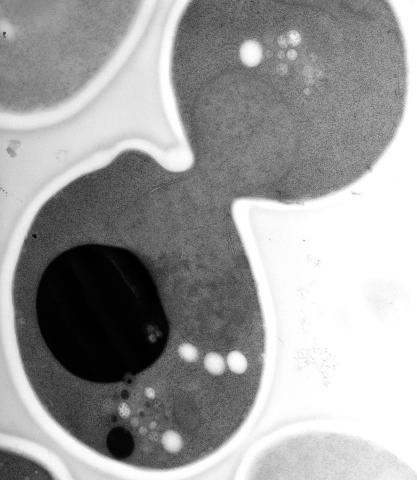
5770: EM of yeast cell division
Cell division is an incredibly coordinated process. It not only ensures that the new cells formed during this event have a full set of chromosomes, but also that they are endowed with all the cellular materials, including proteins, lipids and small functional compartments called organelles, that are required for normal cell activity. This proper apportioning of essential cell ingredients helps each cell get off to a running start.
This image shows an electron microscopy (EM) thin section taken at 10,000x magnification of a dividing yeast cell over-expressing the protein ubiquitin, which is involved in protein degradation and recycling. The picture features mother and daughter endosome accumulations (small organelles with internal vesicles), a darkly stained vacuole and a dividing nucleus in close contact with a cadre of lipid droplets (unstained spherical bodies). Other dynamic events are also visible, such as spindle microtubules in the nucleus and endocytic pits at the plasma membrane.
These extensive details were revealed thanks to a preservation method involving high-pressure freezing, freeze-substitution and Lowicryl HM20 embedding.
This image shows an electron microscopy (EM) thin section taken at 10,000x magnification of a dividing yeast cell over-expressing the protein ubiquitin, which is involved in protein degradation and recycling. The picture features mother and daughter endosome accumulations (small organelles with internal vesicles), a darkly stained vacuole and a dividing nucleus in close contact with a cadre of lipid droplets (unstained spherical bodies). Other dynamic events are also visible, such as spindle microtubules in the nucleus and endocytic pits at the plasma membrane.
These extensive details were revealed thanks to a preservation method involving high-pressure freezing, freeze-substitution and Lowicryl HM20 embedding.
Matthew West and Greg Odorizzi, University of Colorado
View Media
2725: Supernova bacteria
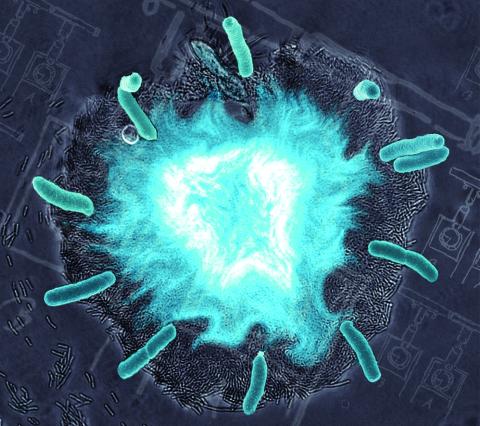
2725: Supernova bacteria
Bacteria engineered to act as genetic clocks flash in synchrony. Here, a "supernova" burst in a colony of coupled genetic clocks just after reaching critical cell density. Superimposed: A diagram from the notebook of Christiaan Huygens, who first characterized synchronized oscillators in the 17th century.
Jeff Hasty, UCSD
View Media

1336: Life in balance
1336: Life in balance
Mitosis creates cells, and apoptosis kills them. The processes often work together to keep us healthy.
Judith Stoffer
View Media
6891: Microtubules in African green monkey cells
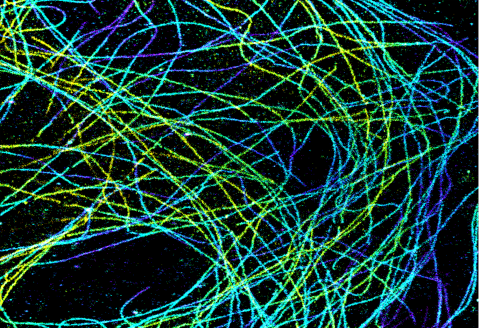
6891: Microtubules in African green monkey cells
Microtubules in African green monkey cells. Microtubules are strong, hollow fibers that provide cells with structural support. Here, the microtubules have been color-coded based on their distance from the microscope lens: purple is closest to the lens, and yellow is farthest away. This image was captured using Stochastic Optical Reconstruction Microscopy (STORM).
Related to images 6889, 6890, and 6892.
Related to images 6889, 6890, and 6892.
Melike Lakadamyali, Perelman School of Medicine at the University of Pennsylvania.
View Media
3341: Suicidal Stem Cells
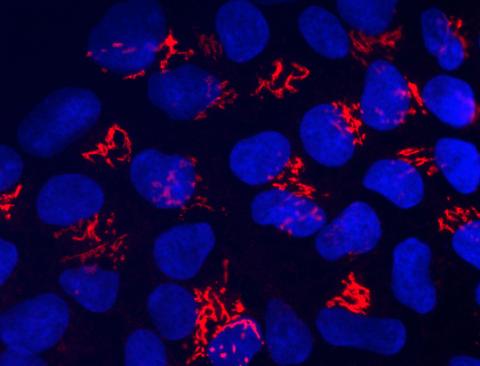
3341: Suicidal Stem Cells
Embryonic stem cells store pre-activated Bax (red) in the Golgi, near the nucleus (blue). Featured in the June 21, 2012, issue of Biomedical Beat.
Mohanish Deshmukh
View Media
1330: Mitosis - prophase
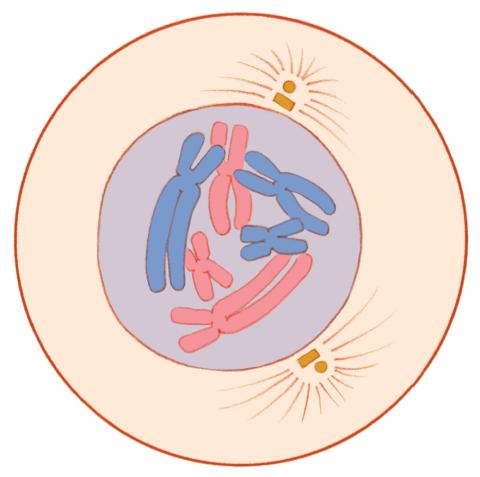
1330: Mitosis - prophase
A cell in prophase, near the start of mitosis: In the nucleus, chromosomes condense and become visible. In the cytoplasm, the spindle forms. Mitosis is responsible for growth and development, as well as for replacing injured or worn out cells throughout the body. For simplicity, mitosis is illustrated here with only six chromosomes.
Judith Stoffer
View Media
2433: Fruit fly sperm cells
2433: Fruit fly sperm cells
Developing fruit fly spermatids require caspase activity (green) for the elimination of unwanted organelles and cytoplasm via apoptosis.
Hermann Steller, Rockefeller University
View Media
3746: Serum albumin structure 3
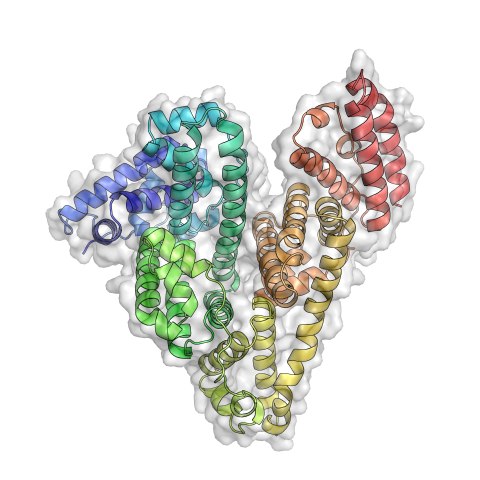
3746: Serum albumin structure 3
Serum albumin (SA) is the most abundant protein in the blood plasma of mammals. SA has a characteristic heart-shape structure and is a highly versatile protein. It helps maintain normal water levels in our tissues and carries almost half of all calcium ions in human blood. SA also transports some hormones, nutrients and metals throughout the bloodstream. Despite being very similar to our own SA, those from other animals can cause some mild allergies in people. Therefore, some scientists study SAs from humans and other mammals to learn more about what subtle structural or other differences cause immune responses in the body.
Related to entries 3744 and 3745.
Related to entries 3744 and 3745.
Wladek Minor, University of Virginia
View Media
6519: Human fibroblast undergoing cell division
6519: Human fibroblast undergoing cell division
During cell division, cells physically divide after separating their genetic material to create two daughter cells that are genetically identical to the parent cell. This process is important so that new cells can grow and develop. In this image, a human fibroblast cell—a type of connective tissue cell that plays a key role in wound healing and tissue repair—is dividing into two daughter cells. A cell protein called actin appears gray, the myosin II (part of the family of motor proteins responsible for muscle contractions) appears green, and DNA appears magenta.
Nilay Taneja, Vanderbilt University, and Dylan T. Burnette, Ph.D., Vanderbilt University School of Medicine.
View Media
2635: Mitochondria and endoplasmic reticulum
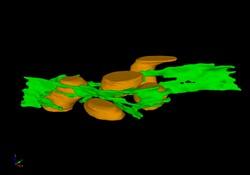
2635: Mitochondria and endoplasmic reticulum
A computer model shows how the endoplasmic reticulum is close to and almost wraps around mitochondria in the cell. The endoplasmic reticulum is lime green and the mitochondria are yellow. This image relates to a July 27, 2009 article in Computing Life.
Bridget Wilson, University of New Mexico
View Media
2808: Cell proliferation in a quail embryo
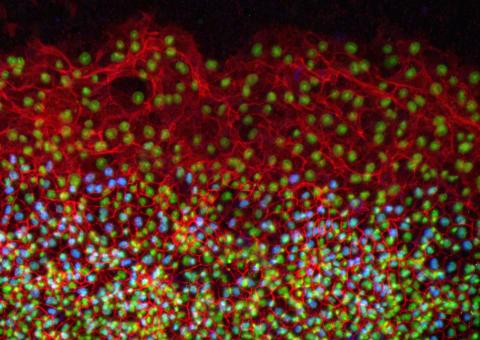
2808: Cell proliferation in a quail embryo
Image showing that the edge zone (top of image) of the quail embryo shows no proliferating cells (cyan), unlike the interior zone (bottom of image). Non-proliferating cell nuclei are labeled green. This image was obtained as part of a study to understand cell migration in embryos. More specifically, cell proliferation at the edge of the embryo was studied by examining the cellular uptake of a chemical compound called BrDU, which incorporates into the DNA during the S-phase of the cell cycle. Here, the cells that are positive for BrDU uptake are labeled in cyan, while other non-proliferating cell nuclei are labeled green. Notice that the vast majority of BrDU+ cells are located far away from the edge, indicating that edge cells are mostly non-proliferating. An NIGMS grant to Professor Garcia was used to purchase the confocal microscope that collected this image. Related to image 2807 and video 2809.
Andrés Garcia, Georgia Tech
View Media
2553: Alternative splicing (with labels)
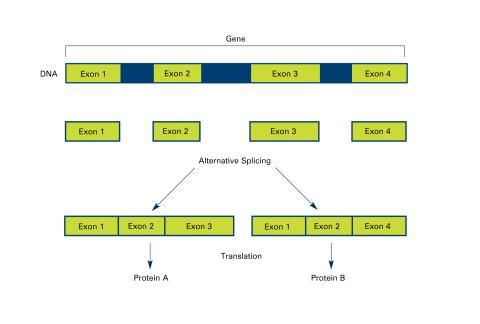
2553: Alternative splicing (with labels)
Arranging exons in different patterns, called alternative splicing, enables cells to make different proteins from a single gene. Featured in The New Genetics.
See image 2552 for an unlabeled version of this illustration.
See image 2552 for an unlabeled version of this illustration.
Crabtree + Company
View Media

6588: Cell-like compartments emerging from scrambled frog eggs 2
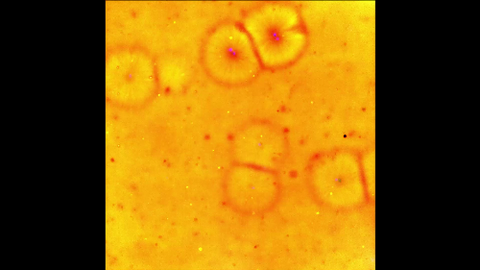
6588: Cell-like compartments emerging from scrambled frog eggs 2
Cell-like compartments spontaneously emerge from scrambled frog eggs, with nuclei (blue) from frog sperm. Endoplasmic reticulum (red) and microtubules (green) are also visible. Regions without nuclei formed smaller compartments. Video created using epifluorescence microscopy.
For more photos of cell-like compartments from frog eggs view: 6584, 6585, 6586, 6591, 6592, and 6593.
For videos of cell-like compartments from frog eggs view: 6587, 6589, and 6590.
Xianrui Cheng, Stanford University School of Medicine.
View Media
6933: Zebrafish head vasculature video
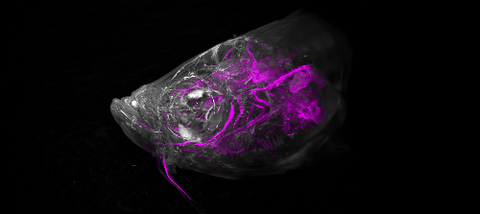
6933: Zebrafish head vasculature video
Various views of a zebrafish head with blood vessels shown in purple. Researchers often study zebrafish because they share many genes with humans, grow and reproduce quickly, and have see-through eggs and embryos, which make it easy to study early stages of development.
This video was captured using a light sheet microscope.
Related to image 6934.
This video was captured using a light sheet microscope.
Related to image 6934.
Prayag Murawala, MDI Biological Laboratory and Hannover Medical School.
View Media
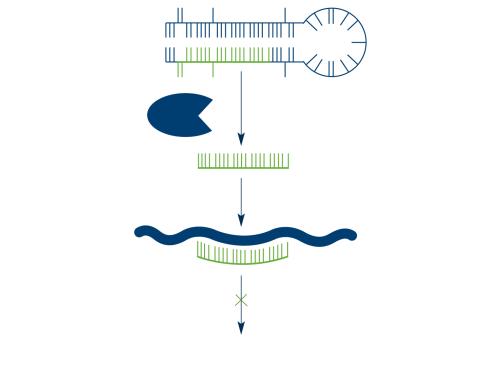
2556: Dicer generates microRNAs
2556: Dicer generates microRNAs
The enzyme Dicer generates microRNAs by chopping larger RNA molecules into tiny Velcro®-like pieces. MicroRNAs stick to mRNA molecules and prevent the mRNAs from being made into proteins. See image 2557 for a labeled version of this illustration. Featured in The New Genetics.
Crabtree + Company
View Media

6355: H1N1 Influenza Virus
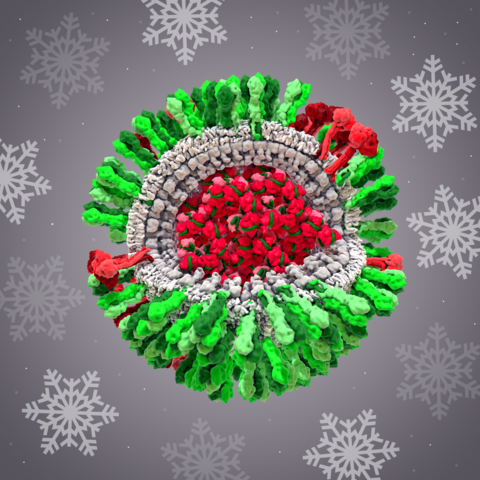
6355: H1N1 Influenza Virus
CellPack image of the H1N1 influenza virus, with hemagglutinin and neuraminidase glycoproteins in green and red, respectively, on the outer envelope (white); matrix protein in gray, and ribonucleoprotein particles inside the virus in red and green. Related to image 6356.
Dr. Rommie Amaro, University of California, San Diego
View Media
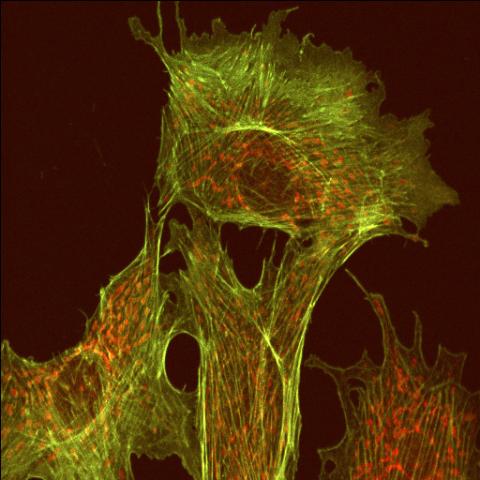
1102: Endothelial cell
1102: Endothelial cell
This image shows two components of the cytoskeleton, microtubules (green) and actin filaments (red), in an endothelial cell derived from a cow lung. The cystoskeleton provides the cell with an inner framework and enables it to move and change shape.
Tina Weatherby Carvalho, University of Hawaii at Manoa
View Media
3597: DNA replication origin recognition complex (ORC)
3597: DNA replication origin recognition complex (ORC)
A study published in March 2012 used cryo-electron microscopy to determine the structure of the DNA replication origin recognition complex (ORC), a semi-circular, protein complex (yellow) that recognizes and binds DNA to start the replication process. The ORC appears to wrap around and bend approximately 70 base pairs of double stranded DNA (red and blue). Also shown is the protein Cdc6 (green), which is also involved in the initiation of DNA replication. Related to video 3307 that shows the structure from different angles. From a Brookhaven National Laboratory news release, "Study Reveals How Protein Machinery Binds and Wraps DNA to Start Replication."
Huilin Li, Brookhaven National Laboratory
View Media
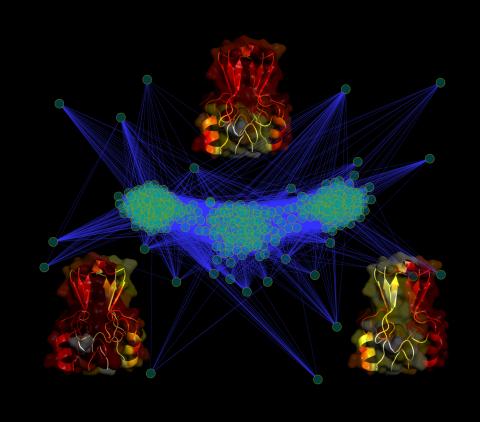
3295: Cluster analysis of mysterious protein
3295: Cluster analysis of mysterious protein
Researchers use cluster analysis to study protein shape and function. Each green circle represents one potential shape of the protein mitoNEET. The longer the blue line between two circles, the greater the differences between the shapes. Most shapes are similar; they fall into three clusters that are represented by the three images of the protein. From a Rice University news release. Graduate student Elizabeth Baxter and Patricia Jennings, professor of chemistry and biochemistry at UCSD, collaborated with José Onuchic, a physicist at Rice University, on this work.
Patricia Jennings and Elizabeth Baxter, University of California, San Diego
View Media
